Abstract
CCAAT/enhancer-binding protein-β (C/EBPβ) is a mediator of cell survival and tumorigenesis. When C/EBPβ−/− mice are treated with carcinogens that produce oncogenic Ras mutations in keratinocytes, they respond with abnormally elevated keratinocyte apoptosis and a block in skin tumorigenesis. Although this aberrant carcinogen-induced apoptosis results from abnormal upregulation of p53, it is not known whether upregulated p53 results from oncogenic Ras and its ability to induce p19Arf and/or activate DNA-damage response pathways or from direct carcinogen-induced DNA damage. We report that p19Arf is dramatically elevated in C/EBPβ−/− epidermis and that C/EBPβ represses a p19Arf promoter reporter. To determine whether p19Arf is responsible for the proapoptotic phenotype in C/EBPβ−/− mice, C/EBPβ−/−;p19Arf−/− mice were generated. C/EBPβ−/−;p19Arf−/− mice responded to carcinogen treatment with increased p53 and apoptosis, indicating p19Arf is not essential. To ascertain whether oncogenic Ras activation induces aberrant p53 and apoptosis in C/EBPβ−/− epidermis, we generated K14-ER:Ras;C/EBPβ−/− mice. Oncogenic Ras activation induced by 4-hydroxytamoxifen did not produce increased p53 or apoptosis. Finally, when C/EBPβ−/− mice were treated with differing types of DNA-damaging agents, including alkylating chemotherapeutic agents, they displayed aberrant levels of p53 and apoptosis. These results indicate that C/EBPβ represses p53 to promote cell survival downstream of DNA damage and suggest that inhibition of C/EBPβ may be a target for cancer cotherapy to increase the efficacy of alkylating chemotherapeutic agents.
Similar content being viewed by others
Main
CCAAT/enhancer-binding protein-β (C/EBPβ), a member of the basic leucine zipper transcription factor (bZIP) family, is an important mediator of cell proliferation, differentiation, and survival of several cell types and also functions in inflammatory responses, metabolism, cellular transformation, oncogene-induced senescence, and tumorigenesis.1, 2 C/EBPβ promotes mouse keratinocyte cell survival in response to carcinogen treatment.3, 4 C/EBPβ also functions in the survival of CCl4-treated hepatic stellate cells5 and is required for proliferation and growth factor-dependent survival of Myc/Raf-transformed macrophages through an autoregulatory pathway involving IGF-1.6 Increased expression of C/EBPβ in Wilms tumors is associated with primary tumor relapse, and acute loss of C/EBPβ in a cell line derived from a human metastatic Wilms tumor results in spontaneous apoptosis.7 Similarly, C/EBPβ is necessary to sustain growth and survival of anaplastic lymphoma kinase-positive anaplastic large-cell lymphoma cells.8 These studies demonstrate that C/EBPβ is an important mediator of cell survival and that the role of C/EBPβ in cell survival may provide a mechanism through which C/EBPβ promotes tumorigenesis.
C/EBPβ is stimulated by Ras in keratinocytes,3 and C/EBPβ cooperates with oncogenic Ras to induce cellular transformation, which requires cell cycle-dependent phosphorylation of C/EBPβ on Ser64 and Thr189.3, 9 C/EBPβ is required for Ras-induced skin tumorigenesis as C/EBPβ−/− mice are completely resistant to mouse skin tumorigenesis initiated by topically applied carcinogens such as 7,12-dimethylbenz[a]anthracene (DMBA),3 which functions through its ability to produce oncogenic Ras mutations.10 DMBA treatment of mouse skin produces benign papillomas of which greater than 95% contain an activating A → T mutation in codon 61 of Ras.10 Carcinogen-induced oncogenic mutation of codon 61 can occur in 0.1–5% of Ras genes by error-prone repair within 1–3 days after treatment.11 Treatment of C/EBPβ−/− mouse skin with DMBA results in aberrant increases in p53 levels and function, which is responsible for the vast anomalous induction of apoptosis in 1–2% of C/EBPβ−/− keratinocytes within 1 day of treatment.4 Collectively, these studies suggest that C/EBPβ may be required for the survival of DMBA-initiated oncogenic Ras-containing tumor precursor cells in mouse skin through the regulation of p53 levels and function.
Although it is known that the aberrant increase in p53 is responsible for the proapoptotic phenotype in DMBA-treated C/EBPβ−/− mice, it is not known whether this increase is due to oncogenic Ras. p53 is stabilized and activated in response to different types of stress including oncogenic stress, DNA damage, hypoxia, and nutrient deprivation.12 Oncogenic Ras regulates p53 levels and activity through multiple mechanisms.13 An important, well-established mechanism involves the induction of p19Arf.12 Oncogenic Ras induces p19Arf gene expression through regulation of transcription factors such as Dmp1, AP-1 (JunD, c-Jun, FRA1), and E2F1.14, 15, 16 Induced p19Arf functions by inhibiting the negative feedback loop between p53 and Mdm2, resulting in p53 stabilization and eventually growth arrest or apoptosis.17
Oncogenic Ras can also induce p53-dependent senescence both in culture and in vivo, in part, through activation of DNA-damage response pathways.18, 19, 20 The mechanism through which oncogenic Ras activates DNA-damage response pathways is not clear, but may be due to aberrant DNA replication, resulting in the generation of DNA strand breaks followed by activation of DNA-damage response kinases, ATM/ATR.20 ATM/ATR are the central components of the DNA-damage response induced by both replication stress and DNA-damaging agents that directly alter DNA structure.21 ATM/ATR activation results in phosphorylation of p53, through direct and indirect mechanisms.21 ATM and ATR also contribute to p53 activation by phosphorylating Mdm2, MdmX, and/or E2F1, modulating their ability to regulate p53.21 These post-translational modifications result in increased p53 stability and activity.21
The fact that C/EBPβ is a downstream effector of oncogenic Ras coupled with ample evidence linking oncogenic Ras to p53 prompted us to investigate whether carcinogen-induced aberrant increases in p53 and apoptosis in C/EBPβ−/− mice are a result of oncogenic Ras-induced signaling. We demonstrate that p19Arf is upregulated in C/EBPβ−/− mouse epidermis. However, oncogenic Ras activation and upregulated p19Arf expression are not required for the atypical induction of p53 and apoptosis in C/EBPβ−/− mice. Instead, we found that DNA damage results in anomalous increases in p53 and apoptosis, indicating that C/EBPβ functions downstream of DNA damage to repress p53 and apoptosis.
