Abstract
Many vertebrate and insect viruses possess antiapoptotic genes that are required for their infectivity. This led to the hypothesis that apoptosis is an innate immunoresponse important for limiting virus infections. The role of apoptosis may be especially important in insect antiviral defense because of the lack of adaptive immunity. However, the cellular mechanism that elicits apoptosis in response to viral infection in insects has not been determined. Using an in vivo infection system with the mosquito baculovirus CuniNPV (Culex nigripalpus nucleopolyhedrovirus), we demonstrated that michelob_x (mx), the mosquito ortholog of Drosophila proapoptotic gene reaper, is specifically induced in larval midgut cells following viral infection. Interestingly, the dynamics of mx induction corresponds with the outcome of the infection. In the permissive mosquito C. quinquefasciatus, a slow induction of mx failed to induce prompt apoptosis, and the infected cells eventually undergo necrosis with heavy loads of encapsulated viruses. In contrast, in the refractory mosquito Aedes aegypti, a rapid induction of mx within 30 min p.i. is followed by apoptosis within 2–6 h p.i., suggesting a possible role for apoptosis in limiting viral infection. When the execution of apoptosis was delayed by caspase inhibitors, viral gene expression became detectable in the A. aegypti larvae.
Similar content being viewed by others
Main
The theory that apoptosis has a very important role in virus infections was mainly supported by evidence that many viruses possess one or multiple genes that interfere with cellular apoptosis during the infection process.1 Studies on viruses have uncovered a multitude of viral arsenals that can manipulate cellular apoptotic response in essentially all aspects and levels of the process. For instance, both the intrinsic and extrinsic cell death regulatory pathways are targeted by proteins and/or small RNAs encoded by mammalian viruses.2, 3 The extensiveness of viral interference of apoptosis ranges from manipulating upstream sensing and regulatory mechanisms to blocking the enzymatic activity of downstream effectors such as caspases.
Several very important cell death regulators were initially identified in viruses. For instance, IAP (inhibitor of apoptosis) was originally identified in lepidopteran baculoviruses.4, 5 Viruses mutated for iap induce rapid cell death in infected cells.6, 7 Viral IAPs not only can block cell death associated with viral infection but also apoptosis induced by other cytotoxic stimuli. Independently, genetic study in Drosophila melanogaster identified reaper-like genes as the pivotal regulators of programmed cell death.8 Subsequent genetic and biochemical analysis revealed that reaper-like proapoptotic genes function as IAP antagonists. One of the cellular iap genes, diap1, is ubiquitously expressed and required for the survival of cells. Essentially all cells in the developing embryo undergo apoptosis when functional Diap1 is absent.9, 10 Selective cell death during Drosophila development is mainly achieved by specific expression of the IAP antagonists reaper, hid, grim, and sickle. With the exception of HID, whose proapoptotic activity is subject to post-translational modification,11 IAP antagonists such as reaper are mainly regulated at the transcriptional level. In addition to mediating developmental cell death, IAP antagonists are also responsible for mediating cell death in response to environmental stimuli. For example, the expression of reaper in D. melanogaster can be activated/induced by X ray, UV irradiation, or hormonal surges.8, 12, 13
As insects lack adaptive immunity, it has been postulated that apoptosis would have an even more important role in antiviral response. Indeed, apoptosis has been observed during pathogen infection of mosquitoes and has been associated with host susceptibility to viral infection. It has been documented that ingestion of blood containing West Nile virus induces apoptosis in the midgut of a refractory Culex pipiens strain.14 In contrast, necrosis has been associated with Western Equine Encephalomyelitis virus infection in susceptible C. tarsalis strains.15 Although these evidences strongly suggest that proapoptotic response may have a very important role in determining vector compatibility, detailed mechanistic study has been hindered by the lack of knowledge about the underlying genetic mechanisms mediating proapoptotic response against viral infection.
The genome projects of Anopheles gambiae and Aedes aegypti revealed that, compared with D. melanogaster, these arbovirus vectors have expanded families of IAPs and caspases.16, 17 This was speculated as an adaptation to repeated exposure to pathogens associated with blood ingestion, although expansion of caspases has also been found in other Drosophila species.18 The A. gambiae genome project did not initially annotate any IAP antagonists because of the fast divergence of their sequences. The missing IAP antagonist was uncovered using an advanced bioinformatics approach, which identified michelob_x (mx) as the reaper-like IAP antagonist in both Anopheles and Aedes mosquitoes.19 Another IAP antagonist that is related to mx was subsequently characterized in A. aegypti.20 Despite the low sequence similarity of the entire gene, the functional domain, that is, the IAP-binding motif (IBM), was very well conserved, and the functional mechanism of Mx appears to be very similar to that of Reaper.
The identification of IAP antagonists in mosquitoes allowed us to ask whether reaper-like genes are involved in proapoptotic response to viral infection. We took advantage of the mosquito baculovirus CuniNPV (Culex nigripalpus nucleopolyhedrovirus) because of the accessibility of this system and the established insect pathology associated with CuniNPV infection.21 CuniNPV is originally isolated from the mosquito C. nigripalpus.22 It is related to lepidopteran and hymenopteran baculoviruses, but genomic sequence comparison indicated that there is a large evolutionary distance between CuniNPV and lepidopteran baculoviruses.23 CuniNPV infects only epithelial cells of the larval midgut, has a restricted host range, and mainly infects Culex spp. within the subgenus Culex.21 None of the examined Aedes mosquitoes, including A. aegypti, is susceptible to CuniNPV infection.21 CuniNPV can exist either as the occluded form or the budded form. The virus exists outside the mosquito in the occluded form, which allows the virus to survive under harsh environmental conditions. Ingested occluded virus initiates the infection in the presence of the divalent cation magnesium. Not all larval midgut cells are receptive to CuniNPV infection, which is limited to a particular group of resorbing/secreting cells in the gastric caeca and the posterior midgut.23 Once inside the midgut, the virus can spread from infected cells to neighboring cells via the budded form.
