Abstract
Increasing evidence has indicated that long noncoding RNAs (lncRNAs) are of great importance in different cell contexts. However, only a very small number of lncRNAs have been experimentally validated and functionally annotated during human hematopoiesis. Here, we report an lncRNA, HOTAIRM1, which is associated with myeloid differentiation and has pivotal roles in the degradation of oncoprotein PML-RARA and in myeloid cell differentiation by regulating autophagy pathways. We first revealed that HOTAIRM1 has different variants that are expressed at different levels in cells and that the expression pattern of HOTAIRM1 is closely related to that of the PML-RARA oncoprotein in acute promyelocytic leukemia (APL) patients. We further revealed that the downregulation of HOTAIRM1 could inhibit all-trans retinoic acid (ATRA) -induced degradation of PML-RARA in APL cells and repress the process of differentiation from promyelocytic to granulocytic cells. More importantly, we found that HOTAIRM1 regulates autophagy and that autophagosome formation was inhibited when HOTAIRM1 expression was reduced in the cells. Finally, through the use of a dual luciferase activity assay, AGO2 RNA immunoprecipitation and RNA pull-down, HOTAIRM1 was revealed to act as a microRNA sponge in a pathway that included miR-20a/106b, miR-125b and their targets ULK1, E2F1 and DRAM2. We constructed a human APL-ascites SCID mouse model to validate the function of HOTAIRM1 and its regulatory pathway in vivo. This is the first report showing that a lncRNAs regulates autophagy and the degradation of the PML-RARA oncoprotein during the process of myeloid cell differentiation blockade, suggesting that lncRNAs may be the potential therapeutic targets for leukemia.
Similar content being viewed by others
Main
Long noncoding RNAs (lncRNAs), which are more than 200 nt in length, are very poorly understood and often dismissed as merely transcriptional ‘noise’.1, 2 However, a number of lncRNAs have been found to have important functions in multiple major biological processes in specific cell types, tissues and developmental conditions.3, 4, 5 Of note, lncRNA transcripts has key roles in the circuitry controlling ES cell and muscle differentiation through their binding to multiple chromatin regulatory proteins or ncRNAs.6, 7, 8, 9, 10 Moreover, recent studies have found that several lncRNAs participate in the autophagy pathway. Autophagy is an evolutionarily highly conserved process for maintaining cellular homeostasis through the cellular digestion of excessive, damaged, or aged proteins and intracellular organelles within double-membrane vesicles called autophagosomes.11, 12, 13, 14 Because of its dynamic and inducible catabolic process that is activated by environmental and hormonal cues, autophagy can promote the rapid cellular changes necessary for proper differentiation.15, 16, 17, 18 For example, the lncRNAs FLJ11812 and APF regulate autophagic cell death in human umbilical vein endothelial cells and cardiomyocytes respectively.19, 20 Interestingly, lncRNAs can act as competing endogenous RNAs (ceRNAs) for microRNA (miRNA) and can in turn regulate critical cellular processes though the post-transcriptional silencing of protein-coding genes.21, 22, 23 By inhibiting the target genes of miRNAs, lncRNAs have important roles in biological processes and in cancer, including functions in muscle differentiation,8 autophagy19 and cardiovascular diseases.20
HOTAIRM1, an lncRNA located between HOXA1 and HOXA2, has been reported to be involved in the differentiation of the myeloid cell line NB4 that is induced by all-trans retinoic acid (ATRA).24 The knockdown of HOTAIRM1 selectively attenuates the induction of the differentiation gene CD11b and results in a larger population of cells that maintain cell cycle progression from G1 to S phases.24, 25 A recent study also revealed that the level of HOTAIRM1 expression at diagnosis provided relevant prognostic information for intermediate-risk acute myeloid leukemia (AML),26 with a very low level in acute promyelocytic leukemia (APL), which is divided into M3 subtype in French–American–British (FAB) classification systems. It has been shown that APL patients are characterized by a specific t (15;17) chromosomal translocation, resulting in the fusion of the promyelocytic gene (PML) and the retinoic acid receptor alpha gene (RARA) into the oncoprotein PML-RARA,27 which represses promyelocytic to granulocytic differentiation.28, 29, 30 Studies have shown that granulocytic differentiation and the degradation of PML-RARA are induced due to increased autophagic flux following treatment of APL cells with ATRA.31, 32 Therefore, we asked whether HOTAIRM1 is involved in the pathogenesis of APL, autophagy and PML-RARA protein degradation.
On the basis of annotations in the Ensembl database, we also found that other variants of HOTAIRM1 exist, suggesting that HOTAIRM1 could have a more complicated regulatory network. Given the importance of HOTAIRM1 in the differentiation of myeloid cells and the development of related diseases, we first detected and validated the existence of HOTAIRM1 variants and revealed a novel mechanism by which HOTAIRM1 regulates autophagy via a specific miRNA signature during the process of myeloid cell differentiation and the degradation of the PML-RARA oncoprotein in APL.
Results
Identification of HOTAIRM1 variants in myeloid cells
Previous studies have shown that HOTAIRM1 is a 483-nt antisense transcript in the HOXA cluster.24 However based on data from the Ensembl database, five other variants have also been identified. As shown in Figure 1a, HOTAIRM1 transcripts spliced into noncoding RNAs consisting of one, two or even three exons. Because two variant sequences were nearly identical, these six variants can be divided into five types (HOTAIRM1-1 to HOTAIRM1-5). To identify the existence of these variants, we next performed rapid amplification of cDNA ends (RACE) to experimentally validate their 5’- and 3’-ends in the NB4 cell line, and the results indeed showed different starting and ending sites (Figures 1b and c). We further investigated the relative expression of these five variants in NB4 cells following treatment with1 μM ATRA at three time points from 24 to 72 hours (h). The relative expression levels of three variants, HOTAIRM1-1, HOTAIRM1-3 and HOTAIRM1-5, were considerably higher than those of the other two variants, HOTAIRM1-2 and HOTAIRM1-4 (Figure 1d). Thus in the following experiments, we primarily focused on the HOTAIRM1-1, 3 and 5 variants.
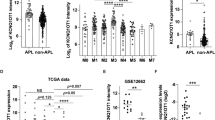
Identification of HOTAIRM1 in myeloid cells. (a) Schematic drawing of five different variants was acquired from the Ensembl online database. Five pairs of specific primers were designed for qRT-PCR. (b) 5’RACE was performed to identify HOTAIRM1 variants in NB4 cells. Two reverse primers termed R1 and R2 were designed for inner PCR to obtain the 5’-ends of HOTAIRM1 variant cDNA in cells. Those most similar to the predicted 5’-ends of HOTAIRM1 variants in Ensembl are marked by green arrows; the remainders are nonspecific products marked by red arrows. (c) The inner PCR for 3’RACE was performed by two forward primers that bind to the two specific sites of the variants. HOTAIRM1 variants are marked in green; those marked in red are nonspecific products. (d) The relative expression of five different variants was normalized to GAPDH after 24, 48 and 72 h of 1 μM ATRA treatment in NB4 cells. (e and f) The expression of three HOTAIRM1 variants and PML-RARA in 19 pairs of patients with t (15;17) APL before or after therapy. Relative expression was analyzed by the 2-△CT method normalized to GAPDH. Experiments were performed in triplicate, and the error bars represent S.D. for all panels. CR, patients in complete remission. (g) Correlation of HOTAIRM1 variants and PML-RARA expression in 19 pairs of APL patients. Experiments were performed in triplicate, and the correlation was analyzed by Pearson’s r-test
The NB4 cell line was derived from an APL patient,33 and the knockdown of HOTAIRM1 selectively attenuates NB4 cell differentiation following induction with ATRA.24, 25 Therefore, we asked whether HOTAIRM1 was associated with the therapeutic outcomes of APL patients receiving drug treatment. We tested 19 paired AML-M3 patients at diagnosis and complete remission (AML-M3-CR) and found that the levels of HOTAIRM1 expression were significantly increased in AML-M3-CR patients compared with AML-M3 patients at diagnosis (Figure 1e). This expression pattern was consistent in unpaired AML-M3 and AML-M3-CR patient samples (Supplementary Figure S1). Notably, the three variants showed the same expression patterns.