Results
p19Arf is specifically deregulated in C/EBPβ−/− mouse epidermis and primary epidermal keratinocytes
To begin to determine whether C/EBPβ regulates p53 through a mechanism involving p19Arf, we first examined p19Arf expression in untreated C/EBPβ−/− and wild-type mouse epidermis. Surprisingly, we observed a dramatic 35-fold increase in p19Arf mRNA levels in the epidermis of C/EBPβ−/− mice compared to wild type (Figure 1a). p19Arf protein levels were also significantly increased in epidermal lysates from C/EBPβ−/− mice (Figure 1b). This increase in p19Arf was independent of increased Ras levels and was associated with a modest increase in p53 protein levels (Figure 1b). Our observation that p19Arf is highly expressed in C/EBPβ−/− mouse epidermis was unexpected as p19Arf is not detectable in most normal adult tissues.22, 23 The INK4a-Arf locus encodes both p19Arf and p16INK4a.24 Although they have unique promoters, p19Arf and p16INK4a tumor suppressor genes are often coordinately regulated.25 To determine whether p19Arf expression is specifically upregulated in C/EBPβ−/− mice, or whether p16INK4a expression is similarly altered, we examined p16INK4a mRNA levels from untreated C/EBPβ−/− and wild-type mouse epidermis (Figure 1c). p16INK4a expression was not significantly altered, indicating that p19Arf is specifically deregulated in C/EBPβ−/− mice. Analysis of the proximal 2.6 kb of the p19Arf promoter revealed the identification of several putative C/EBP-binding sites. Cotransfection of C/EBPβ and a 2.6-kb p19Arf promoter reporter demonstrated that C/EBPβ significantly repressed the activity of the p19Arf promoter reporter by 40% (Figure 1d), suggesting that C/EBPβ may directly repress p19Arf expression.
p19Arf is specifically deregulated in C/EBPβ−/− mouse epidermis. (a) Relative p19Arf mRNA levels in C/EBPβ+/+ and C/EBPβ−/− mouse epidermis. *Significantly different from C/EBPβ+/+ as determined by Student's t-test (N=8, P<0.05). Data represent mean±S.E. (b) Representative immunoblot analysis of p19Arf, C/EBPβ Ras, and p53 from lysates isolated from C/EBPβ+/+ and C/EBPβ−/− mouse epidermis. β-actin levels are shown as a loading control. (c) Relative p16INK4a mRNA levels in C/EBPβ+/+ and C/EBPβ−/− mouse epidermis (N=4). Data represent mean±S.E. No significant difference was observed (P=0.376). (d) Balb/MK2 keratinocytes were cotransfected with a p19Arf promoter reporter construct and either empty vector or a C/EBPβ expression vector. Data represent mean±S.E. *Significantly different from basal activity of the p19Arf promoter reporter, as determined by two-factor ANOVA (N=3 independent experiments, P<0.05)
Although not expressed at detectable levels in most normal adult tissues, p19Arf expression is readily induced by ‘culture shock’ when primary cells are placed in culture.26 To investigate whether p19Arf expression is also deregulated in C/EBPβ−/− keratinocytes in response to ‘culture shock’, we examined p19Arf mRNA and protein levels in wild-type and C/EBPβ−/− primary epidermal keratinocytes. C/EBPβ−/− primary keratinocytes displayed a consistent, significant increase in both p19Arf mRNA and protein levels over that observed in wild-type keratinocytes (Figure 2a and b). These results indicate the deregulation of p19Arf is due to a keratinocyte-intrinsic defect.
p19Arf is deregulated in C/EBPβ−/− primary epidermal keratinocytes and is most highly expressed in a sub-population of keratinocytes within C/EBPβ−/− mouse skin. (a) Relative p19Arf mRNA levels in C/EBPβ+/+ and C/EBPβ−/− primary keratinocytes, over time. Data represent mean±S.E. Two-factor ANOVA demonstrated significant interaction between genotype and time (P<0.05). *Significantly different from C/EBPβ+/+ at indicated time points (N=3, P⩽0.05). (b) Representative immunoblot analysis of p19Arf from lysates harvested from C/EBPβ+/+ and C/EBPβ−/− primary keratinocytes. A nonspecific band is shown as a loading control. (c) Representative p19Arf immunohistochemical staining of C/EBPβ+/+ and C/EBPβ−/− mouse epidermis. Arrows indicate p19Arf-positive nucleolar staining. Four skin sections from each of six independent C/EBPβ+/+ and C/EBPβ−/− mice were examined for p19Arf-positive staining
We performed p19Arf immunohistochemistry to localize the expression of p19Arf in skin. p19Arf was detected in C/EBPβ−/− mouse skin, but not in wild-type or in the negative control p19Arf−/− mouse skin (Figure 2c). p19Arf was expressed in a sub-population of C/EBPβ−/− suprabasal keratinocytes located within the upper infundibulum area of the hair follicle. p19Arf localized to nuclei and displayed distinct nucleolar staining. These results demonstrate that p19Arf is greatly increased in C/EBPβ−/− mouse epidermis and primary keratinocytes and is most highly expressed in keratinocytes within the infundibulum of the hair follicle.
p19Arf is not required for carcinogen-induced anomalous increases in p53 and apoptosis in C/EBPβ−/− mice
As p19Arf levels are dramatically increased in C/EBPβ−/− mouse epidermis, we hypothesized that C/EBPβ−/− epidermal keratinocytes may be ‘primed’ to rapidly stabilize and activate p53 following DMBA-induced oncogenic Ras activation, resulting in increased cell death compared to wild-type epidermis. To determine whether elevated p19Arf in C/EBPβ−/− mouse epidermis is involved in the anomalous increases in p53 and apoptosis in response to DMBA treatment, C/EBPβ−/−;p19Arf−/− compound mice were generated. Following DMBA treatment, the number of p53-positive and apoptotic epidermal keratinocytes was quantified in wild-type, p19Arf−/−, C/EBPβ−/−, and C/EBPβ−/−;p19Arf−/− compound mouse epidermis (Figure 3a and b). C/EBPβ−/− mice displayed a significant increase in p53 and apoptosis compared with wild type following carcinogen treatment.4 C/EBPβ−/−;p19Arf−/− compound mice displayed similar anomalous increases in p53 and apoptosis levels compared with C/EBPβ−/− mice, indicating that upregulation of p19Arf in C/EBPβ−/− mouse epidermis is not responsible for the abnormal induction of p53 or apoptosis following DMBA treatment. Representative photomicrographs of p53-immunostained and H and E-stained skin sections showing apoptotic keratinocytes of each of the four DMBA-treated genotypes are shown in Figure 3c.
p19Arf is not required for carcinogen-induced anomalous increases in p53 and apoptosis in C/EBPβ−/− mice. C/EBPβ+/+, p19Arf−/−, C/EBPβ−/−, and C/EBPβ−/−; p19Arf−/− mice were treated topically with 400 nmol DMBA and 20 h later, skin sections were prepared. Skin sections were used to quantify (a) p53-positive basal keratinocytes and (b) apoptotic basal keratinocytes in mouse skin. Data represent mean±S.E. Four skin sections from each of at least three independent mice for each genotype were examined for p53-positive staining or apoptosis. There were no significant difference observed in p53-positive keratinocytes per cm (P=0.769) or apoptotic keratinocytes per cm (P=0.505) between C/EBPβ−/− and C/EBPβ−/−;p19Arf−/− mice; whereas, both genotypes displayed a significant increase in p53-postive keratinocytes and apoptotic keratinocytes compared with C/EBPβ+/+ or p19Arf−/− control mice as determined by ANOVA (N⩾3, P<0.05). (c) Representative p53 immunohistochemical staining (left) and H and E staining (right) of C/EBPβ+/+, p19Arf−/−, C/EBPβ−/−, and C/EBPβ−/−; p19Arf−/− mouse skin sections collected 20 h after treatment with DMBA. Arrows indicate p53-positive (left) or apoptotic basal epidermal keratinocytes (right)
Oncogenic Ras activation is not sufficient to trigger atypical increases in p53 and apoptosis in C/EBPβ−/− mice
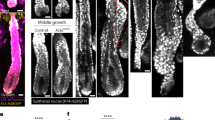
Previous studies from our laboratory demonstrated that forced expression of oncogenic Ras in C/EBPβ−/− primary keratinocytes in culture does not elicit increased apoptosis (data not shown). One possible explanation for this result is that an in vivo epidermal/dermal microenvironment is required for oncogenic Ras to induce aberrant apoptosis in C/EBPβ−/− epidermal keratinocytes. To elucidate whether activation of oncogenic Ras signaling, itself, can elicit aberrant increases in p53 and apoptosis in C/EBPβ−/− mice, we generated K14-ER:Ras;C/EBPβ−/− compound mice. K14-ER:Ras mice are genetically engineered mice that utilize the keratin 14 (K14) promoter to target the expression of the ER:oncogenic RasV12 fusion protein to basal keratinocytes of mouse epidermis. The activity of ER-Ras is regulated through topical treatment with the synthetic ER antagonist, 4-hydroxytamoxifen (4-OHT).27 As shown in Figure 4a and b, the toggle on of oncogenic Ras with a single topical application of 4-OHT induced oncogenic Ras-mediated proliferation in K14-ER:Ras and K14-ER:Ras;C/EBPβ−/− mouse epidermis, as determined by a significant increase in the number of 5-bromo-2-deoxyuridine (BrdU)-positive interfollicular basal epidermal keratinocytes.