In this study, we showed that mx is induced in larval midgut cells following exposure to a mosquito baculovirus CuniNPV. More importantly, the dynamics of this induction is different in the susceptible C. quinquefasciatus versus the refractory A. aegypti.21 The relatively timid and delayed response of mx in C. quinquefasciatus (mx_Cu.qu) is associated with necrosis, whereas the robust and immediate induction of mx in A. aegypti (mx_Ae.ae) is followed by apoptosis within 6 h of viral exposure, suggesting that apoptosis may contribute to limiting viral infection in Aedes.
Results
Molecular cloning and characterization of mx in C. quinquefasciatus
Mx was originally identified in Anopheles and Aedes mosquito genomes as the ortholog of Drosophila Reaper using an integrated bioinformatics strategy and verified via functional assays.19 A similar bioinformatics approach was applied to identify potential IAP antagonists in the C. pipiens genome. Using the sequence information, we were able to clone the mx ortholog (mx_Cu.qu) gene from a cDNA made from C. quinquefasciatus larvae.
Mx_Cu.qu is ∼80% identical to its orthologs in A. aegypti (Mx_Ae.ae) or A. albopictus (Mx_Ae.al). The three mx orthologs in the Culicinae tribe share considerable similarity beyond the IAP-binding motif (Figure 1a). In contrast, they share little similarity with the mx ortholog in A .gambiae except the IBM. Given the evolution history of these groups, we would expect a significant difference between the subfamilies Anophelinae and Culicinae, which may also be partially reflected in the Mx protein distance tree (Figure 1b).
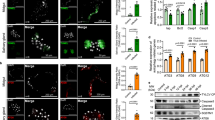
Gene cloning and characterization of mx_Cu.qu. (a) Alignment of protein sequences of Mx orthologs in mosquitoes and Reaper from D. melanogaster. The blue triangles denote the relative position of the intron in Mx sequences (there is no intron in Reaper). The three Mx orthologs from Culicinae share considerable similarity besides the IBM (red line). However, they share little similarity with Mx from Anopheles or the Drosophila Reaper. (b) Distance tree of these orthologs. (c) Expression of Mx_Cu.qu kills C6/36 cells. However, when amino acids 2–4 are removed (−IBM), the protein has little, if any, proapoptotic activity. (d) Mx_Cu.qu-induced cell death is almost completely blocked by co-transfection of Diap1 and significantly inhibited by co-transfection of p35. Numbers in parentheses denote the ratio of respective genes. (e) C6/36 cells killed by expression of mx_Cu.qu or mx_Ae.ae show typical apoptotic morphology including cell fragmentation (arrow) and phagocytosis by neighboring cells (arrow head). Cells co-transfected with pIE-lacZ and pIE-mx_Cu.qu were fixed 20 h later and processed for X-Gal staining
Like the mx orthologs in Aedes and Anopheles, Mx_Cu.qu induces rapid cell death when expressed in C3/36 cells (Figure 1c). This proapoptotic activity is largely, if not totally, dependent on the N-terminal IBM. Removing the first three amino acids of this motif (2–4; ‘AIA’) abolishes most of the killing ability of Mx_Cu.qu (Figure 1c).
The proapoptotic activity of Mx_Cu.qu and Mx_Ae.ae appear to be very similar when assayed in C6/36 cells. Cell killing induced by either Mx_Cu.qu or Mx_Ae.ae is significantly suppressed by co-transfection of Diap1, and to a less degree, by co-transfection of the viral inhibitor P35 (Figure 1d). Diap1 is the major antiapoptotic cellular protein in Drosophila and is highly conserved from insects to mammals. P35 is the viral caspase inhibitor that was originally identified from lepidopteran baculovirus.24 The fact that Mx_Cu.qu-induced cell death can be blocked by either Diap1 or P35 indicates that the functional mechanism of Mx_Cu.qu is the same as previously characterized for Mx orthologs in Anopheles and Aedes and Reaper in Drosophila.
Cell death induced by expressing mx_Cu.qu or mx_Ae.ae in C6/36 has typical apoptotic hallmarks, that is, nuclear condensation, fragmentation, and so on. Interestingly, C6/36 cells killed by expression of mx_Cu.qu or mx_Ae.ae appear to be quickly phagocytosed by neighboring cells (Figure 1e). Since pIE-lacZ was co-transfected with pIE-mx_Cu.qu, cells expressing mx_Cu.qu or mx_Ae.ae are also β-galactosidase (β-gal) positive. At 20 h after transfection, most of the remaining blue β-gal-positive cells in mx_Cu.qu-transfected samples appear to be in the later stages of apoptosis. These β-gal-positive cells are often rounded up and/or fragmented. Small phagosomes with β-gal staining can be clearly seen in cells surrounding the fragmented β-gal-positive cell, indicating that dying or dead cells were quickly phagocytosed before the breakdown of the lacZ protein. A similar phenomenon was observed for reaper and hid-induced cell death in Drosophila embryos where most dying cells were quickly phagocytosed by hemocyte/macrophages.8, 25, 26 However, not all Drosophila cell lines have endocytotic capability.27 The fact that C6/36 cells are actively attracted to apoptotic cells and aggressively phagocytose fragmented apoptotic bodies suggests that this cell line, similar to the Drosophila S2 cell line, may have a mesoderm origin and display hemocyte/macrophage functionality.
The mx gene is induced in midgut cells infected by CuniNPV
Using quantitative real-time PCR (Q-PCR) we first monitored the expression of mx following CuniNPV infection in pools of whole larvae. We found that in C. quinquefasciatus there was a lack of significant increase of mx_Cu.qu expression during the initial phase of infection, that is, before 8 h p.i. However, 48 h after the initiation of virus infection, the level of mx_Cu.qu reached significantly higher levels (Figure 2a). At this time, the level of mx_Cu.qu in infected larvae was about six times higher than the uninfected controls.