Because of the crucial importance of the PML-RARA oncoprotein in the occurrence and progression of APL, we also investigated the association of PML-RARA and HOTAIRM1 expression levels in 19 pairs of AML-M3 and AML-M3-CR patients. The results showed that PML-RARA expression levels were decreased in AML-M3-CR patients following ATRA treatment (Figure 1f) and that PML-RARA expression negatively correlated with the expression of the three HOTAIRM1 variants (Figure 1g). Thus, we hypothesized that the downregulation of HOTAIRM1 may be important for the effects of PML-RARA on cell differentiation during APL or may affect therapy-induced clearance of PML-RARA.
HOTAIRM1 enhances ATRA-induced degradation of PML-RARA in APL cells
The above observation revealed a close association between the expression of HOTAIRM1 and PML-RARA, suggesting that both may have significant roles in the occurrence of APL. We therefore asked whether PML-RARA could directly regulate HOTAIRM1 or whether PML-RARA was regulated by HOTAIRM1. First, we used the U937-PR9 cell line, which carries a PML-RARA gene that can be induced with ZnSO4 treatment. However, when PML-RARA expression was induced by 200 μM Zn2+ for 12 h, no apparent effect on HOTAIRM1 expression was observed (Figure 2a and Supplementary Figure S2A). We then used siRNA to decrease PML-RARA expression in the NB4 cell line, and we again failed to observe a significant effect on the relative expression of HOTAIRM1 (Figure 2b). These data indicated that HOTAIRM1 is not regulated by PML-RARA. Next, to determine whether HOTAIRM1 could regulate PML-RARA, we established a long-term, stable shRNA lentiviral vector with an effective shRNA sequence targeting the common regions of the three variants of HOTAIRM1 (hereafter termed NB4-Lv-shHOTAIRM1 cells or NB4-Lv-NC cells) (Supplementary Figure S2B). We then investigated whether the inhibition of HOTAIRM1 could attenuate the ATRA-induced clearance of PML-RARA. It was shown that knockdown of HOTAIRM1 could significantly increase PML-RARA protein levels (Figure 2c). Interestingly, the knockdown of HOTAIRM1 could also slightly increase PML-RARA mRNA level (Supplementary Figure S2C), suggesting that HOTAIRM1 might also affect PML-RARA mRNA expression, which deserve to be further demonstrated. Furthermore, we designed two effective siRNAs targeting HOTAIRM1 to detect the PML-RARA protein levels in ATRA-induced NB4 cells (Supplementary Figure S2D and S2E). Wright–Giemsa staining showed that NB4-Lv-shHOTAIRM1 cells lack ATRA-induced changes in nuclear morphology (Figure 2d). Consistently, the expression of the granulocytic differentiation cell-surface marker CD11b was decreased in NB4-Lv-shHOTAIRM1 cells, as measured by flow cytometry (FCM) (Figure 2e). These data show that low levels of HOTAIRM1 in APL patients may be linked to the PML-RARA and granulocytic differentiation. However, it is not clear how HOTAIRM1 regulates PML-RARA status, nor is it clear which pathways may be involved.
HOTAIRM1 is associated with PML-RARA oncoprotein and APL differentiation following therapeutic treatment. (a) HOTAIRM1 expression was not affected by increased PML-RARA following a 12 h 200 μM ZnSO4 induction in U937-PR9 cells. (b) HOTAIRM1 was not influenced by the knockdown of PML-RARA 48 h after the transfection of NB4 cells with an effective siRNA. (c) Western blot analysis of PML-RARA in NB4-Lv-NC and NB4-Lv-shHOTAIRM1 cells treated with ATRA. The PML-RARA/GAPDH densitometric ratio was recorded by quantity one. Quantification of data from three independent experiments (right) (*P<0.05). (d) Wright–Giemsa staining was performed to evaluate the differentiation of NB4 cells 48 h after ATRA treatment when HOTAIRM1 was knocked down; staining was observed under a × 40 objective lens. Scale bar, 10 μm. (e) Flow cytometric analysis of iTGAM/cD11b cell-surface expression in NB4-Lv-NC or NB4-Lv-shHOTAIRM1 cells followed by treatment with or without ATRA. Values are derived from three independent experiments, and data are reported as the mean±S.D. (**P<0.01)
Analysis of the lncRNA–miRNA interaction indicates that HOTAIRM1 promotes PML-RARA degradation via an autophagy pathway
Recent progresses have indicated that lncRNAs can act as ceRNAs or natural miRNA sponges that can communicate with and coregulate each other by competitive binding to shared miRNAs.8, 19, 20, 34 Notably, a previous study reported that HOTAIRM1 expression correlated with a specific miRNAs signature in AML patients.26 Thus, we hypothesized that HOTAIRM1 may act as a ceRNA in myeloid cells. Using DIANA TOOLS software,35, 36 together with assessing △G by BiBiServ2-RNAhybrid,37 we predicted the potential binding sites of HOTAIRM1 on various miRNAs. As shown in Table 1, three miRNAs, miR-20a, miR-106b and miR-125b, were predicted to be the potential miRNAs targeting of HOTAIRM1. Importantly, previous studies have demonstrated that these three miRNAs are all involved in autophagy-related pathways through the direct targeting of autophagy-associated genes.38, 39 Given the importance of autophagy pathway in the degradation of PML-RARA and myeloid differentiation,16, 32, 31 We therefore hypothesized that HOTAIRM1 may function through an miRNA-associated autophagy pathway.
To validate this hypothesis, we used NB4-Lv-NC and NB4-Lv-shHOTAIRM1 cells to investigate the effects on autophagy. Ultrastructural analysis as detected by transmission electron microscopy (TEM) revealed that autophagosome formation was inhibited in NB4-Lv-shHOTAIRM1 cells, especially after 1 μM ATRA was added for 48 h (Figure 3a). To further test autophagic flux, we performed immunofluorescent labeling for LC3B, and evaluated the results by laser scanning confocal microscopy. The results also provided evidence that autophagy was attenuated in NB4-Lv-shHOTAIRM1 cells in the presence of ATRA, which normally induces autophagy dramatically (Figure 3b). The ratio of LC3B cleavage and turnover (LC3B-II) was also measured (Supplementary Figure S3A), which is widely regarded as an important hallmark for monitoring autophagic flux.13, 40 To further validate the effect on autophagy-lysosomal biogenesis, LC3B-II, GABARAP-II and p62 expression levels were detected by western blotting (Figures 3c and e and Supplementary Figure S3B). The protein levels of both LC3B-II and GABARAP-II (mRNA of which was not regulated by HOTAIRM1, Supplementary Figure S3C) were obviously repressed in NB4-Lv-shHOTAIRM1 cells with ATRA treatment (Figures 3c, d and Supplementary Figure S3D), and then the GABARAP-II also significantly changed when co-treated with ATRA and bafilomycin A1 (an autophagy pathway inhibitor) (Figure 3d, lanes 5 and 6), although LC3B-II was slight (Figure 3c).