Oncogenic Ras activation is not sufficient to trigger atypical increases in p53 and apoptosis in C/EBPβ−/− mice. (a) Representative BrdU immunohistochemical staining of C/EBPβ+/+ and K14-ER:Ras mice collected before (0 h) or after treatment with 4-OHT (24 h). Arrows represent BrdU-positive interfollicular basal keratinocytes. (b) Quantitation of BrdU-positive basal keratinocytes in skin sections collected from untreated (0 h) or 4-OHT-treated (24 h) mice. Data represent mean±S.E. Two-way ANOVA demonstrated significance for genotype, time, and the interaction between genotype and time. K14-ER:Ras and K14-ER:Ras;C/EBPβ−/− mice displayed significant increases in BrdU-positive keratinocytes compared with C/EBPβ+/+ or C/EBPβ−/− control mice 24 h after treatment (N⩾3, P<0.05). (c) p53-positive keratinocytes were quantitated in skin sections collected 12 h after treatment with 4-OHT. Data represent mean±S.E. (N⩾3). No significant differences were observed using two-factor ANOVA (P=0.554). (d) Keratinocyte apoptosis was quantitated in C/EBPβ+/+, C/EBPβ−/−, K14-ER:Ras, and K14ER:Ras;C/EBPβ−/− mice treated with 4-OHT. Data represent the mean±S.E. (N⩾3, for all time points). No significant differences were observed as determined by two-factor ANOVA (P=0.251). For comparison, keratinocyte apoptosis was quantitated in C/EBPβ+/+ and C/EBPβ−/− mouse skin collected 20 h after treatment with 400 nmol DMBA. C/EBPβ−/− mice displayed a significant increase in apoptosis compared with C/EBPβ+/+ as determined by Student's t-test (N=3, P<0.05)
To determine whether activation of oncogenic Ras elicits altered regulation of p53 and apoptosis in C/EBPβ−/− mice, we examined p53 and apoptosis levels in skin sections from C/EBPβ+/+, C/EBPβ−/−, K14-ER:Ras, and K14-ER:Ras;C/EBPβ−/− mice treated with 4-OHT (Figure 4c and d). The toggle on of oncogenic Ras by 4-OHT treatment did not trigger an increase in p53 or apoptosis in K14-ER:Ras;C/EBPβ−/− mice. The number of apoptotic keratinocytes following oncogenic Ras activation was very low, especially compared with the apoptotic response observed in C/EBPβ−/− mice after treatment with DMBA (Figure 4d). Moreover, multiple treatments with 4-OHT were not sufficient to induce aberrant apoptosis in K14-ER:Ras;C/EBPβ−/− mice, despite the induction of significant epidermal hyperplasia consisting of four nucleated cell layers compared to one to two in C/EBPβ+/+ mice and a BrdU interfollicular basal keratinocyte labeling index of approximately 35% compared to 3–5% in C/EBPβ+/+ mice. These data indicate that activation of oncogenic ER-Ras does not induce atypical increases in p53 and apoptosis in K14-ER:Ras;C/EBPβ−/− mouse epidermis.
DNA-damaging agents induce aberrant increases in p53 and apoptosis in C/EBPβ−/− mouse epidermis
To determine whether direct carcinogen-induced DNA damage elicits altered regulation of p53 and apoptosis in C/EBPβ−/− mice, we examined the effects of various types of DNA-damaging agents in C/EBPβ−/− mice compared with C/EBPβ+/+. As shown in Figure 5a, topical treatment with N-methyl-N′-nitro-N-nitrosoguanidine (MNNG) produced a significant increase in the number of p53-positive keratinocytes in C/EBPβ−/− mouse epidermis compared with C/EBPβ+/+ mice. C/EBPβ−/− mice also displayed an anomalous increase in apoptosis when compared to wild type (Figure 5b). In general, these increases were observed across the entire 16-h time course. As MNNG is known to produce oncogenic Ras mutations in mouse skin,28 we examined the effects of ultraviolet B (UVB) radiation in C/EBPβ−/− mice. UVB produces DNA damage mostly in the form of cyclobutane pyrimidine dimers and 6, 4 photoproducts that do not involve mutation of Ras. As shown in Figure 5c, p53 was rapidly and significantly increased in C/EBPβ−/− mice compared with C/EBPβ+/+ mice as early as 3 h after UVB treatment. Similarly, UVB exposure induced a significant increase in apoptosis in C/EBPβ−/− mice compared with wild type (Figure 5d). Similar results were obtained using terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling (TUNEL) staining (data not shown). These results demonstrate that different types of DNA damage induce anomalous increases in p53 and apoptosis in C/EBPβ−/− mice.