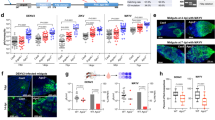
Induction of mx_Cu.qu and mx_Ae.ae following CuniNPV infection. (a) The level of mx_Cu.qu and mx_Ae.ae in pooled larvae (15–20 for each time points) following CuniNPV infection of C. quinquefasciatus and A. aegypti, respectively. Expression level was first normalized with house-keeping genes before calculating the fold induction. Data are presented as mean±S.D. of at least three independent experiments. (*P-value<0.05 and ***P-value<0.001.) (b) Expression of mx_Cu.qu in the midguts of CuniNPV-infected or control (Ctrl) C. quinquefasciatus larvae was monitored with FISH. While there is little induction of mx_Cu.qu at early infection (2 h p.i.), there is very high level of mx_Cu.qu in infected cells at 24 and 48 h p.i. (c) Expression of mx_Ae.ae in the midguts of Ctrl or CuniNPV-infected A .aegypti larvae was monitored with FISH. There is a rapid induction of mx_Ae.ae in cells at the gastric caeca at 2 h p.i. The number of mx_Ae.ae-positive cells returned to the background level by 8 h p.i. For each time point, at least 50 midguts were analyzed in two to three independent experiments, and 80–100% examined guts presented the features as represented in b and c
In sharp contrast, Q-PCR analysis indicated that there was a quick response of mx_Ae.ae induction in the first 2 h following exposure to CuniNPV in the refractory A. aegypti larvae (Figure 2a). Significant induction of mx_Ae.ae was found within 1 h of exposure to the virus (Supplementary Figure S1). The elevated level of mx_Ae.ae in virus-challenged larvae was transient and retreated to background or previous infection level at about 8 h p.i.
The dynamics of mx induction in both C. quinquifasciatus and A. aegypti were verified via fluorescent in situ hybridization (FISH) with cRNA probes to mx_Cu.qu and mx_Ae.ae, respectively. Consistent with the Q-PCR data, only timid induction of mx_Cu.qu could be detected in a few cells in the gastric caeca at early stage of virus infection (2 h) in CuniNPV-infected C. quinquifasciatus larvae (Figure 2b). However, the mRNA level of mx_Cu.qu is very high at later stages of infection. At 48 h p.i., it appears that most cells in the gastric caeca and the posterior midgut have very high levels of mx_Cu.qu (Figure 2b and Supplementary Figure S2). These cells also display necrotic nuclei morphology (described below).
In contrast, there is a rapid induction of mx_Ae.ae in the gastric caeca of A. aegypti larvae exposed to the same treatment of CuniNPV. The number of cells expressing mx_Ae.ae, as detected by FISH, reached a peak at about 2 h p.i. At 4 h p.i., the number of mx_Ae.ae-positive cells had decreased significantly. By 8 h p.i., there was no difference in mx_Ae.ae FISH signal between virus-treated and control A. aegypti larvae (Figure 2c).
Necrosis, not apoptosis, is associated with CuniNPV infection in the susceptible mosquito C. quinquefasciatus
Previous histological analysis has shown that many cells at the gastric caeca and posterior midgut of infected C. nigripalpus or C. quinquefasciatus larvae were heavily infected with CuniNPV occlusion bodies (OBs) and display signs of necrosis at the final stages of infection.23, 28 When TUNEL (terminal deoxynucleotidyl transferase dUTP nick end labeling) was applied to monitor possible apoptotic response to viral infection in C. quinquefasciatus larvae, there was no significant level of apoptosis at either early stage (2–8 h p.i.) or late stages (24 and 48 h p.i.) of the infection (data not shown).
FISH analysis with cRNA probes for CuniNPV genes allowed us to pinpoint virus-infected cells even before the final stages of infection. Counter stain with DAPI indicated that the nuclei of infected cells, that is, cells with CuniNPV gene expression, were greatly hypertrophied at 48 h p.i. (Figure 3a). These nuclei were swollen to 2–4 times the diameter compared with corresponding cells in uninfected larvae. Our FISH analysis confirmed that before 8 h p.i., only a small group of cells at the gastric caeca and posterior midgut were infected (i.e., have viral gene expression). However, by 48 h p.i., most cells at the posterior midgut and gastric caeca were positive for viral gene expression and had swollen nuclei.23 This indicates that the virus initially infects only a small group of cells but later spreads horizontally to infect other cells, presumably via budded virus. Interestingly, while most of the cells in the posterior midgut were positive for viral gene expression, none of the cells at the anterior midgut was positive for viral gene expression.
Necrosis in CuniNPV-infected C. quinquefasciatus larvae. (a) Cells infected with CuniNPV at 48 h p.i. were identified with a pool of fluorescein-labeled cRNA probes against CuniNPV genes, cun24, cun75, and cun85. Virus-infected cells at this time demonstrated hypertrophied nuclei (arrow) compared with uninfected cells (arrow head). Note that there is clear separation between cells that were positive for viral gene expression versus those that were negative (dashed line). More than 100 midguts were examined in several independent experiments; all cells positive for viral gene expression at this time point have hypertrophied nuclei (additional images in Supplementary Figure S2). (b) Cells with hypertrophied nuclei have compromised cell membrane integrity. Live larvae at 48 h p.i. were exposed to 1 μg/ml PI in culture media for 10 min before the midguts were dissected out, fixed with paraformaldehyde, and counterstained using DAPI. All the cells with hypertrophied nuclei (arrow) are also permeable to PI, indicating necrotic cell death. In contrast, cells in the anterior midgut (arrow head) have normal nuclei and intact membrane. More than 40 midguts were observed in three independent experiments, and essentially all of them showed the described association between nuclei morphology and permeability to PI
To confirm that virus-infected cells in C. quinquefasciatus larvae undergo necrotic instead of apoptotic cell death, we examine the membrane permeability of these infected cells. When live infected C. quinquefasciatus larvae were incubated for 10 min in normal culture media containing 1 μg/ml propidium iodide (PI), all of these swollen cells at posterior midgut were labeled with PI, whereas none of the uninfected cells in the anterior midgut were labeled (Figure 3b). The swollen morphology and the compromised membrane integrity of these cells are characteristics of necrosis. Indeed, tissues of the infected specimens at 48 h p.i. were very fragile and easily disintegrated when touched or pulled during dissection. Eventually, at about 60–72 h p.i., most mosquitoes died and tissues disintegrated into the culture media.