HOTAIRM1 regulates the autophagy pathway. (a) The ultrastructure of NB4-Lv-NC and NB4-Lv-shHOTAIRM1 cells was observed by transmission electron microscopy after 1 μM ATRA treatment for 48 h. The number of autophagosomes observed by transmission electron microscopy was calculated (right, n= 10 cells). Scale bar, 2 μm. (b) Laser scanning confocal microscopy after staining with an LC3B antibody. NB4-lv-NC and NB4-Lv-shHOTAIRM1 cells treated with 1 μM ATRA for 48 h. The number of LC3B puncta per cell was calculated by Image-Pro Plus (right, n=100 cells) (***P<0.001). Scale bar, 10 μm. A representative image from three independent experiments is shown. (c–e) Western blot analysis of LC3B, GABARAP and p62 in NB4-Lv-NC and NB4-Lv-shHOTAIRM1 cells treated with ATRA and Bafilomycin A1. The LC3B-II/GAPDH, p62/GAPDH and GABARAP-II/GAPDH densitometric ratio were recorded by quantity one
To further validate the function of HOTAIRM1 regulating autophagic flux in cells, we subsequently employed the exogenous LC3B (Supplementary Figure S3E) by the lentivirus system and performed the tandem mRFP-GFP-LC3 florescence analysis.41 As shown in Figure 4a and Figure 4b, mRFP-LC3B dots per cell significantly decreased in ATRA-induced NB4 cells transfected with siRNAs targeting HOTAIRM1 than that in the negative controls, while, co-treatment with ATRA and bafilomycin A1, the accumulated yellow dots (due to the increased pH by bafilomycin A1 in autolysosomes) significantly reduced after HOTAIRM1 knockdown. Moreover, the protein level of GFP tagged LC3B-II levels was apparently inhibited in mRFP-GFP-LC3 NB4 cells transfected with siRNAs under ATRA or bafilomycin A1 treatment (Figure 4c). These results together suggested that HOTAIRM1 could involve in the process of autophagosome formation.
HOTAIRM1 regulates exogenous LC3B and contributes to the degradation of PML-RARA. (a) siRNAs targeting HOTAIRM1 were transient transfected into stable mRFP-GFP-LC3 NB4 cells which were treated with ATRA (1 μM, 48 h) and bafilomycin A1 (25 nM, 12 h). Cells were then fixed with 4% PFA followed by confocal microscopy. Scale bar, 5 μm. (b) The number of yellow LC3 dots and red LC3 dots per cell in each condition was quantified by Image-Pro Plus. Total LC3 dots were the addition of the number of yellow LC3 dots with red LC3 dots. More than 30 cells were counted in each condition and data (mean±S.D.) were representative of three independent experiments. (c) Western blotting with antibody GFP after transient transfection of siRNAs targeting HOTAIRM1 in stable mRFP-GFP-LC3 NB4 cells treated with ATRA (1 μM, 48 h) and bafilomycin A1 (25 nM, 12 h). Total lysates were prepared and subjected to immunoblot analysis. The LC3B-II/GAPDH densitometric ratio was recorded by quantity one. (d) Autophagy regulates the ATRA-induced degradation of PML-RARA. NB4-Lv-NC and shHTOAIRM1 cells were treated with ATRA (1 μM, 48 h) and either the autophagy inducer rapamycin (100 nM, 20 h) or the autophagy inhibitors chloroquine (40 μM, 6 h) and bafilomycin A1 (25 nM, 12 h). PML-RARA was assayed by western blotting, and GAPDH expression served as a loading control. (e) The PML-RARA/GAPDH densitometric ratio was recorded by quantity one. Quantification of data from three independent experiments, and the data was reported as the mean±S.D.
Furthermore, two autophagy pathway inhibitors (chloroquine and bafilomycin A1) and one autophagy inducer (rapamycin) were employed to test PML-RARA expression in NB4-Lv-NC and NB4-Lv-shHOTAIRM1 cells with or without ATRA treatment. The results showed that PML-RARA was decreased in NB4-Lv-shHOTAIRM1 cells following dual treatment with ATRA and rapamycin; however, PML-RARA was increased following co-treatment with chloroquine or bafilomycin A1 (Figures 4d and e). Together, these results support the hypothesis that the downregulation of HOTAIRM1 is associated with attenuation in autophagy and retardation in the degradation of PML-RARA in NB4 cells.
HOTAIRM1, a natural sponge, regulates the expression of autophagy-associated genes by competing with miRNA-binding sites
The data suggested that HOTAIRM1 enhances PML-RARA degradation by regulating autophagosome formation, and bioinformatic analyses have suggested that HOTAIRM1 may function as a ceRNA. To validate that HOTAIRM1 functions as a ceRNA in autophagy pathway, we first performed an RNA-protein pull-down assay and showed that AGO2 (a component of miRNA-silencing complex) was detected in the protein complex binding with HOTAIRM1 (Figure 5a). We also performed an RNA immunoprecipitation assay to uncover the RNA that bound to AGO2, and the results showed that HOTAIRM1 was considerably enriched within the precipitate (Figure 5b and Supplementary Figure S4A), indicating that HOTAIRM1 indeed binds to AGO2.
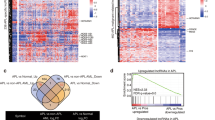
HOTAIRM1 regulates autophagy-associated gene expression by competing with the binding sites of miRNAs. (a) HOTAIRM1 transcripts bind to AGO2 directly. AGO2 was assayed by western blotting after acquiring the possible protein complex binding to HOTAIRM1. Proteins bound to the antisense of HOTAIRM1 served as loading controls. (b) RNA immunoprecipitation was performed to acquire the RNA that interacted with AGO2 protein. The qRT-PCR product of HOTAIRM1 was tested by agarose gel electrophoresis. (c) Luciferase reporter assays analyzing the binding of HOTAIRM1 to miRNAs. NC and miRNAs duplexes were co-transfected with psiCHECH-2 plasmids containing the 59 nt of HOTAIRM1-WT or HOTAIRM1-MUT. The firefly luciferase activity of each sample was normalized to the Renilla luciferase activity (n=3 independent experiments performed in triplicate). (d) Luciferase reporter assays analyzing the genes potentially influenced by HOTAIRM1. NC and two HOTAIRM1-siRNAs were co-transfected with psiCHECH-2 plasmids with the 59-nt of miRNAs targeting the wild-type or mutant gene position in HEK-293 T cells. The firefly luciferase activity of each sample was normalized to the Renilla luciferase activity. Values are derived from n=3 independent experiments, and data are reported as the mean±S.D. (e) Western blotting detecting the proteins expression levels of genes regulated by HOTAIRM1. ULK1, E2F1 and DRAM2 were all downregulated when HOTAIRM1 was knocked down in NB4 cells with or without 1 μM ATRA treatment; GAPDH was used as a loading control. The densitometric ratio normalized to GAPDH was recorded by quantity one. Values are derived from n=3 independent experiments, and data are reported as the mean±S.D. (down). (f–h) The qRT-PCR testing of DRMA1, LC3B and ULK1, three direct targets of E2F1. Experiments were performed in triplicate and are reported as the mean±S.D. (i) The qRT-PCR testing of miRNAs in NB4-Lv-NC and NB4-Lv-shHOTAIRM1 cells after treatment with ATRA (1 μM, 48 h). Experiments were performed in triplicate and normalized to GAPDH. (j) The detection of PML-RARA by western bloting after transient overexpression of miR-20a and miR-106b mimics in NB4 cells. Experiments were performed in triplicate. PML-RARA/GAPDH densitometric ratios were recorded. Differences in c-i were considered significant at *P<0.05, **P<0.01 and ***P<0.001
To validate whether HOTAIRM1 directly targets the three miRNAs, we performed a dual luciferase reporter assay by inserting 59-nt sequences (the fragment of HOTAIRM1 that binds to the mature miR-20a/106b and miR-125b sequences) and mutated sequences of the miRNA seed regions into psiCHECK-2 vectors (Supplementary Figure S4B). As shown in Figure 5c, the activity of HOTAIRM1 was significantly reduced compared with the mutated HOTAIRM1 when miR-20a, miR-106b and miR-125b were transiently transfected into HEK-293 T cells.