DNA-damaging agents induce aberrant increases in p53 and apoptosis in C/EBPβ−/− mouse epidermis. C/EBPβ+/+ and C/EBPβ−/− mice were treated topically with 2.5 μmol of MNNG (a and b) or 100 mJ/cm2 of UVB (c and d), and the treated dorsal skin was collected at the indicated times following treatment. Skin sections were used to quantify p53-positive basal keratinocytes (a and c) or apoptotic interfollicular basal keratinocytes (b and d). Data represent mean±S.E. Two-way ANOVA demonstrated significance for the increase in p53-positive keratinocytes for genotype, time, and the interaction between genotype and time following MNNG and UVB treatment. Two-way ANOVA demonstrated significance for increased apoptosis for genotype, time, and the interaction between genotype and time following MNNG and UVB treatment. *Significantly different from C/EBPβ+/+ at the same time point (N⩾3 for all time points, P<0.05). (e) C/EBPβ+/+, p53−/−, C/EBPβ−/−, and C/EBPβ−/−; p53−/− mice were treated topically with 100 mJ/cm2 of UVB, and the treated dorsal skin was collected 15 h following treatment. Skin sections were used to quantify apoptotic interfollicular basal keratinocytes. Data represent mean±S.E. One-way ANOVA demonstrated significance for increased apoptosis for genotype following UVB treatment. *Significantly different from each genotype (N⩾3 for all time points, P<0.05). (f) Representative p53 immunohistochemical staining or H and E staining (apoptosis) of C/EBPβ+/+ and C/EBPβ−/− mouse epidermis collected either before (0 h) or after treatment with UVB (6 h). Arrows indicate p53-positive or apoptotic basal epidermal keratinocytes, respectively
Previous studies demonstrated that the DMBA-induced proapoptotic phenotype in C/EBPβ−/− mice was dependent on p53 levels and function.4 DMBA functions as a carcinogen by inducing activating oncogenic Ras mutations.10 Our results suggest that increased apoptosis in C/EBPβ−/− mice is a result of DNA damage independent of oncogenic Ras. Thus, we wanted to determine whether p53 is required for increased apoptosis in C/EBPβ−/− mice following treatment with a DNA-damaging agent that functions independently of oncogenic Ras. To investigate this, we treated C/EBPβ+/+, p53−/−, C/EBPβ−/−, and C/EBPβ−/−;p53−/− mice with UVB and determined the number of apoptotic keratinocytes 15 h after treatment (Figure 5e). C/EBPβ−/−;p53−/− mice responded with low levels of apoptosis, similar to that observed for C/EBPβ+/+ and p53−/− control mice. In contrast, C/EBPβ−/− mice displayed a significant increase in apoptosis in response to UVB, similar to results shown in Figure 5d. Representative images of p53 immunohistochemical staining and apoptosis (H and E staining) of C/EBPβ+/+ and C/EBPβ−/− mice before or after UVB exposure are shown (Figure 5f). These results demonstrate that aberrant apoptosis in C/EBPβ−/− mice following UVB-induced DNA damage is p53-dependent.
Intraperitoneal injection of cyclophosphamide in C/EBPβ−/− mice induces aberrant increases in p53 and apoptosis in skin keratinocytes
Our results indicate the loss of C/EBPβ functions to increase the sensitivity of cells to apoptosis induced by DNA-damaging agents. On the basis of these results, we hypothesized that the loss of C/EBPβ would enhance the effectiveness of alkylating chemotherapeutic agents. To examine this, we injected cyclophosphamide intraperitoneally into C/EBPβ+/+ and C/EBPβ−/− mice and examined p53 levels and apoptosis at the remote skin location. Cyclophosphamide is a chemotherapeutic drug given as treatment for various types of cancer including leukemia, lymphoma, breast, ovarian, and bladder. Intraperitoneal injection of cyclophosphamide in C/EBPβ−/− mice resulted in a significant induction of p53 and apoptosis in the skin interfollicular basal keratinocytes over that observed in similarly treated C/EBPβ+/+ mice (Figure 6a and b). These results indicate that the loss of C/EBPβ cooperates with the alkylating chemotherapeutic agent, cyclophosphamide, to enhance cell death.
Intraperitoneal injection of cyclophosphamide in C/EBPβ−/− mice induces aberrant increases in p53 and apoptosis in skin keratinocytes. C/EBPβ+/+ and C/EBPβ−/− mice were treated (i.p.) with 250 mg per kg body weight of cyclophosphamide, and the dorsal skin was collected at the indicated times following treatment. Skin sections were used to quantify (a) p53-positive or (b) apoptotic interfollicular basal keratinocytes. Data represent mean±S.E. Two-way ANOVA demonstrated significance for the increase in p53-positive keratinocytes and increase in apoptosis for genotype, time, and the interaction between genotype and time following cyclophosphamide treatment. *Significantly different from C/EBPβ+/+ at the same time point (N⩾3 for all time points, P<0.05)
C/EBPβ ablation is associated with increased post-translational modification of p53, and p53 transactivation activity is inhibited by C/EBPβ
DMBA treatment of C/EBPβ−/− mice results in increased p53 protein levels without any increase in p53 mRNA levels,4 suggesting that the aberrant increase in p53 protein in the epidermis of DMBA-treated C/EBPβ−/− mice is not due to increased transcription of p53. To further address the role of C/EBPβ in p53 regulation, we conducted ChIP experiments on C/EBPβ+/+ primary keratinocytes to determine whether C/EBPβ was bound to the p53 promoter at a site reported to be a bona fide C/EBP binding site.29 We were unable to demonstrate any binding of C/EBPβ to this area of the p53 promoter. In addition, we did not observe any differential regulation by C/EBPβ of a mouse p53 promoter reporter, which contained the 1.2 kb proximal region of the p53 promoter with or without the above-mentioned C/EBP site (data not shown).29 We conclude that it is unlikely that C/EBPβ directly regulates transcription of p53 in keratinocytes. We next examined whether post-translational modifications associated with enhanced p53 stabilization (mouse S23=human S20) and transactivation (mouse S389=human S392) were differentially present in DMBA-treated C/EBPβ+/+ versus C/EBPβ−/− mice. Using phospho-specific p53 antibodies, we observed that the phosphorylation of both S23 and S389 were significantly increased in DMBA-treated C/EBPβ−/− mice (Figure 7a). These results suggest that the aberrant increases in p53 protein and function in DMBA-treated C/EBPβ−/− mice involve alterations in post-translational modifications of p53. Finally, using a p53-responsive promoter reporter, we observed that C/EBPβ could repress p53 transcriptional activity in keratinocytes (Figure 7b). These data are consistent with previous findings in human endometrial stromal cells.30 Our results indicate that the ablation of C/EBPβ is associated with post-translational modifications of p53, and C/EBPβ can inhibit the transcriptional activity of p53 in keratinocytes.
C/EBPβ ablation is associated with increased post-translational modification of p53, and p53 tranactivational activity is inhibited by C/EBPβ. (a) C/EBPβ+/+ and C/EBPβ−/− mice were treated topically with vehicle control (acetone) or 400 nmol DMBA, and the treated shaved dorsal skin was collected 20 h after treatment. The table contains the number of S23-p53-positive (human S20) or S389-p53-positive (human S392) basal epidermal keratinocytes for the indicated genotypes following treatment. Data represent mean±S.E. *Significantly different from C/EBPβ+/+ following the same treatment as determined by Student's t-test (N=3, P<0.01). (b) Balb/MK2 keratinocytes were cotransfected with a p53-responsive promoter reporter construct (P133/70) or a negative control promoter reporter construct lacking the p53-responsive promoter (P133/41) and empty vector, a C/EBPβ expression vector, a p53 expression vector, or C/EBPβ and p53 expression vector(s) together. Data represent mean±S.E. *Significantly different from basal activity of the p53-responsive promoter reporter, as determined by Student's t-test (N=6 per group from two independent experiments, P<0.05)
Discussion
As described in the introduction, there is a substantial body of evidence that suggested the enhanced proapoptotic response in C/EBPβ−/− epidermal keratinocytes is likely a result of carcinogen-induced oncogenic Ras activation. Using K14-ER:Ras;C/EBPβ−/− mice, we observed that conditional activation of oncogenic Ras12V in epidermal basal keratinocytes of C/EBPβ−/− mouse epidermis was insufficient to induce anomalous increases in p53 and apoptosis. Our results indicate that the proapoptotic phenotype in carcinogen-treated C/EBPβ−/− mice is not due to oncogenic Ras. Instead, we observed that treatment with a diverse group of DNA-damaging agents, including UVB radiation, MNNG, DMBA, and cyclophosphamide all resulted in abnormal increases in p53 and apoptosis. These findings indicate that C/EBPβ participates in the DNA-damage response network and functions to promote cell survival by repressing p53 levels and function in response to DNA damage. Although a parallel increase in p53 and apoptosis in C/EBPβ−/− mouse epidermis was observed following exposure to DNA-damaging agents, apoptosis was found to persist after p53 levels returned to control levels. As induction of p53 is an early event in apoptosis and apoptosis requires approximately 4 h from commitment to termination,31 this could contribute to the sustained apoptosis that is observed in C/EBPβ−/− mice.