Rapid induction of mx_Ae.ae is followed by apoptosis in the refractory mosquito A. aegypti
The dynamics of induction of mx was very different between the susceptible C. quinquefasciatus larvae and the refractory A. aegypti larvae. Cells expressing high level of mx_Ae.ae could be detected within 1 h of infection in the gastric caeca and reach a peak at 2 h p.i. TUNEL analysis indicated that this rapid induction of mx_Ae.ae was followed by a wave of apoptosis in the midgut epithelial cells. TUNEL-positive cells could be observed in the gastric caeca at 2 h p.i. (Figure 4). This wave of apoptosis in CuniNPV-infected A. aegypti larvae appears to be relatively synchronized. The number of TUNEL-positive cells is the highest at about 2–4 h p.i. (Figure 4a). By 8 h p.i., few cells in the gastric caeca or posterior midgut can be found to be TUNEL positive.
Rapid apoptosis following the mx_Ae.ae induction in A. aegypti larvae exposed to CuniNPV. (a) TUNEL assay in A. aegypti midgut. At 2–4 h after CuniNPV infection (2–4 h p.i.), significant increase of TUNEL-positive cells can be detected in over 80% of the examined midguts. In contrast, few TUNEL-positive cells were detectable in the control animal (Ctrl) or the midgut of infected animal at late time points (24 h p.i.). (b) Colocalization analysis indicated that TUNEL-positive cells are the ones that express mx_Ae.ae. Early-stage TUNEL-positive cells (open arrow), indicated by relatively normal nuclei morphology and intact cytoplasm, have high levels of FISH signal for mx_Ae.ae. The mx_Ae.ae signal is not visible in later-stage TUNEL-positive cells with significantly condensed nuclei (solid arrow), which is likely because of the shrinkage of cytoplasm and/or degradation of macromolecules at this stage. Some mx-expressing cells are not TUNEL positive, which is likely because of the lag between mx expression and the activation of caspase-dependent DNase
The quick apoptotic process following mx_Ae.ae induction helps to explain the sharp decline of mx_Ae.ae–positive cells after 2 h p.i. Colocalization analysis confirmed that TUNEL-positive cells, especially those at early apoptotic stages before the onset of significant nuclei condensation, are also positive for mx_Ae.ae (Figure 4b). The timeline of the events indicates that cells undergo apoptosis shortly after the expression of mx_Ae.ae. This is not surprising given that in Drosophila embryogenesis, the expression of reaper in cells destined to die is followed immediately by apoptosis within approximately 1–2 h.8
Inhibition of apoptosis leads to expression of viral genes in the refractory mosquito A. aegypti
The rapid induction of mx and apoptosis following CuniNPV infection in the larval midgut suggests that cells invaded by the virus underwent apoptosis and were quickly eliminated from the midgut epithelium. Under normal conditions, we were not able to detect viral gene expression in A. aegypti larvae exposed to CuniNPV at any time point by either FISH or Q-PCR. This could potentially be attributed to at least three possibilities. First, maybe the virus could not enter the cell because of incompatibility of cell surface receptors. Second, virus that entered the cell could not replicate because of incompatibility of the cellular transcription system. Third, it could be that the quick apoptotic response prevented viral gene expression from reaching a detectable level. To discern these possibilities, we tested whether blocking or delaying apoptosis following CuniNPV infection in A. aegypti larvae could have an impact on the infection process.
The refractory A. aegypti larvae were infected with CuniNPV and split into two populations after the addition of virus and MgCl2. To one series of samples, pan caspase inhibitors z-VAD-FMK, or Q-VD-OPH, or a combination of both was added to the medium. As a control, the same amount of DMSO (the solvent for both z-VAD and Q-VD) was added to the parallel samples. This treatment of both z-VAD and/or Q-VD was able to suppress the cell death induced by transfection of mx in a tissue culture-based assay (Supplementary Figure S3).
From both the caspase inhibitor-treated and control populations, larvae were collected at the indicated time points and processed for Q-PCR and FISH analyses. Immediate-early (ie) gene cun103 is one of the first genes expressed following CuniNPV infection of C. quinquefasciatus. Although it is not detected in A. aegypti larvae treated with DMSO, it was reliably detectable in caspase inhibitor-treated samples (Figure 5a). The expression of cun103 was detectable via Q-PCR at 8, 24, and up to 30 h p.i. In most of the independent trials, the expression of cun103 diminished after 48 h p.i. Only in one out of five trials, we detected persistent cun103 expression in a 72 h p.i. sample. Correspondingly, structural (nucleocapsid) genes cun024 and cun035 were detectable in that sample, indicating that at least in one larva, late-stage viral expression was achieved by inhibiting/delaying apoptosis (Supplementary Figure S4).
Suppressing apoptosis in A. aegypti leads to expressions of viral genes. (a) Suppressing apoptosis allowed the expression of ie viral genes. None of the three ie CuniNPV genes, cun16, cun86, and cun103, could be detectable by Q-PCR in A. aegypti larvae exposed to CuniNPV and treated with DMSO only. In contrast, all the three genes could be detected in A. aegypti larvae treated with capsase inhibitors z-VAD-FMK (z-VAD). Bottom panels are gel pictures of the Q-PCR products. (b) FISH was performed with a pool of cRNA probes against CuniNPV genes, cun16, cun86, and cun103. Top panel, no viral gene could be detected in the midgut of any of the DMSO-treated larvae (over 100 examined) at 24 h after CuniNPV infection. In contrast, viral gene expression was detectable in ∼20% of the midguts dissected from caspase inhibitor-treated larvae
FISH analysis confirmed that about 10–20% of larvae treated with caspase inhibitors have cells with detectable viral gene expression at 24 h p.i. (Figure 5b). The viral gene signals were confined to the nuclei, similar to what we observed with early-stage infection of CuniNPV in the permissive C. quinquefasciatus (Supplementary Figure S5). However, repeated trials with different caspase inhibitors or combinations of caspase inhibitors indicated that the efficacy of caspase inhibitor in transforming the resistant A. aegypti larvae was limited. We were never able to improve the infection rate to more than 20%, nor could we reliably detect the expression of virus structural genes in every trial.