Previous studies have shown that ULK1, E2F1 and DRAM2 are important targets of miR-20a, miR-106b and miR-125b, respectively, and are indispensable genes for the autophagy process.15, 38, 39, 42, 43, 44 Thus, we performed a dual luciferase reporter assay by co-transfecting HEK-293 T cells with HOTAIRM1-siRNAs, (Supplementary Figure S2D), and a psiCHECK-2 vector with 59-nt 3’-untranslated regions) sequences of DRAM2, ULK1 or E2F1 inserted (Supplementary Figure S4C); the cells expressed endogenous HOTAIRM1 and the three miRNAs (Figure 5d and Supplementary Figure S4D). Western blot analyses and the immunofluorescence experiment showed that the knockdown of HOTAIRM1 inhibited the translated levels of the miRNA targeting genes (Figure 5e and Supplementary Figure S4E). The mRNA levels of DRAM1, LC3B and ULK1, which were regulated by E2F1 directly,43 were also reduced following the knockdown of HOTAIRM1 in the NB4 cell line (Figures 5f and h). A ChIP assay was further performed and the results showed that the binding of E2F1 on LC3B promoter was indeed reduced in ATRA-induced NB4-Lv-shHOTAIRM1 cells (Supplementary Figure S4F). The promoter activity assay also showed that the luciferase activity of LC3B promoter was reduced notably when knockdown of HOTAIRM1 in HEK-293 T cells (Supplementary Figure S4G), all together suggesting that HOTAIRM1 could regulate the autophagy genes and their downstream signaling pathway through an miRNA fine tune mechanism.
Finally, we investigated whether HOTAIRM1 promotes PML-RARA degradation via these miRNAs. We detected the levels of miRNAs in NB4-Lv-NC and NB4-Lv-shHOTAIRM1 cells after treatment with ATRA by quantitative real-time PCR (qRT-PCR); the levels of the miRNAs were increased in the NB4-Lv-shHOTAIRM1 cells (Figure 5i), which were responsible for the increase of PML-RARA protein levels after the transient overexpression of miR-20a and miR-106b mimics and miR-125b in NB4 cells (Figure 5j and ref. 39). Thus, it can be concluded that HOTAIRM1 regulates PML-RARA degradation through an miRNA-mediated pathway that inhibits autophagy-related genes.
The inhibition of HOTAIRM1 blocks PML-RARA autophagic degradation and promyelocyte differentiation in vivo
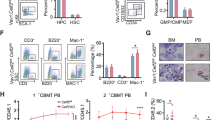
HOTAIRM1 has been shown to be responsible for PML-RARA proteolysis and may be involved in the treatment of APL. For the purpose of furthering our understanding of the function of HOTAIRM1 in APL pathogenesis, we established a human APL-ascites SCID mouse model to reveal the function of lncRNA in disease therapy. As shown in Figure 6a, we obtained the APL-ascites SCID mice 20 days after injecting 1 × 106 NB4, NB4-Lv-NC or NB4-Lv-shHOTAIRM1 cells. Then, we withdrew the ascetic fluid and cultured the leukemic cells in RPMI 1640 medium with 1 μM ATRA for 48 h (named NB4-M, NC-M and shRNA-M). Next, we performed western blotting, FACS analysis and Wright–Giemsa stains. As shown in Figure 6b, LC3B-II, a molecular marker of autophagosome formation, was strongly reduced in shRNA-M cells compared with NC-M cells, and this effect is more apparent following ATRA treatment. Consistent with the attenuation of autophagy, we observed a simultaneous accumulation of PML-RARA in shRNA-M cells (Figure 6c). Furthermore, we tested the level of CD11b expression by FACS analysis to monitor cell differentiation (Figure 6d), and we also performed Wright–Giemsa staining to observe the nuclear types (Figure 6e). These data demonstrated that HOTAIRM1 is involved in the auto-lysosome pathway during ATRA-induced APL eradication. Collectively, we established a complex network including lncRNAs, miRNAs, auto-lysosome formation and PML-RARA degradation (Figure 7).
Downregulation of HOTAIRM1 in a human APL-ascites SCID mouse model. (a) Schematic representation of the ascites induced by injecting NB4, NB4-Lv-NC and NB4-Lv-shHOTAIRM1. (b and c) Western blot analysis of LC3B and PML-RARA in NB4-Lv-NC and NB4-Lv-shHOTAIRM1 ascites cells of mice before and after treatment with 1 μM ATRA for 48 h. LC3B-II/GAPDH and PML-RARA/GAPDH densitometric ratio were recorded by quantity one. (d) Flow cytometric analysis of iTGAM/CD11b cell-surface expression in NB4-Lv-NC and NB4-Lv-shHOTAIRM1 injected ascites cells of NOD-SCID mice followed by treatment with or without ATRA. Experiments were performed in triplicate and are reported as the mean±S.D. (***P<0.001) (e) Wright–Giemsa staining was performed to evaluate the differentiation of ascites cells 48 h following ATRA treatment; staining was observed under a × 40 objective lens. Scale bar, 10 μm
Discussion
Recently, studies have shown that lncRNAs function in major biological processes, including embryonic stem (ES) cell pluripotency, chromatin remodeling and cancer progression and development.3, 6, 45, 46, 47 Although a number of lncRNAs have been identified, only a very few have been experimentally validated and functionally annotated in human hematopoiesis.48 HOTAIRM1 has been reported to be involved in myeloid differentiation and associated with the prognosis and classification of myeloid leukemia; however, the underlying mechanism has not been determined. In this study, we first showed that the expression pattern of HOTAIRM1, variants with differing expression levels in NB4 cells, is closely related to the expression of the PML-RARA in APL patients. Importantly, we demonstrated that HOTAIRM1 regulated the degradation of PML-RARA by binding to endogenous miRNAs that repress the autophagy pathway. This is the first time that an lncRNA can have an important role in ATRA-induced APL differentiation, suggesting that lncRNAs may be a potential leukemia therapeutic target.
The evolutionary study of lncRNA repertoires has revealed that lncRNAs do not exhibit a high interspecies conservation as protein-coding genes.49 Furthermore, RNA secondary and tertiary structures may be preserved throughout evolution and may serve as functional units, which may explain the lack of sequence conservation among many lncRNAs.50, 51 A previous study has shown that the complex transcriptional activity of HOTAIRM1 among species breaks homology at the DNA sequence level.24 In this study, we found various additional transcriptional patterns of HOTAIRM1 with many variant forms within a single species. Although HOTAIRM1 was less conserved, we found a surprisingly high level of conservation within the miRNA-binding regions of HOTAIRM1 variants within human cells. This conservation of other lncRNAs has also been observed in other studies.19, 8, 52, 20 Furthermore, the minimum free energy (mfe) of RNA-RNA hybridizations calculated based on secondary structures by RNAhybrid analysis37 showed that HOTAIRM1 and miR-20a/106b or miR125b had effective and convincible interactions. This variant analysis indicated that the biologically functional domain was determined by the common region, the last exons or the 3’-end of HOTAIRM1. The functions of the three variants in regulating PML-RARA degradation through autophagy strongly support this conserved cellular function of HOTAIRM1.