Regulation of p53 is complex, involving at least 10 positive and negative feedback loops that modulate p53 expression, stability, and activity.32 It is not yet known how C/EBPβ participates within the DNA-damage signaling cascade to repress p53 levels and function. C/EBPβ can function as a transcriptional repressor.33, 34, 35, 36 Basal p53 mRNA levels were increased 2.5-fold in C/EBPβ−/− mouse epidermis compared with wild-type mice, suggesting a repressor function for C/EBPβ in the regulation of basal p53 mRNA levels.4 However, DMBA treatment did not further increase p53 mRNA levels, indicating that the increased p53 protein levels in DMBA-treated C/EBPβ−/− mouse epidermis are not due to upregulation of p53 mRNA.4 On the basis of these results and results presented in the current study, it is unlikely that C/EBPβ regulates the transcription of p53 in epidermal keratinocytes in response to DNA damage. DNA-damage response pathways may modulate C/EBPβ levels or activity, thereby altering the ability of C/EBPβ to regulate the expression of genes or activity of proteins involved in post-translational modification of p53. Accordingly, C/EBPβ protein levels are induced in keratinocytes following exposure to UVB.37 Elevated levels of C/EBPβ may repress p53 by regulating the expression of genes that contribute to phosphorylation, acetylation, or sumoylation of p53 following DNA damage. Our results indicate ablation of C/EBPβ is associated with increased post-translational modifications of p53, which are known to enhance p53 stability and activity. Additional studies are required to determine whether these changes are a cause or consequence of elevated p53. C/EBPβ has been shown to interact with p53, disrupting p53 DNA binding and transcriptional activity,30 and we provide evidence that this mechanism of inhibition of p53 transcriptional activity by C/EBPβ also occurs in keratinocytes. This mechanism could contribute to the negative regulation of p53 by C/EBPβ and loss of C/EBPβ could result in abnormally elevated p53-mediated apoptosis observed in C/EBPβ−/− mice treated with DNA-damaging agents.
Although we demonstrate that direct DNA damage is an important signal in eliciting C/EBPβ-mediated repression of p53 to promote keratinocyte cell survival, additional mechanisms in other cell types contribute to C/EBPβ-mediated cell survival.5, 6 For example, CCl4 treatment, which induces DNA damage through the generation of free radicals, results in phosphorylation of C/EBPβ by ribosomal protein 6-kinase, producing a functional XEXD caspase substrate/inhibitor box that is critical for survival of hepatic stellate cells.5 In addition, C/EBPβ is required for growth factor-dependent survival of Myc/Raf-transformed macrophages through an autoregulatory pathway involving IGF-1.6 Activation of several oncogenes including Myc, Ras, Raf, cyclin E, Cdc25A, and E2F1 can engage DNA-damage response pathways, resulting in activation of p53 and the induction of apoptosis or senescence.18, 20, 38 The important decision of cell survival or cell death in response to different stimuli is likely controlled by more than one signal. Thus, it is possible that there is overlap of these pathways in their ability to contribute to C/EBPβ-mediated cell survival depending on the cellular and genetic context.
Our results reveal that p19Arf is upregulated in untreated C/EBPβ−/− mouse epidermis and primary keratinocytes. The magnitude of p19Arf expression in C/EBPβ−/− mouse epidermis was unexpected because p19Arf is not detectably expressed during embryonic development and is found at very low levels and only in a small subset of normal adult tissues, such as the testes.22, 23 Localization of p19Arf in C/EBPβ−/− mouse skin demonstrated that p19Arf was detectable in the nucleoli of keratinocytes in the infundibulum of the hair follicle. We observed that p19Arf is not upregulated in the epidermis of mice in which C/EBPα has been ablated (unpublished results), demonstrating that the regulation of p19Arf is C/EBP isoform specific to C/EBPβ.
The biological significance of deregulated p19Arf in C/EBPβ−/− mouse skin and exactly how C/EBPβ regulates p19Arf expression remains to be determined. p19Arf is known for its role as a tumor suppressor gene, through its ability to inhibit Mdm2 and thereby derepress p53.17 However, p19Arf functions through p53-independent mechanisms, including the direct repression of E2F1 or c-Myc transcriptional activity.17 Although our results indicate that p19Arf is not involved in the p53-dependent proapoptotic phenotype in DMBA-treated C/EBPβ−/− mice, it is unclear whether or not p19Arf functions to influence C/EBPβ−/− mouse skin homeostasis or tumorigenesis. p19Arf expression is under strict regulation, as revealed by the increasing number of transcription factors such as Pokemon, E2F3b, p53, Twist, Bmi1, Tbx2/3, and JunD, which repress p19Arf.25 Our results suggest that C/EBPβ may also contribute to the negative regulation of p19Arf through a mechanism involving repression of the p19Arf proximal promoter.
In summary, our results indicate that C/EBPβ represses p53 to promote cell survival downstream of DNA damage. These findings uncover a novel link between the DNA-damage response pathway, C/EBPβ, and the repression of p53 to promote cell survival. The fact that the loss of C/EBPβ significantly enhances the proapoptotic effect of systemically administered cyclophosphamide, an alkylating chemotherapeutic, suggests that C/EBPβ may be a molecular target for cotherapy. The inhibition of C/EBPβ in p53-proficient tumor cells could cooperate with alkylating chemotherapeutic agents to enhance apoptosis and tumor regression.
Materials and Methods
Animals
B6.129-C/EBPβ−/− mice, B6.129-C/EBPβ−/−;p53−/−, and p53−/− mice have been previously described.3, 4 B6.129-Cdkn2atm1Cjs (p19Arf−/−) mice were obtained from the Mouse Models of Human Cancers Consortium (MMHCC), generously made available by Charles Sherr (St. Jude's Research Hospital, TN, USA).39 Mice were genotyped as described by the MMHCC PCR protocol for strain no. 01XG7. p19Arf−/− female mice were crossed with C/EBPβ−/− male mice. Offsprings (C/EBPβ+/−;p19Arf+/−) were crossed to generate all genotypes utilized in these studies. K14-ER:Ras mice were a kind gift from Dr. Paul Khavari (Stanford University, CA, USA).27 K14-ER:Ras female mice were crossed with C/EBPβ−/− male mice to generate K14-ER:Ras;C/EBPβ+/− mice. K14-ER:Ras;C/EBPβ+/− mice were crossed with B6.129-C/EBPβ+/− mice to generate all genotypes used in these studies. Mice were genotyped for ER:Ras by PCR using the following primers: forward, CACCACCAGCTCCACTTCAGCACATT; reverse, CGCACCAACGTGTAGAAGGCATCCTC. The dorsal hair of mice (7–11 weeks old) was clipped with electric clippers at least 2 days before use. Mice were treated as described in the text. DMBA (catalog no. 40818, Fisher Scientific USA – Acros Organics, Pittsburgh, PA, USA) was dissolved in acetone (400 nmol/200 μl). 4-OHT (catalog no. H7904) and MNNG (catalog no. 12994-1) were purchased from Sigma (St. Louis, MO, USA). 4-OHT was dissolved in 95% ethanol (1 mg/100 μl). MNNG was dissolved in acetone (2.5 μmol/200 μl of acetone). Mice were treated with cyclophosphamide monohydrate (Sigma, catalog no. C0768) prepared in phosphate-buffered saline (PBS, 25 mg/ml) by intraperitoneal injection at a final dose of 250 mg per kg body weight. The UV lamp used in this study, purchased from UVP Inc. (catalog no. 34-0039-01; Upland, CA, USA), emits wavelengths between 280 and 350 nm with a spectrum peak at 312 nm. The light intensity of the lamp was measured using the IL-1400A radiometer (International Light, Newburyport, MA, USA) equipped with the SEL240/UVB-1 sensor, which detects wavelengths between 265 and 332 nm. Mice were treated with 100 mJ/cm2 of UVB radiation.