Discussion
A significant contribution to our knowledge about the role of apoptosis in virus–host interaction came from studying lepidopteran baculoviruses such as AcMNPV (Autographa californica multicapsid nucleopolyhedrovirus).29 Several antiapoptotic genes have been characterized in lepidopteran baculoviruses, including iap, p35, and p49.4, 30 While injecting wild-type virus to moth larvae usually leads to necrosis, injecting viruses with mutated p35 induced massive apoptosis.31 Consequently, the infectivity of the mutant virus was much lower than that of the wild-type virus. Most of these previous studies relied on directly injecting budded viruses into the hemocoel. However, in natural conditions, baculovirus infection of insects is initiated in the midgut epithelia by occluded viruses. Ours is the first evidence showing that infection by the occluded baculovirus induces proapoptotic response. In addition, the correlation between the cellular outcome of CuniNPV-infected midgut epithelial cells and the organism susceptibility to viral infection strongly suggests that apoptosis has a very important role in limiting viral infection.
Apoptosis in the mosquito midgut epithelia has long been observed accompanying the infection of a variety of pathogens.14, 32 However, we know little as to which regulatory pathway is responsible for mediating apoptosis against pathogen infection.33 Our study here indicated that the mosquito ortholog of reaper, mx, is activated following CuniNPV infection. This, to our knowledge, is the first evidence suggesting that reaper-like genes, known for their pivotal role in regulating programmed cell death during development, are also involved in the innate immune response against viral infection.
In Drosophila, reaper-like genes (hid, grim, and sickle) are required for most developmental cell death.34 The reaper, grim, and sickle are exclusively expressed in cells destined to die during development or following cytotoxic stimuli such as irradiation. Transcriptional regulation of mx also appears to be tightly regulated in the mosquito larval midgut. In the absence of viral infection, only a few cells have detectable levels of mx expression and presumably reflects the routine turnover of midgut epithelial cells. The induction of mx in CuniNPV-infected C. quinquefasciatus larval midgut is limited to virus-infected tissues, that is, gastric caeca and posterior midgut. As we have reported previously, like reaper, mx is also transcriptionally responsive to UV irradiation.35 Thus it appears that although the protein sequence of Mx has diverged significantly from that of Reaper, their regulation at the transcriptional level share much similarity.
Our study strongly indicates that the induction of mx is very likely responsible for the proapoptotic response following CuniNPV infection. However, direct assessment of this hypothesis was hindered by the lack of technical means to specifically block the surge of mx_Ae.ae expression following viral infection. There has not been any reported success with RNAi in Aedes larvae. Our attempts with double-stranded RNA, using a strategy that worked in A. gambiae larvae,36 failed to produce any damping effect on the induction of mx_Ae.ae following CuniNPV infection. Besides potential problems associated with the efficacy of RNAi in Aedes larvae, it is also questionable whether the RNAi mechanism, which can bring down the level of constitutively expressed genes, can be effective in suppressing the quick induction of mx_Ae.ae associated with viral infection. It is possible that because the induction is so robust and followed quickly by apoptosis, RNAi mechanism could not function fast enough to have a significant, if any, impact on the rapid increase of mx_Ae.ae. It might be necessary to first identify the enhancer(s) mediating viral infection-induced mx_Ae.ae and applying this to genetically engineered mosquito to truly test the hypothesis that the induction of mx_Ae.ae is responsible for the apoptosis of midgut cells following CuniNPV infection.
A race to apoptosis?
Our study highlights the importance of the timing of proapoptotic response on the cellular and organismal outcome. In the refractory host A. aegypti, the induction of mx_Ae.ae and apoptosis happened so fast that viral gene expression was never detectable in the process. In contrast, in the susceptible host C. quinquefasciatus, the delayed and timid induction of mx_Cu.qu did not reach to detectable level until viral genes were detectable (Figure 6). The data presented here do not exclusively suggest that the quick induction of apoptosis is responsible for the refractory phenotype, as there are many other differences between the two species. However, the fact that viral gene can be detected in A. aegypti when apoptosis is delayed/suppressed suggests that apoptosis is responsible for preventing viral gene expression.
The race to apoptosis through the mosquito reaper. This is a schematic presentation of our findings. On CuniNPV infection, there is a competing ‘race’ between the host cell to express cellular proapoptotic gene(s) to eliminate the infected cell and the virus to express early genes to block apoptosis and initiate proliferation. A prompt induction of mx and apoptosis leads to elimination of the infected cell before the viral genes are detectable in the refractory mosquito A. aegypti. A delay in this process, either due to insufficient proapoptotic response or interference of caspase inhibitors, could subjugate the cell under viral control and render the organism susceptible to the virus
The extremely high level of mx_Cu.qu in CuniNPV-infected midgut cells in C. quinquefasciatus indicates that the virus has a very efficient way to block mx-induced apoptosis. In the tissue culture system, mx-induced cell death can be effectively blocked by co-expression of iap. However, there was no clear ortholog of either iap or p49 in CuniNPV. A predicted viral protein, CUN75, has marginal similarity to AcMNPV P35, but lacks a conserved reactive site loop that is required for caspase-inhibition activity.37 We did not find any antiapoptotic effect of cun75 when co-transfected with mx_Cu.qu or mx_Ae.ae. As the only dipteran baculovirus with genomic information, CuniNPV is distantly related to lepidopteran baculoviruses and lacks identifiable orthologs to many important genes such as the ie genes.38 It is very likely that CuniNPV utilizes a yet unknown, but powerful, mechanism to block the apoptotic pathway downstream of mx activation.