The PML-RARA oncoprotein is of crucial importance both pathologically and therapeutically in APL.53, 54, 55 PML-RARA can bind to the promoter regions of target genes and also have a rather diverse repertoire of binding sites.56 Recently published data have provided evidence that 14q32 hypermethylation is implicated in the pathogenesis of APL and leads to the overexpression of miRNAs clustered on chromosome 14q32.57 Furthermore, a group of miRNAs that targets primarily cancer proteins is repressed directly by PML-RARA.58 In this study, we showed that HOTAIRM1 regulated PML-RARA degradation through the autophagy pathway. This is the first report that an lncRNA is involved in ATRA-induced PML-RARA degradation, and this finding provides a new potential therapeutic target for APL.
Recent studies have found that several lncRNAs derived from different transcriptional elements of the genome participate in the regulation of the auto-lysosome pathway. Of note, APF, an antisense lncRNA derived from the Pdpr gene coding region, regulates autophagic cell death in cardiomyocytes through ATG7.20 Moreover, FLJ11812, a 3’-untranslated region-associated lncRNA, could promote autophagy exposed to a novel autophagy inhibitor, 3-benzyl-5-((2-nitrophenoxy) methyl)–dihydrofuran-2(3 h)-one (3BDO).19 In contrast with a previous report, our study showed that HOTAIRM1 is an autophagy-related lncRNA derived from intergenic regions in the genome, implying that autophagy-associated lncRNAs could be derived from 3’-untranslated region and antisense of genes and also long intergenic noncoding RNAs. However, lncRNAs from other regions, introns or promoter-associated lncRNAs remain to be discovered. Therefore, further exploration to discover additional autophagy-related lncRNAs in the genome is required. Furthermore, previous data have shown that ceRNA transcripts, which derived from protein-coding or noncoding RNA transcripts, are able to cross-regulate other transcripts that share common response elements of miRNAs.52 Although a set of lncRNAs have been reported to be ceRNAs, none were reported to have a role in oncoprotein degradation through autophagic pathways. This study is the first to show that HOTAIRM1, a natural sponge for three miRNAs that targets three protein-coding genes, cross-regulated ULK1 and E2F1 by an miRNA-mediated silencing pathway and controlled the level of autophagy.
It has been demonstrated that in mouse ES cells, the knockdown of lncRNAs induces specific early differentiation, suggesting that lncRNAs are normally a barrier to this differentiation.6 However in the specific terminal differentiation lineages, very few lncRNAs have been implicated in this complex process. Interestingly, the two ceRNAs H19 and linc-MD1 show conservation of function during muscle differentiation, repressing differentiation and correlating with the anticipation of the differentiation program, respectively.9, 8 It was controversial whether or not the functions of lncRNAs in two different stages of specific differentiation lineages were consistent. In this study, the knockdown of HOTAIRM1 caused retardation of myeloid cell differentiation, which was consistent with the function of linc-MD1 but different from the role of lncRNAs in specific early differentiation lineages. The functions of lncRNAs in different differentiation stages are largely unclear. Moreover, a previous study showed that several lncRNAs were implicated in genetic disorders and cancers that were caused by a blockade in differentiation.46, 59, 60 Autophagy has an important role in the processes of myeloid cell differentiation and PML-RARA degradation in APL.16, 32, 31 Our study is the first to show that HOTAIRM1, an lncRNA involved in the autophagic program, accelerated granulocytic differentiation during therapeutic treatment of APL, revealing a novel pathway through which lncRNAs can affect differentiation.
In summary, we provide a new regulatory network involving HOTAIRM1 and miRNAs mediated autophage pathways in the degradation of oncoprotein. HOTAIRM1 functions as a ceRNA to mediate significant autophagy-associated genes through an miRNA pathway. More importantly, we revealed that the up-regulated expression of HOTAIRM1 could enhance ATRA-induced PML-RARA degradation by affecting autophagic flux and could control myeloid cell differentiation in APL cells. Conclusively, our work indicates that HOTAIRM1 is an lncRNA that may be a novel potential therapeutic target for leukemia.
Materials and Methods
Patients and samples
A total of 79 bone marrow samples, including 54 APL samples at diagnosis and 25 APL samples after therapy, from the First and Second Affiliated Hospital of Sun Yat-sen University were enrolled in this study. Patient characteristics are shown in Table 2. Informed consent to perform the biological studies was obtained from parents and guardians. This study was approved by the ethics committee of the affiliated hospitals of Sun Yat-sen University.
Cell lines and cultures
Leukemia cells (NB4 and U937-PR9cells) and HEK-293 T cells were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA.) and were cultured in RPMI Medium Modified (HyClone, South Logan, UT, USA) and DMEM (HyClone), respectively, containing 10% fetal bovine serum (Gibco, ThermoFisher Scientific, Waltham, MA, USA). The cells were cultured in a humidified atmosphere containing 5% CO2 at 37 °C. ATRA (Sigma-Aldrich, St. Louis, MO, USA) was dissolved in ethanol and used at a final concentration of 1 μM. Rapamycin (Sigma-Aldrich) and Bafilomycin A1 (Sangon Biotech, Shanghai, China) were dissolved in DMSO with the final concentrations of 100 nM and 25 nM, respectively. Chloroquine (Sigma-Aldrich) was dissolved in ultrapure water at a final concentration of 40 μM. Wright-Giesma staining was used to test for changes in cell morphology.
FACS analysis
Cell differentiation was assessed by detecting the surface antigen expression of ITGAM/CD11b (eBioscience, San Diego, CA, USA) by flow cytometric analysis using a BD FACSAria cytometer (BD Biosciences, San Jose, CA, USA).
5’RACE and 3’RACE
Total RNA from NB4 cells was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s guidelines. Then, the 5’- and 3’-ends of cDNA were acquired using a 5’-Full RACE Kit with TAP (Takara, Dalian, Liaoning, China) and a 3’-Full RACE Core Set with PrimeScript RTase (Takara), respectively, in accordance with the manufacturer’s instructions. PCR products were obtained and then cloned into pEASY-T3 (TransGen Biotech, Beijing, China) for further sequencing.
DIANA TOOLS and BiBiServ2 analysis
The online lncRNA databases DIANA35, 36 and BiBiServ2 (ref. 37) were used to predict the relationships between lncRNAs and miRNAs. Transcripts of HOTAIRM1 termed n410904 from the NONCODE database (http://www.noncode.org/index.php) were used to search the predicted targets in the LncBase of DIANA (http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=lncBase/index) under a threshold set to 0.85, and the results were further validated in RNAhybrid of the BiBiServ2 database(http://bibiserv.techfak.uni-bielefeld.de/), which has been established based on classical RNA secondary structure prediction algorithms.