Cell lines and cell culture
Balb/MK2 mouse keratinocytes (Bernard Weissman, University of North Carolina, NC, USA) were cultured as described previously.37 Mouse primary epidermal keratinocytes were isolated from newborn C/EBPβ+/+ or C/EBPβ−/− littermates as described previously.37 Isolated keratinocytes were plated at 3 × 106 cells per well in six-well culture dishes in complete media (Ca2+-free EMEM supplemented with 10% non-Chelex-treated FBS, 10 ng of hEGF per ml, 100 U of penicillin per ml, 100 μg of streptomycin per ml, and 250 ng of amphotericin B per ml) for 4 h. Cultures were gently washed with Mg2+- and Ca2+-free PBS and re-fed with keratinocyte serum-free medium (Ca2+-free K-SFM supplemented with 2.5 μg of hEGF, 25 mg bovine pituitary extract, 5 μg of gentimycin per ml, and calcium chloride to a final concentration of 0.05 mM). Three independent pools of isolated primary keratinocytes were examined for p19Arf mRNA and protein expression at each time point. Primary keratinocytes collected at time 0 were isolated as above; 1 × 106 cells were pelleted by centrifugation and subsequently used to harvest total RNA or protein lysates as described.
Construction of pGL4.10-p19Arf 2.6 kb promoter reporter and luciferase assay
pGL2-p19Arf 2.6 kb promoter reporter was kindly provided by Kazushi Inoue (Wake Forest University, NC, USA). The p19Arf 2.6 kb promoter fragment was cloned into pGL4.10 using KpnI and BglII. Balb/MK2 keratinocytes were cotransfected in 12-well plates with 300 ng per well of pGL4.10-p19Arf 2.6 kb or pGL4.10 and the indicated amounts of pcDNA3.1(−)-C/EBPβ expression plasmids using Lipofectamine (Invitrogen, Carlsbad, CA, USA) in triplicate. Cells were washed with ice-cold PBS, lysed in 1 × Cell Culture Lysis Buffer (Promega, Madison, WI, USA), harvested by scraping, and cleared by centrifugation before analyzing for luciferase activity. Data were normalized to total protein and activity of the empty reporter construct. The average relative fold change compared to p19Arf basal promoter activity, arbitrarily set to 1, for three independent experiments are shown±S.E. A two-factor ANOVA was conducted on the relative fold-change values. To test for repression, a t-test, using the pooled ANOVA error, was carried out for each treatment to test the null hypothesis that mean relative fold change=1 versus the alternative that mean relative fold change <1.
The p53-responsive promoter reporter constructs (P133/70) and the negative control lacking the p53-responsive promoter (P133/41) have been described previously40 and were a kind gift from Thomas Eling (National Institute of Environmental Health Sciences, Research Triangle Park, NC, USA). Balb/MK2 keratinocytes were cotransfected in 12-well plates with 400 ng per well of P133/70 or P133/41 and the indicated expression vectors at 200 ng per well (empty vector (pcDNA3.1), C/EBPβ, p53, or C/EBPβ and p53) using FastTransfect reagent (catalog no. E2431, Promega). Total DNA transfected was maintained at a constant level for all the wells using pcDNA3.1. Cells were lysed 48 h post-transfection and processed as described above. Data are described as the average fluorescence units normalized to total protein±S.E.
Quantitative real-time RT-PCR
Total RNA was collected and prepared for quantitative real-time RT-PCR as previously described.4 Detection of p19Arf mRNA levels was conducted using a Cdkn2a (p19Arf) TaqMan® Gene Expression Assay (catalog no. Mm01257348_m1; Applied Biosystems, Foster City, CA, USA). Sequences for p16INK4a primers and dual-labeled probe were provided by Bryan Betz (National Institute of Environmental Health Sciences), synthesized by Sigma Genosys, and used at a final concentration of 0.5 μM (forward primer: CCCAACGCCCCGAACT, reverse primer: GTGAACGTTGCCCATCATCA, probe: [FAM]TTTCGGTCGTACCCCGATTCAGGTG[TAM]). Each independent sample was run in triplicate. Expression levels for in vivo samples were normalized to 18S (Applied Biosystems) and expression levels for primary keratinocyte samples were normalized to GAPDH (Applied Biosystems). Data were analyzed using the comparative CT method and presented relative to the control, C/EBPβ+/+, or C/EBPβ+/+ at time zero. N=4–8 mice/genotype for in vivo analysis and N=3 independent pooled keratinocytes/time for the primary keratinocyte experiment.
Preparation of protein lysates
Isolation of protein lysates from epidermal homogenates was performed as previously described,4 with the exception that samples were homogenized on ice before sonication. For preparation of protein lysates from cultured primary keratinocytes, cells were washed with cold PBS, harvested by scraping, and collected by centrifugation. Cells were lysed on ice in radioimmunoprecipitation buffer (1% Nonidet P-40, 0.5% sodium deoxycholate, 0.1% sodium dodecyl sulfate (SDS), 1 mM phenylmethylsulfonyl fluoride, 1 mM sodium orthovanadate, and 1 × protease inhibitor cocktail (Roche, Basel, Switzerland) in PBS), sonicated and cleared by centrifugation at 14 000 × g for 10 min. Supernatants were stored at −80 °C until use. Protein concentration was determined by Bio-Rad Protein Assay reagent (Bio-Rad, Hercules, CA, USA).
Immunoblot analysis
Equal amounts of protein (10–40 μg) were dissolved in SDS sample buffer, boiled, and separated on 12% SDS-polyacrylamide gel electrophoresis. Immunoblot analysis was conducted as described previously.4 Membranes were probed with antibody for p19Arf (1 : 400, sc-32748), C/EBPβ (1 : 2000, sc-150), H-Ras (1 : 2000, sc-520), or p53 (1 : 2000, sc-6243) from Santa Cruz Biotechnology (Santa Cruz, CA, USA) or β-actin (1 : 20 000, A-5441) from Sigma. The membranes were washed and probed with a horseradish peroxidase-conjugated secondary antibody for rat (1 : 2000, sc-2006; Santa Cruz Biotechnology), rabbit (1 : 2500, NA934 V; GE Healthcare UK Ltd, Buckinghamshire, England), or mouse (1 : 10 000, NXA931; GE Healthcare UK Ltd).