Apoptosis and mosquito–host compatibility
Correlation between apoptosis and resistant phenotype has been found with refractory Culex mosquito exposed to the West Nile virus.14 Our comparison of the CuniNPV infection in the A. aegypti and C. quinquefasciatus also suggests that apoptosis has an important role in eliminating CuniNPV-infected midgut cells and convey resistance to viral infection.
Partially blocking apoptosis by feeding the refractory A. aegypti larvae with caspase inhibitors allowed reliable detection of several ie viral genes. This indicates that the virus was able to enter into the nuclei of the midgut cells and successfully initiate gene expression when apoptosis was blocked or delayed. The fact that the application of caspase inhibitor alone was not reliable in allowing expression of structural genes may be because of several possibilities. First, there may be additional antiviral responses that can block the developmental program of the virus. However, it is also possible that while caspase inhibitor partially blocks the activity of Mx-activated caspases, it did not block the other antivirus mechanisms elicited by Mx. It has been found that Reaper can nonspecifically inhibit protein synthesis,39, 40 an activity that is independent of caspase activation. It remains to be seen as to whether Mx also has this activity and can inhibit viral protein synthesis. It is notable that in the tissue culture system, while co-expressing iap was very effective in blocking mx-induced cell death (Figure 1), application of very high level of caspase inhibitor only partially blocked/delayed cell death (Supplementary Figure S3).
In summary, our study found that mx, the mosquito ortholog of the proapoptotic gene reaper, is transcriptionally activated in cells infected by CuniNPV and that the cellular outcome correlates with organism resistance/susceptibility. This should pave the way for more mechanistic studies addressing the role of apoptosis in determining host susceptibility or vector compatibility for human pathogens. For instance, as controlled expression of mx_Ae.ae has been successfully used as an effector for sterile insect approaches,41 it will be very interesting to see whether adding viral responsiveness to such a system could also make the vector more resistant to viral infection.
Materials and Methods
Bioinformatics, gene cloning, and cell death assay
Data-mining strategy, plasmid construction, and in vitro cell death assay were performed as described previously.19 Briefly, a motif search program implemented in C was customized to search for the IBM motifs in the genomic and EST sequences from mosquito genomes. An intronless cDNA for mx_Cu.qu was then obtained by reverse transcriptase PCR using RNA extracted from CuniNPV-infected mosquito C. quinquefasciatus that serves as the template for PCR reaction. The primers used here are 5′-ACCGGCCGGCGCTGGTTACGTGATTCT-3′ and 3′-CGGGATCCATACACTTTGCGGAGCAGA-5′. PCR product was then subcloned into the pIE-3 vector for transfection assay, and cloned into pBS for synthesizing cRNA probe. The other plasmids used in this study, including Pie-LacZ, Pie-mx_Ae.ae, Pie-p35, and Pie-diap1, are described previously.19
Mosquito rearing and CuniNPV infection
C. quinquefasciatus (Gainesville, FL strain, maintained since 1995) and A. aegypti (Orlando, FL strain, maintained since 1952) were reared in the insectary of the Mosquito and Fly Research Unit at the Center for Medical, Agricultural, and Veterinary Entomology, USDA-ARS, Gainesville, FL, USA. CuniNPV viral OBs were purified from infected C. quinquefasciatus larvae as described previously.23 The standard assay involves groups of 100 three- to four-day-old C. quinquefasciatus and A. aegypti larvae obtained from the laboratory colony. Larvae are exposed at a dose of 1.2 × 109 OBs per ml in 100 ml of deionized water with 15 mM MgCl2 plus 50 mg alfalfa and potbelly pig chow mixture (2 : 1). Groups without the addition of the virus serve as controls.
RNA extraction and Q-PCR
Total RNA was extracted from frozen larvae with RNeasy Mini Kit (QIAGEN, Valencia, CA, USA) according to the protocol provided by the manufacturer. RNA samples were treated with DNase I to remove genomic DNA. cDNA was prepared by reverse transcription of total RNA with a High-Capacity cDNA Archive Kit (Applied Biosystems, Foster City, CA, USA). Q-PCR was performed with an ABI 7500 Fast thermocycler (Applied Biosystems) following protocols provided by the manufacturer. Triplicates were measured for each gene/sample combination.
To control for variation of mx expression levels during normal development, each pool of larvae was hatched on the same day and randomly separated into control and viral treatment groups. A sample of 10–12 larvae was taken from each group at a given time point and frozen at −80 °C. The cellular expression levels of mx were measured as relative to the glyceraldehyde 3-phosphate dehydrogenase or actin levels before the calculation of the ratio between CuniNPV-treated and control samples.
PI staining
Live mosquito larvae were removed from the culture media, washed briefly with water, and transferred to 6-well plates with 5 ml of fresh culture media containing 1 μg/ml PI (Sigma, St. Louis, MO, USA). After 10 min of incubation, the staining was stopped by transferring the larvae to fresh media without PI. The larvae were than dissected quickly, and the midgut and attached tissues were fixed in 4% paraformaldehyde in PBS. The fixed samples were washed three times in PBS. Samples were mounted with the Vectorshield mounting medium (Vector Lab, Burlingame, CA, USA) containing DAPI. The pictures were taken with a Leica upright fluorescent microscope (Leica, Bannockburn, IL, USA) using OpenLab software (Improvision, Coventry, UK).
TUNEL assay
Apoptosis was detected using the FragEL Kit (Calbiochem, Gibbstown, NJ, USA) with a modified protocol. Midguts were dissected and fixed with 4% paraformaldehyde, 100 mM PIPES buffer, pH 7.4, 2 mM MgSO4, and 1 mM EGTA for 20 min at room temperature. They were washed with TBS (20 mM Tris-HCl, pH 8.0, and 140 mM NaCl), followed by 5 min incubations in a TBS-methanol step gradient from 0, 25, 50, 75 to 100% methanol and back to TBS buffer following a reversed methanol series of concentrations. The tissue was then incubated for 5 min at room temperature with 20 μg/ml protease K in TBS, and the reaction was stopped by washing with 4% paraformaldehyde in TBS. This was followed by another 10 min fixation with 4% paraformaldehyde in TBS. The sample was equilibrated for 20 min with equilibration buffer at room temperature and incubated with TdT labeling reaction mixture for 1 h at 37 °C. The samples were then mounted with Vectorshield mounting medium (Vector Lab) containing DAPI. The pictures were taken with a Leica upright fluorescent microscope using OpenLab software.