RNA isolation and quantitative real-time PCR
Real-time PCR was performed to quantify mRNA expression using SYBR Premix Ex Taq (TliRNaseH Plus) (Takara) according to the manufacturer’s instructions. The qRT-PCR was performed to detect mature miRNAs and HOTAIRM1 expression. Briefly, RNA was reverse-transcribed to cDNA using a PrimeScript RT reagent kit with gDNA Eraser (Perfect Real Time) (Takara) and amplified with specific miRNA RT and PCR amplification primers. U6 and GAPDH served as internal controls. For PML-RARA, primers were designed as previously reported.61 PML breakpoints were rapidly assigned by qRT-PCR using a common PML-RARA reverse primer in conjunction with PML intron 3 (bcr3) breakpoint-specific forward primers. The oligonucleotide sequences are shown in Supplementary Table S1.
Luciferase constructs
A human LC3B promoter fragment, demonstrated in previous work, was amplified from NB4 genomic DNA, verified by sequencing and cloned into pGL3-basic (Promega, Madison, WI, USA) to generate the construct. The 3’-UTR segment comprising 59 bp of the HOTAIRM1, E2F1, DRMA2 or ULK1 3’-UTR was synthesized and inserted into the psiCHECK2 control vector (Promega) for the miRNA functional analysis. The core sequence of HOTAIRM1 wild-type targeting miR-125b and miR-20a/106b was 5′-UGGAGUUGGGGGUUUCUGUAGGCACUUUA-3′. The targets point mutated vectors were mutated at five points of the 8 bp sequences complementary to the seed sections of miRNAs. The mutated core sequence of HOTAIRM1 targeting miR-125b and miR-20a/106b were 5′-GUUUGAGUUCCC-3′ and 5′-UUUCUGUAGCGUGAUUA-3′ respectively (bold bases were mutated). All the wild types and point mutations were confirmed by DNA sequencing. The lentivirus carrying mRFP-GFP-LC3 was purchased from Hanbio Co. Ltd (Hanbio, Shanghai, China).
Transient transfection
For the luciferase reporter assay, HEK-293 T cells were seeded in 96-well sterile plastic culture plates at a density of 1 × 104 cells per well with complete growth medium. The cells were transfected using Lipofectamine 3000 (Invitrogen) according to the manufacturer’s protocol. The following were purchased from Guangzhou RiboBio Co., Ltd (Guangzhou, Guangdong, China): miR-20amimics, miR-106b mimics and miR-125b mimics and control miRNA. Small interfering RNAs (siRNAs) against human HOTAIRM1 transcripts and the negative control RNA duplex (RiboBio) were purchased from Guangzhou RiboBio Co., Ltd. The nucleotide sequence for HOTAIRM1-siRNA1 and HOTAIRM1-siRNA2 were 5′-AGAAACUCCGUGUUACUCAUU-3′ (ref. 24) and 5′- GCCAGAAACCAGCCAUAGU-3′ (designed by RiboBio) respectively. The siRNA1 sequences targeting HOTAIRM1 were designed and expressed as shRNAs: sh-HOTAIRM1. Two siRNAs against human E2F1 were designed according to published data.43 Cells were transfected using the Neon Transfection System (Invitrogen) with 100 pmol of oligonucleotides in 10 μl reactions according to the manufacturer’s guidelines.
Dual luciferase activity assay
In the 3’-UTR assay, transient transfections were performed in HEK-293 T cells with 100 nM miR-20a, miR-106b or miR-125b mimics, or siRNA, or negative control vector; 0.1 μg of psiCHECK2 control or psiCHECK2-HOTAIRM1, psiCHECK2-ULK1, psiCHECK2-E2F1, or psiCHECK2-DRAM2; or psiCHECK-point mutated vector. For promoter analysis, 293 T cells at ~80% confluence were transfected with pGL3-Pro-LC3B (500 ng) and the internal control Renilla luciferase plasmid pRL-TK (10 ng, Promega) using Lipofectamine 3 000 (Invitrogen). To examine the effects of HOTAIRM1 on LC3B promoter activity, various combinations of pcDNA3.1 vectors expressing HOTAIRM1 variants (50 ng) or siRNAs targeting HOTAIRM1 (and E2F1) were co-transfected. Firefly and Renilla luciferase activities were measured consecutively 24 h following transfection using the Dual-Luciferase Reporter Assay (Promega) according to the manufacturer’s instructions.
Chromatin immunoprecipitation (ChIP)
ChIP was performed using the ChIP assay kit (Millipore, Billerica, MA, USA) according to the manufacturer’s instructions. DNA/protein crosslinking was achieved by incubating the cells for 20 min at 37 °C in 1% formaldehyde. After sonication, chromatin was immunoprecipitated overnight with 6 μg of anti-E2F1 antibody (Santa Cruz, Dallas, TX, USA) or 2 μg of normal IgG (Millipore). Genomic regions of LC3B and ULK1 containing the E2F1-binding sites were amplified by qRT-PCR using the primers as shown in Supplementary Table S1.
Western blot
Protein extracts were boiled in RIPA buffer (Beyotime, Shanghai, China) and separated in a SDS–polyacrylamide electrophoresis gel. The proteins were then transferred to a polyvinylidene fluoride membrane (Millipore) and probed with anti-RARA (Santa Cruz), anti-AGO2 (Abnova, Taipei, Taiwan), anti-p62 (Santa Cruz), anti-GABARAP (Proteintech, Chicago, IL, USA), anti-GFP (TransGen Biotech), anti-ULK1 (Novus Biologicals, Littleton, CO, USA), anti-LC3B (Novus Biologicals), anti-DRAM2 (Sigma-Aldrich), anti-E2F1 (Millipore) and anti-GAPDH (Sigma-Aldrich) antibodies.
Animal models
NB4 cells were transfected with Lv-shHOTAIRM1 (lentiviral vector expressing HOTAIRM1 shRNA) or Lv-NC (lentiviral vector expressing the negative control) according to the recommended protocol (GeneChem, Shanghai, China). The transfected cells (NB4-Lv-shRNA and NB4-Lv-NC) were sorted by FCM (MoFlo XDP instrument, Beckman Coulter, Brea, CA, USA) and GFP-positive cells were used for in vivo experiments. Five-week-old NOD-SCID mice were maintained under specific pathogen-free conditions in the Laboratory Animal Center of Sun Yat-sen University, and all experiments were performed according to our Institutional Animal Guidelines. The human APL-ascites SCID mouse model was used as previously described, with modifications.62 The tumor xenografts were established by a single intra peritoneal transplantation of 1 × 106 NB4 cells in 0.2 ml of RPMI-1640 medium into NOD-SCID mice.
Immunofluorescence and Transmission electron microscopy
The experiments are performed as our previous described procedures.39 EM images were obtained from thin sections using a JEM1400 electron microscope (JEOL, Akishima, Tokyo, Japan). The relative cytosolic areas in cross-sections of 10 cells were analyzed by Image-Pro Plus software. And fluorescence signals obtained using anti-LC3B antibody (Novus Biologicals), anti-E2F1 antibody (Santa Cruz), anti-ULK1 antibody (Novus Biologicals) and anti-DRAM2 antibody (Sigma-Aldrich) were acquired and analyzed by Carl Zeiss LSM 780 microscope (Carl Zeiss, Jena, Germany). At least 10 cells from each group were analyzed.
RNA immunoprecipitation and RNA pull-down
RNA immunoprecipitation was performed using a Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore) according to the manufacturer’s instructions. RNA for the in vitro experiments was transcribed using a TranscriptAid T7 High Yield Transcription Kit (ThermoFisher Scientific) according to the manufacturer’s instructions. The 3’-end biotin-labeled RNA production and RNA pull-down assay were carried out using a Pierce Magnetic RNA-Protein Pull-Down Kit according to manufacturer’s instructions (ThermoFisher Scientific). All of the RNA was purified by a Thermo GeneJET RNA Purification Kit (ThermoFisher Scientific).