Immunohistochemical staining
Untreated or treated, dorsal skin was excised and fixed for 24 h in 10% neutral-buffered formalin. Four independent areas of skin were collected, embedded in paraffin, and cut into sections (≈5 μM). For p53 immunohistochemical staining, sections were deparaffinized, treated with 3% H2O2, and subjected to antigen retrieval with citrate buffer (pH 6.0) in a 95 °C water bath for 30 min. Sections were blocked with normal goat serum for 30 min at room temperature and then incubated with rabbit polyclonal anti-p53 antibody (catalog no. sc-6243, 1 : 500), purchased from Santa Cruz Biotechnology, at 4 °C for 24 h. For phospho-specific p53 immunohistochemical staining, the following antibodies were purchased from Cell Signaling Technology Inc. (Danvers, MA, USA): rabbit polyclonal anti-phospho-p53-(Ser20) (catalog no. 9287, 1 : 200) and anti-phospho-p53-(Ser392) (catalog no. 9281, 1 : 300). Staining for phospho-specific p53 was conducted in the same way as for wild-type p53. Sections were subsequently incubated with biotinylated goat anti-rabbit IgG at room temperature for 30 min. Staining was detected using the rabbit Vectastain Elite ABC kit (catalog no. PK-6101) from Vector Laboratories (Burlingame, CA, USA) and 3,3′-diaminobenzidine (DAB) (BioGenex, San Ramon, CA, USA) following the manufacturers’ protocols. The sections were counterstained with hematoxylin, dehydrated, and mounted. Data are expressed as the average number of p53-positive staining interfollicular basal keratinocytes per centimeter of mouse skin±S.E. p19Arf immunohistochemical staining was modified from a previously described method23 by Bryan Betz (National Institute of Environmental Health Sciences). Briefly, paraffin-embedded sections from mouse skin were deparaffinized and subjected to antigen retrieval in antigen decloaker citrate buffer (pH 6.0) (Biocare, Concord, CA, USA) for 45 min in the decloaking chamber (Biocare). After cooling, slides were treated with 3% H2O2 to quench endogenous peroxide activity. Sections were blocked with normal rabbit serum in 0.1% Triton X-100/PBS for approximately 1 h at room temperature. Rat polyclonal anti-p19Arf primary antibody (catalog no. 05-929), purchased from Upstate (Chicago, IL, USA), was diluted 1 : 15 in 0.1% Triton X-100/PBS and added to sections for incubation at 4 °C overnight. Sections were subsequently incubated with biotinylated rabbit anti-rat IgG at room temperature for 30 min. Staining was detected using the rat Vectastain Elite ABC kit (catalog no. PK-6104) and DAB, following the manufacturers’ protocols. The sections were counterstained with hematoxylin, dehydrated, and mounted. In vivo BrdU labeling and immunohistochemical staining were conducted by administering 100 mg per kg body weight of BrdU in 200 ml of PBS by i.p. injection 1 h before killing the mice. Skin sections were deparaffinized, incubated in 2 M HCl for 30 min at 37 °C, washed in borate buffer for 3 min at room temperature, followed by digestion in 0.001% trypsin for 3 min at 37 °C. Endogenous peroxidase activity was inhibited by incubation in 3% H2O2 for 10 min. Sections were blocked in horse serum for 20 min and then incubated in anti-BrdU IgG primary antibody (catalog no. 347580; BD Biosciences, San Jose, CA, USA) at a dilution of 1 : 25 for 1 h at room temperature. Slides were further processed using the Vectastain Elite mouse ABC kit (catalog no. PK-6102). Immunohistochemical staining for BrdU was quantified and summarized as the average number of BrdU-positive interfollicular basal keratinocytes per centimeter of mouse skin±S.E., N⩾3.
Detection of apoptotic keratinocytes
Mouse skin sections stained with H and E were used to quantify the presence of apoptotic keratinocytes. As previously described,3, 4 apoptotic keratinocytes in the interfollicular basal epidermis were scored positive if all three of the following criteria were present: dark pyknotic nuclei, cytoplasmic eosinophilia, and absence of cellular contacts. TUNEL staining was conducted as previously described4 to confirm the apoptosis results obtained using H and E-stained sections. Data are presented as the average number of apoptotic keratinocytes per centimeter of mouse skin±S.E.
Statistical analysis
Statistical analysis was performed as described in the text using SASv9.1.3 Service Pack 4.
Abbreviations
- 4-OHT:
-
4-hydroxytamoxifen
- BrdU:
-
5-bromo-2-deoxyuridine
- C/EBPβ:
-
CCAAT/enhancer-binding protein-β
- DMBA:
-
7,12-dimethylbenz[a]anthracene
- K14:
-
keratin 14
- MNNG:
-
N-methyl-N′-nitro-N-nitrosoguanidine
- TUNEL:
-
terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling
- UVB:
-
ultraviolet B radiation
References
Ramji DP, Foka P . CCAAT/enhancer-binding proteins: structure, function and regulation. Biochem J 2002; 365 (Part 3): 561–575.
Sebastian T, Johnson PF . Stop and go: anti-proliferative and mitogenic functions of the transcription factor C/EBPbeta. Cell Cycle 2006; 5: 953–957.
Zhu S, Yoon K, Sterneck E, Johnson PF, Smart RC . CCAAT/enhancer binding protein-beta is a mediator of keratinocyte survival and skin tumorigenesis involving oncogenic Ras signaling. PNAS 2002; 99: 207–212.
Yoon K, Zhu S, Ewing SJ, Smart RC . Decreased survival of C/EBPbeta-deficient keratinocytes is due to aberrant regulation of p53 levels and function. Oncogene 2007; 26: 360–367.
Buck M, Poli V, Hunter T, Chojkier M . C/EBPbeta phosphorylation by RSK creates a functional XEXD caspase inhibitory box critical for cell survival. Mol Cell 2001; 8: 807–816.
Wessells J, Yakar S, Johnson PF . Critical prosurvival roles for C/EBP beta and insulin-like growth factor I in macrophage tumor cells. Mol Cell Biol 2004; 24: 3238–3250.
Li W, Kessler P, Yeger H, Alami J, Reeve AE, Heathcott R et al. A gene expression signature for relapse of primary wilms tumors. Cancer Res 2005; 65: 2592–2601.
Piva R, Pellegrino E, Mattioli M, Agnelli L, Lombardi L, Boccalatte F et al. Functional validation of the anaplastic lymphoma kinase signature identifies CEBPB and BCL2A1 as critical target genes. J Clin Invest 2006; 116: 3171–3182.
Shuman JD, Sebastian T, Kaldis P, Copeland TD, Zhu S, Smart RC et al. Cell cycle-dependent phosphorylation of C/EBP{beta} mediates oncogenic cooperativity between C/EBP{beta} and H-RasV12. Mol Cell Biol 2004; 24: 7380–7391.
Quintanilla M, Brown K, Ramsden M, Balmain A . Carcinogen-specific mutation and amplification of Ha-ras during mouse skin carcinogenesis. Nature 1986; 322: 78–80.