FISH
Probes were synthesized using DIG- or Fluorescein-RNA Labeling Mix (Roche, Madison, WI, USA). Midguts were dissected, fixed, and processed as described above. After prefixing with 4% paraformaldehyde in PBT_DEPC (0.3% Triton in PBS made with DEPC pretreated double-distilled water) for 30 min, the tissue was incubated for 7 min with 50 μg/ml protease K in PBT_DEPC, and reaction was stopped by washing with 4% paraformaldehyde. Samples were incubated with probes diluted in hybridization buffer (50% formamide, 25% 2 × SSC, 20 μg/ml yeast tRNA, 100 μg/ml ssRNA, 50 μg/ml heparin, and 0.1% Tween-20). Hybridization was performed overnight at 60 °C. If necessary, HRP-conjugated anti-DIG or anti-FITC (Roche) antibody (depends on what marker the probes carry) was applied after hybridization, followed by signal amplification using the Tyramid Signal Amplification Kit (PerkinElmer, Waltham, MA, USA).
Blocking apoptosis with z-VAD and Q-VD-OPH
Immediately following the administration of virus and MgCl2, exposed A. aegypti larvae were removed from the culture pan together with the virus containing media to 24-well or 12-well culture dishes. z-VAD-fmk (R&D Bioscience, Minneapolis, MN, USA) and Q-VD-OPH (BioVision, Mountain View, CA, USA) were added to individual set of the wells for final concentration of 100 and 50 μM respectively. The combination of 50 μM z-VAD-fmk and 20 μM Q-VD-OPH was also applied. Meanwhile, equal amounts of DMSO (solvent for z-VAD and Q-VD-OPH) were added to a parallel set of wells as control. Larvae were collected at discrete time points following the infection and processed for Q-PCR or FISH as described above. To assess the inhibitory efficiency of above caspase inhibitors, cultured C6/36 cells were transfected with 0.2 μg lacZ and 0.2 μg Mx in the presence or absence of corresponding caspase inhibitors. The number of viable (lacZ positive) cells were counted at 24 h after transfection.
Abbreviations
- mx :
-
michelob_x
- IAP:
-
inhibitor of apoptosis
- β-gal:
-
β-galactosidase
- IBM:
-
IAP-binding motif
- CuniNPV :
-
Culex nigripalpus nucleopolyhedrovirus
- AcMNPV :
-
Autographa californica multicapsid nucleopolyhedrovirus
- OB:
-
occlusion body
- Q-PCR:
-
quantitative real-time PCR
- FISH:
-
fluorescent in situ hybridization
- TUNEL:
-
terminal deoxynucleotidyl transferase dUTP nick end labeling
- PI:
-
propidium iodide
References
Benedict CA, Norris PS, Ware CF . To kill or be killed: viral evasion of apoptosis. Nat Immunol 2002; 3: 1013–1018.
Galluzzi L, Brenner C, Morselli E, Touat Z, Kroemer G . Viral control of mitochondrial apoptosis. PLoS Pathog 2008; 4: e1000018.
Thomson BJ . Viruses and apoptosis. Int J Exp Pathol 2001; 82: 65–76.
Clem RJ, Miller LK . Control of programmed cell death by the baculovirus genes p35 and iap. Mol Cell Biol 1994; 14: 5212–5222.
Crook NE, Clem RJ, Miller LK . An apoptosis-inhibiting baculovirus gene with a zinc finger-like motif. J Virol 1993; 67: 2168–2174.
Li QJ, Liston P, Moyer RW . Functional analysis of the inhibitor of apoptosis (iap) gene carried by the entomopoxvirus of Amsacta moorei. J Virol 2005; 79: 2335–2345.
Means JC, Muro I, Clem RJ . Silencing of the baculovirus Op-iap3 gene by RNA interference reveals that it is required for prevention of a apoptosis during Orgyia pseudotsugata M nucleopolyhedrovirus infection of Ld652Y cells. J Virol 2003; 77: 4481–4488.
White K, Grether ME, Abrams JM, Young L, Farrell K, Steller H . Genetic control of programmed cell death in Drosophila. Science 1994; 264: 677–683.
Goyal L, McCall K, Agapite J, Hartwieg E, Steller H . Induction of apoptosis by Drosophila reaper, hid and grim through inhibition of IAP function. EMBO J 2000; 19: 589–597.
Hawkins CJ, Wang SL, Hay BA . A cloning method to identify caspases and their regulators in yeast: identification of Drosophila IAP1 as an inhibitor of the Drosophila caspase DCP-1. Proc Natl Acad Sci USA 1999; 96: 2885–2890.
Bergmann A, Agapite J, McCall K, Steller H . The Drosophila gene hid is a direct molecular target of Ras-dependent survival signaling. Cell 1998; 95: 331–341.
Jiang C, Lamblin AF, Steller H, Thummel CS . A steroid-triggered transcriptional hierarchy controls salivary gland cell death during Drosophila metamorphosis. Mol Cell 2000; 5: 445–455.
Zhou L, Steller H . Distinct pathways mediate UV-induced apoptosis in Drosophila embryos. Dev Cell 2003; 4: 599–605.
Vaidyanathan R, Scott TW . Apoptosis in mosquito midgut epithelia associated with West Nile virus infection. Apoptosis 2006; 11: 1643–1651.
Weaver SC, Lorenz LH, Scott TW . Pathologic changes in the midgut of Culex tarsalis following infection with Western equine encephalomyelitis virus. Am J Trop Med Hyg 1992; 47: 691–701.
Christophides GK, Zdobnov E, Barillas-Mury C, Birney E, Blandin S, Blass C et al. Immunity-related genes and gene families in Anopheles gambiae. Science 2002; 298: 159–165.