Statistical analysis
Data are expressed as the mean±S.D. of three independent experiments. The differences between two groups were analyzed by Wilcoxon matched pairs test. The Kruskal–Wallis test was used for the comparison of three groups, and multiple comparisons were tested using the least significant difference test. Differences were considered significant at *P<0.05, **P<0.01 and ***P<0.001.
Abbreviations
- lncRNA:
-
long noncoding RNA
- AML:
-
acute myeloid leukemia
- APL:
-
acute promyelocytic leukemia
- ATRA:
-
all-trans retinoic acid
- CR:
-
complete remission
- qRT-PCR:
-
quantitative real-time PCR
References
Mercer TR, Dinger ME, Mattick JS . Long non-coding RNAs: insights into functions. Nat Rev Genet 2009; 10: 155–159.
Ponting CP, Oliver PL, Reik W . Evolution and functions of long noncoding RNAs. Cell 2009; 136: 629–641.
Mizutani R, Wakamatsu A, Tanaka N, Yoshida H, Tochigi N, Suzuki Y et al. Identification and characterization of novel genotoxic stress-inducible nuclear long noncoding RNAs in mammalian cells. PLoS One 2012; 7: e34949.
Wang KC, Chang HY . Molecular mechanisms of long noncoding RNAs. Mol Cell 2011; 43: 904–914.
Zhang H, Chen Z, Wang X, Huang Z, He Z, Chen Y . Long non-coding RNA: a new player in cancer. J Hematol Oncol 2013; 6: 37.
Dinger ME, Amaral PP, Mercer TR, Pang KC, Bruce SJ, Gardiner BB et al. Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res 2008; 18: 1433–1445.
Guttman M, Donaghey J, Carey BW, Garber M, Grenier JK, Munson G et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 2011; 477: 295–300.
Kallen AN, Zhou XB, Xu J, Qiao C, Ma J, Yan L et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell 2013; 52: 101–112.
Cesana M, Cacchiarelli D, Legnini I, Santini T, Sthandier O, Chinappi M et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 2011; 147: 358–369.
Ferdin J, Nishida N, Wu X, Nicoloso MS, Shah MY, Devlin C et al. HINCUTs in cancer: hypoxia-induced noncoding ultraconserved transcripts. Cell Death Differ 2013; 20: 1675–1687.
Galluzzi L, Pietrocola F, Bravo-San Pedro JM, Amaravadi RK, Baehrecke EH, Cecconi F et al. Autophagy in malignant transformation and cancer progression. EMBO J 2015; 34: 856–880.
Behrends C, Sowa ME, Gygi SP, Harper JW . Network organization of the human autophagy system. Nature 2010; 466: 68–76.
Klionsky DJ, Abdalla FC, Abeliovich H, Abraham RT, Acevedo-Arozena A, Adeli K et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 2012; 8: 445–544.
Lalaoui N, Johnstone R, Ekert PG . Autophagy and AML—food for thought. Cell Death Differ 2015; 23: 5–6.
Eriksen AB, Torgersen ML, Holm KL, Abrahamsen G, Spurkland A, Moskaug JO et al. Retinoic acid-induced IgG production in TLR-activated human primary B cells involves ULK1-mediated autophagy. Autophagy 2015; 11: 460–471.
Mizushima N, Levine B . Autophagy in mammalian development and differentiation. Nat Cell Biol 2010; 12: 823–830.
Maiuri MC, Kroemer G . Autophagy in stress and disease. Cell Death Differ 2015; 22: 365–366.
Saeki K, Yuo A, Okuma E, Yazaki Y, Susin SA, Kroemer G et al. Bcl-2 down-regulation causes autophagy in a caspase-independent manner in human leukemic HL60 cells. Cell Death Differ 2000; 7: 1263–1269.
Ge D, Han L, Huang S, Peng N, Wang P, Jiang Z et al. Identification of a novel MTOR activator and discovery of a competing endogenous RNA regulating autophagy in vascular endothelial cells. Autophagy 2014; 10: 957–971.
Wang K, Liu C, Zhou L, Wang J, Wang M, Zhao B et al. APF lncRNA regulates autophagy and myocardial infarction by targeting miR-188-3p. Nat Commun 2015; 6: 6779.
Couzin J . Cancer biology. A new cancer player takes the stage. Science 2005; 310: 766–767.
Sanchez-Mejias A, Tay Y . Competing endogenous RNA networks: tying the essential knots for cancer biology and therapeutics. J Hematol Oncol 2015; 8: 30.
Zeng CW, Zhang XJ, Lin KY, Ye H, Feng SY, Zhang H et al. Camptothecin induces apoptosis in cancer cells via microRNA-125b-mediated mitochondrial pathways. Mol Pharmacol 2012; 81: 578–586.
Zhang X, Lian Z, Padden C, Gerstein MB, Rozowsky J, Snyder M et al. A myelopoiesis-associated regulatory intergenic noncoding RNA transcript within the human HOXA cluster. Blood 2009; 113: 2526–2534.
Zhang X, Weissman SM, Newburger PE . Long intergenic non-coding RNA HOTAIRM1 regulates cell cycle progression during myeloid maturation in NB4 human promyelocytic leukemia cells. RNA Biol 2014; 11: 777–787.
Diaz-Beya M, Brunet S, Nomdedeu J, Pratcorona M, Cordeiro A, Gallardo D et al. The lincRNA HOTAIRM1, located in the HOXA genomic region, is expressed in acute myeloid leukemia, impacts prognosis in patients in the intermediate-risk cytogenetic category, and is associated with a distinctive microRNA signature. Oncotarget 2015; 6: 31613–31627.
de The H, Lavau C, Marchio A, Chomienne C, Degos L, Dejean A . The PML-RAR alpha fusion mRNA generated by the t(15;17) translocation in acute promyelocytic leukemia encodes a functionally altered RAR. Cell 1991; 66: 675–684.
Ablain J, de The H . Revisiting the differentiation paradigm in acute promyelocytic leukemia. Blood 2011; 117: 5795–5802.
Di Croce L, Raker VA, Corsaro M, Fazi F, Fanelli M, Faretta M et al. Methyltransferase recruitment and DNA hypermethylation of target promoters by an oncogenic transcription factor. Science 2002; 295: 1079–1082.
Villa R, Pasini D, Arantxa G, Morey L, Occhionorelli M, Viré E et al. Role of the polycomb repressive complex 2 in acute promyelocytic leukemia. Cancer Cell 2007; 11: 513–525.
Wang Z, Cao L, Kang R, Yang M, Liu L, Zhao Y et al. Autophagy regulates myeloid cell differentiation by p62/SQSTM1-mediated degradation of PML-RARα oncoprotein. Autophagy 2014; 7: 401–411.
Isakson P, Bjoras M, Boe SO, Simonsen A . Autophagy contributes to therapy-induced degradation of the PML/RARA oncoprotein. Blood 2010; 116: 2324–2331.
Lanotte M, Martin-Thouvenin V, Najman S, Balerini P, Valensi F, Berger R . NB4, a maturation inducible cell line with t(15;17) marker isolated from a human acute promyelocytic leukemia (M3). Blood 1991; 77: 1080–1086.
Zhang Z, Zhu Z, Watabe K, Zhang X, Bai C, Xu M et al. Negative regulation of lncRNA GAS5 by miR-21. Cell Death Differ 2013; 20: 1558–1568.