Chakravarti D, Mailander PC, Cavalieri EL, Rogan EG . Evidence that error-prone DNA repair converts dibenzo[a,l]pyrene-induced depurinating lesions into mutations: formation, clonal proliferation and regression of initiated cells carrying H-ras oncogene mutations in early preneoplasia. Mutat Res 2000; 456: 17–32.
Horn HF, Vousden KH . Coping with stress: multiple ways to activate p53. Oncogene 2007; 26: 1306–1316.
McMahon M, Woods D . Regulation of the p53 pathway by Ras, the plot thickens. Biochim Biophys Acta 2001; 1471: M63–M71.
Dimri GP, Itahana K, Acosta M, Campisi J . Regulation of a senescence checkpoint response by the E2F1 transcription factor and p14(ARF) tumor suppressor. Mol Cell Biol 2000; 20: 273–285.
Sreeramaneni R, Chaudhry A, McMahon M, Sherr CJ, Inoue K . Ras-Raf-Arf signaling critically depends on the Dmp1 transcription factor. Mol Cell Biol 2005; 25: 220–232.
Weitzman JB, Fiette L, Matsuo K, Yaniv M . JunD protects cells from p53-dependent senescence and apoptosis. Mol Cell 2000; 6: 1109–1119.
Sherr CJ . Divorcing ARF and p53: an unsettled case. Nat Rev Cancer 2006; 6: 663–673.
Di Micco R, Fumagalli M, Cicalese A, Piccinin S, Gasparini P, Luise C et al. Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature 2006; 444: 638–642.
Gorgoulis VG, Vassiliou LV, Karakaidos P, Zacharatos P, Kotsinas A, Liloglou T et al. Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 2005; 434: 907–913.
Bartkova J, Horejsi Z, Koed K, Kramer A, Tort F, Zieger K et al. DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 2005; 434: 864–870.
Wahl GM, Carr AM . The evolution of diverse biological responses to DNA damage: insights from yeast and p53. Nat Cell Biol 2001; 3: E277–E286.
Zindy F, Quelle DE, Roussel MF, Sherr CJ . Expression of the p16INK4a tumor suppressor versus other INK4 family members during mouse development and aging. Oncogene 1997; 15: 203–211.
Bertwistle D, Zindy F, Sherr CJ, Roussel MF . Monoclonal antibodies to the mouse p19(Arf) tumor suppressor protein. Hybrid Hybridomics 2004; 23: 293–300.
Quelle DE, Zindy F, Ashmun RA, Sherr CJ . Alternative reading frames of the INK4a tumor suppressor gene encode two unrelated proteins capable of inducing cell cycle arrest. Cell 1995; 83: 993–1000.
Gil J, Peters G . Regulation of the INK4b-ARF-INK4a tumour suppressor locus: all for one or one for all. Nat Rev Mol Cell Biol 2006; 7: 667–677.
Sherr CJ, DePinho RA . Cellular senescence: mitotic clock or culture shock? Cell 2000; 102: 407–410.
Tarutani M, Cai T, Dajee M, Khavari PA . Inducible activation of Ras and Raf in adult epidermis. Cancer Res 2003; 63: 319–323.
Brown K, Buchmann A, Balmain A . Carcinogen-induced mutations in the mouse c-Ha-ras gene provide evidence of multiple pathways for tumor progression. Proc Natl Acad Sci USA 1990; 87: 538–542.
Boggs K, Reisman D . C/EBPbeta participates in regulating transcription of the p53 gene in response to mitogen stimulation. J Biol Chem 2007; 282: 7982–7990.
Schneider-Merck T, Pohnke Y, Kempf R, Christian M, Brosens JJ, Gellersen B . Physical interaction and mutual transrepression between CCAAT/enhancer-binding protein beta and the p53 tumor suppressor. J Biol Chem 2006; 281: 269–278.
Gavrieli Y, Sherman Y, Ben-Sasson SA . Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J Cell Biol 1992; 119: 493–501.
Harris SL, Levine AJ . The p53 pathway: positive and negative feedback loops. Oncogene 2005; 24: 2899–2908.
Corbi AL, Jensen UB, Watt FM . The alpha2 and alpha5 integrin genes: identification of transcription factors that regulate promoter activity in epidermal keratinocytes. FEBS Lett 2000; 474: 201–207.
Burkart AD, Mukherjee A, Sterneck E, Johnson PF, Mayo KE . Repression of the inhibin alpha-subunit gene by the transcription factor CCAAT/enhancer-binding protein-beta. Endocrinology 2005; 146: 1909–1921.
Di-Poi N, Desvergne B, Michalik L, Wahli W . Transcriptional repression of peroxisome proliferator-activated receptor beta/delta in murine keratinocytes by CCAAT/enhancer-binding proteins. J Biol Chem 2005; 280: 38700–38710.
Sankpal NV, Mayo MW, Powell SM . Transcriptional repression of TFF1 in gastric epithelial cells by CCAAT/enhancer binding protein-beta. Biochim Biophys Acta 2005; 1728: 1–10.
Yoon K, Smart RC . C/EBP{alpha} is a DNA damage-inducible p53-regulated mediator of the G1 checkpoint in keratinocytes. Mol Cell Biol 2004; 24: 10650–10660.
Hong S, Pusapati RV, Powers JT, Johnson DG . Oncogenes and the DNA damage response: Myc and E2F1 engage the ATM signaling pathway to activate p53 and induce apoptosis. Cell Cycle 2006; 5: 801–803.
Kamijo T, Zindy F, Roussel MF, Quelle DE, Downing JR, Ashmun RA et al. Tumor suppression at the mouse INK4a locus mediated by the alternative reading frame product p19ARF. Cell 1997; 91: 649–659.
Baek SJ, Wilson LC, Eling TE . Resveratrol enhances the expression of non-steroidal anti-inflammatory drug-activated gene (NAG-1) by increasing the expression of p53. Carcinogenesis 2002; 23: 425–434.
Acknowledgements
We would like to thank Dr. Cavell Brownie for help with statistical analysis, Bryan Betz for his assistance with p19Arf immunohistochemistry and providing p16INK4a primers and probe sequences, and Sara Dixon for quantification of proliferation in the K14-ER:Ras;C/EBPβ−/− mice. This work was supported by a grant from the National Cancer Institute (CA046637) and a grant from the National Institute of Environmental Health Sciences (ES012473).
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by V De Laurenzi
Rights and permissions
About this article
Cite this article
Ewing, S., Zhu, S., Zhu, F. et al. C/EBPβ represses p53 to promote cell survival downstream of DNA damage independent of oncogenic Ras and p19Arf. Cell Death Differ 15, 1734–1744 (2008). https://doi.org/10.1038/cdd.2008.105
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.105
Keywords
This article is cited by
-
Combined absence of TRP53 target genes ZMAT3, PUMA and p21 cause a high incidence of cancer in mice
Cell Death & Differentiation (2024)
-
Regulation of the bi-directional cross-talk between ovarian cancer cells and adipocytes by SPARC
Oncogene (2019)
-
C/EBPβ deletion in oncogenic Ras skin tumors is a synthetic lethal event
Cell Death & Disease (2018)
-
Identifying conserved UV exposure genes and mechanisms
Scientific Reports (2018)
-
C/EBPα expression is partially regulated by C/EBPβ in response to DNA damage and C/EBPα-deficient fibroblasts display an impaired G1 checkpoint
Oncogene (2009)