Nene V, Wortman JR, Lawson D, Haas B, Kodira C, Tu ZJ et al. Genome sequence of Aedes aegypti, a major arbovirus vector. Science 2007; 316: 1718–1723.
Bryant B, Ungerer MC, Liu QZ, Waterhouse RM, Clem RJ . A caspase-like decoy molecule enhances the activity of a paralogous caspase in the yellow fever mosquito, Aedes aegypti. Insect Biochem Molec 2010; 40: 516–523.
Zhou L, Jiang G, Chan G, Santos CP, Severson DW, Xiao L . Michelob_x is the missing inhibitor of apoptosis protein antagonist in mosquito genomes. EMBO Rep 2005; 6: 769–774.
Bryant B, Blair CD, Olson KE, Clem RJ . Annotation and expression profiling of apoptosis-related genes in the yellow fever mosquito, Aedes aegypti. Insect Biochem Mol Biol 2008; 38: 331–345.
Andreadis TG, Becnel JJ, White SE . Infectivity and pathogenicity of a novel baculovirus, CuniNPV from Culex nigripalpus (Diptera: Culicidae) for thirteen species and four genera of mosquitoes. J Med Entomol 2003; 40: 512–517.
Becnel J, White S, Moser B, Fukuda T, Rotstein M, Undeen A et al. Epizootiology and transmission of a newly discovered baculovirus from the mosquitoes Culex nigripalpus and C. quinquefasciatus. J Gen Virol 2001; 82 (Part 2): 275–282.
Moser B, Becnel J, White S, Afonso C, Kutish G, Shanker S et al. Morphological and molecular evidence that Culex nigripalpus baculovirus is an unusual member of the family Baculoviridae. J Gen Virol 2001; 82 (Part 2): 283–297.
Clem RJ, Fechheimer M, Miller LK . Prevention of apoptosis by a baculovirus gene during infection of insect cells. Science 1991; 254: 1388–1390.
Abrams JM, White K, Fessler LI, Steller H . Programmed cell death during Drosophila embryogenesis. Development 1993; 117: 29–43.
Zhou L, Hashimi H, Schwartz LM, Nambu JR . Programmed cell death in the Drosophila central nervous system midline. Curr Biol 1995; 5: 784–790.
Abrams JM, Lux A, Steller H, Krieger M . Macrophages in Drosophila embryos and L2 cells exhibit scavenger receptor-mediated endocytosis. Proc Natl Acad Sci USA 1992; 89: 10375–10379.
Becnel JJ, White SE, Shapiro AM . Culex nigripalpus nucleopolyhedrovirus (CuniNPV) infections in adult mosquitoes and possible mechanisms for dispersal. J Invertebr Pathol 2003; 83: 181–183.
Clem RJ . Baculoviruses and apoptosis: a diversity of genes and responses. Curr Drug Targets 2007; 8: 1069–1074.
Zoog SJ, Schiller JJ, Wetter JA, Chejanovsky N, Friesen PD . Baculovirus apoptotic suppressor P49 is a substrate inhibitor of initiator caspases resistant to P35 in vivo. EMBO J 2002; 21: 5130–5140.
Clarke TE, Clem RJ . In vivo induction of apoptosis correlating with reduced infectivity during baculovirus infection. J Virol 2003; 77: 2227–2232.
Han YS, Thompson J, Kafatos FC, Barillas-Mury C . Molecular interactions between Anopheles stephensi midgut cells and Plasmodium berghei: the time bomb theory of ookinete invasion of mosquitoes. EMBO J 2000; 19: 6030–6040.
Abraham EG, Jacobs-Lorena M . Mosquito midgut barriers to malaria parasite development. Insect Biochem Molec 2004; 34: 667–671.
Steller H . Regulation of apoptosis in Drosophila. Cell Death Differ 2008; 15: 1132–1138.
Zhang Y, Lin N, Carroll PM, Chan G, Guan B, Xiao H et al. Epigenetic blocking of an enhancer region controls irradiation-induced proapoptotic gene expression in Drosophila embryos. Dev Cell 2008; 14: 481–493.
Zhang X, Zhang J, Zhu KY . Chitosan/double-stranded RNA nanoparticle-mediated RNA interference to silence chitin synthase genes through larval feeding in the African malaria mosquito (Anopheles gambiae). Insect Mol Biol 2010; 19: 683–693.
dela Cruz WP, Friesen PD, Fisher AJ . Crystal structure of baculovirus P35 reveals a novel conformational change in the reactive site loop after caspase cleavage. J Biol Chem 2001; 276: 32933–32939.
Afonso CL, Tulman ER, Lu Z, Balinsky CA, Moser BA, Becnel JJ et al. Genome sequence of a baculovirus pathogenic for Culex nigripalpus. J Virol 2001; 75: 11157–11165.
Holley CL, Olson MR, Colon-Ramos DA, Kornbluth S . Reaper eliminates IAP proteins through stimulated IAP degradation and generalized translational inhibition. Nat Cell Biol 2002; 4: 439–444.
Yoo SJ, Huh JR, Muro I, Yu H, Wang L, Wang SL et al. Hid, Rpr and Grim negatively regulate DIAP1 levels through distinct mechanisms. Nat Cell Biol 2002; 4: 416–424.
Fu G, Lees RS, Nimmo D, Aw D, Jin L, Gray P et al. Female-specific flightless phenotype for mosquito control. Proc Natl Acad Sci USA 2010; 107: 4550–4554.
Acknowledgements
This work was supported by NIH Grants R21AI067555 and R56AI079074. We are very grateful for the insightful comments provided by Dr. Rollie Clem on this manuscript and for the excellent technical support provided by Neil Sanscrainte and Kelly Anderson at USDA/CMAVE at Gainesville, FL, USA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by E Baehrecke
Supplementary Information accompanies the paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Liu, B., Becnel, J., Zhang, Y. et al. Induction of reaper ortholog mx in mosquito midgut cells following baculovirus infection. Cell Death Differ 18, 1337–1345 (2011). https://doi.org/10.1038/cdd.2011.8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2011.8