Paraskevopoulou MD, Georgakilas G, Kostoulas N, Reczko M, Maragkakis M, Dalamagas TM et al. DIANA-LncBase: experimentally verified and computationally predicted microRNA targets on long non-coding RNAs. Nucleic Acids Res 2012; 41: D239–D245.
Paraskevopoulou MD, Georgakilas G, Kostoulas N, Vlachos IS, Vergoulis T, Reczko M et al. DIANA-microT web server v5.0: service integration into miRNA functional analysis workflows. Nucleic Acids Res 2013; 41: W169–W173.
REHMSMEIER M . Fast and effective prediction of microRNA/target duplexes. RNA 2004; 10: 1507–1517.
Wu H, Wang F, Hu S, Yin C, Li X, Zhao S et al. MiR-20a and miR-106b negatively regulate autophagy induced by leucine deprivation via suppression of ULK1 expression in C2C12 myoblasts. Cell Signal 2012; 24: 2179–2186.
Zeng C, Chen Z, Zhang X, Han B, Lin K, Li X et al. MIR125B1 represses the degradation of the PML-RARA oncoprotein by an autophagy- lysosomal pathway in acute promyelocytic leukemia. Autophagy 2014; 10: 1726–1737.
Streeter A, Menzies FM, Rubinsztein DC . LC3-II tagging and western blotting for monitoring autophagic activity in mammalian cells. Methods Mol Biol 2016; 1303: 161–170.
Menzies FM, Moreau K, Puri C, Renna M, Rubinsztein DC . Measurement of autophagic activity in mammalian cells. Curr Protoc Cell Biol 2012; 15: 15–16.
Haim Y, Bluher M, Slutsky N, Goldstein N, Kloting N, Harman-Boehm I et al. Elevated autophagy gene expression in adipose tissue of obese humans: a potential non-cell-cycle-dependent function of E2F1. Autophagy 2015; 11: 2074–2088.
Polager S, Ofir M, Ginsberg D . E2F1 regulates autophagy and the transcription of autophagy genes. Oncogene 2008; 27: 4860–4864.
O'Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT . c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005; 435: 839–843.
Klattenhoff CA, Scheuermann JC, Surface LE, Bradley RK, Fields PA, Steinhauser ML et al. Braveheart, a long noncoding RNA required for cardiovascular lineage commitment. Cell 2013; 152: 570–583.
Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 2010; 464: 1071–1076.
Fang K, Han BW, Chen ZH, Lin KY, Zeng CW, Li XJ et al. A distinct set of long non-coding RNAs in childhood MLL-rearranged acute lymphoblastic leukemia: biology and epigenetic target. Hum Mol Genet 2014; 23: 3278–3288.
Garzon R, Volinia S, Papaioannou D, Nicolet D, Kohlschmidt J, Yan PS et al. Expression and prognostic impact of lncRNAs in acute myeloid leukemia. Proc Natl Acad Sci USA 2014; 111: 18679–18684.
Necsulea A, Soumillon M, Warnefors M, Liechti A, Daish T, Zeller U et al. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 2014; 505: 635–640.
Chu C, Spitale RC, Chang HY . Technologies to probe functions and mechanisms of long noncoding RNAs. Nat Struct Mol Biol 2015; 22: 29–35.
Johnsson P, Lipovich L, Grandér D, Morris KV . Evolutionary conservation of long non-coding RNAs; sequence, structure, function. Biochim Biophys Acta 2014; 1840: 1063–1071.
Tay Y, Rinn J, Pandolfi PP . The multilayered complexity of ceRNA crosstalk and competition. Nature 2014; 505: 344–352.
Grisolano JL, Wesselschmidt RL, Pelicci PG, Ley TJ . Altered myeloid development and acute leukemia in transgenic mice expressing PML-RAR alpha under control of cathepsin G regulatory sequences. Blood 1997; 89: 376–387.
Grignani F, Ferrucci PF, Testa U, Talamo G, Fagioli M, Alcalay M et al. The acute promyelocytic leukemia-specific PML-RAR alpha fusion protein inhibits differentiation and promotes survival of myeloid precursor cells. Cell 1993; 74: 423–431.
Wang ZY, Chen Z . Acute promyelocytic leukemia: from highly fatal to highly curable. Blood 2008; 111: 2505–2515.
Saeed S, Logie C, Stunnenberg HG, Martens JHA . Genome-wide functions of PML–RARa in acute promyelocytic leukaemia. Brit J Cancer 2011; 104: 554–558.
Manodoro F, Marzec J, Chaplin T, Miraki-Moud F, Moravcsik E, Jovanovic JV et al. Loss of imprinting at the 14q32 domain is associated with microRNA overexpression in acute promyelocytic leukemia. Blood 2014; 123: 2066–2074.
Saumet A, Vetter G, Bouttier M, Portales-Casamar E, Wasserman WW, Maurin T et al. Transcriptional repression of microRNA genes by PML-RARA increases expression of key cancer proteins in acute promyelocytic leukemia. Blood 2009; 113: 412–421.
Nie L, Wu HJ, Hsu JM, Chang SS, Labaff AM, Li CW et al. Long non-coding RNAs: versatile master regulators of gene expression and crucial players in cancer. Am J Transl Res 2012; 4: 127–150.
Prensner JR, Chinnaiyan AM . The emergence of lncRNAs in cancer biology. Cancer Discov 2011; 1: 391–407.
Grimwade D, Jovanovic JV, Hills RK, Nugent EA, Patel Y, Flora R et al. Prospective minimal residual disease monitoring to predict relapse of acute promyelocytic leukemia and to direct pre-emptive arsenic trioxide therapy. J Clin Oncol 2009; 27: 3650–3658.
Zhang SY, Zhu J, Chen GQ, Du XX, Lu LJ, Zhang Z et al. Establishment of a human acute promyelocytic leukemia-ascites model in SCID mice. Blood 1996; 87: 3404–3409.
Acknowledgements
We thank the autophage community provides the helpful guidelines and relevant reagents information. This research was supported by the National Natural Science Foundation of China (No. 81270629) and grants from Guangdong province (No. 2014T70833) and Guangzhou (201606080912429).
Author contributions
Z-HC designed and performed the research, analyzed the data and wrote the manuscript. W-TW, WH, KF, Y-MS and S-RL performed the research and analyzed the data. X-QL collected and analyzed the clinical data. Y-QC designed the research and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by D Rubinsztein
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Chen, ZH., Wang, WT., Huang, W. et al. The lncRNA HOTAIRM1 regulates the degradation of PML-RARA oncoprotein and myeloid cell differentiation by enhancing the autophagy pathway. Cell Death Differ 24, 212–224 (2017). https://doi.org/10.1038/cdd.2016.111
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2016.111
This article is cited by
-
Exploring the hidden hot world of long non-coding RNAs in thermophilic fungus using a robust computational pipeline
Scientific Reports (2024)
-
Exosome-encapsulated lncRNA HOTAIRM1 contributes to PM2.5-aggravated COPD airway remodeling by enhancing myofibroblast differentiation
Science China Life Sciences (2024)
-
Targeting HDAC3 to overcome the resistance to ATRA or arsenic in acute promyelocytic leukemia through ubiquitination and degradation of PML-RARα
Cell Death & Differentiation (2023)
-
Non-coding RNAs and Exosomal Non-coding RNAs in Traumatic Brain Injury: the Small Player with Big Actions
Molecular Neurobiology (2023)
-
LncRNA HOTAIRM1 promotes osteogenic differentiation of human bone marrow-derived mesenchymal stem cells by targeting miR-152-3p/ETS1 axis
Molecular Biology Reports (2023)