Abstract
Background:
Selecting the most beneficial treatment regimens for colorectal cancer (CRC) patients remains challenging due to a lack of prognostic markers. Members of the Myosin family, proteins recognised to have a major role in trafficking and polarisation of cells, have recently been reported to be closely associated with several types of cancer and might thus serve as potential prognostic markers in the context of CRC.
Methods:
We used a previously established meta-analysis of publicly available gene expression data to analyse the expression of different members of the Myosin V family, namely MYO5A, 5B, and 5C, in CRC. Using laser-microdissected material as well as tissue microarrays from paired human CRC samples, we validated both RNA and protein expression of Myosin Vb (MYO5B) and its known adapter proteins (RAB8A and RAB25) in an independent patient cohort. Finally, we assessed the prognostic value of both MYO5B and its adapter-coupled combinatorial gene expression signatures.
Results:
The meta-analysis as well as an independent patient cohort study revealed a methylation-independent loss of MYO5B expression in CRC that matched disease progression. Although MYO5B mutations were identified in a small number of patients, these cannot be solely responsible for the common downregulation observed in CRC patients. Significantly, CRC patients with low MYO5B expression displayed shorter overall, disease-, and metastasis-free survival, a trend that was further reinforced when RAB8A expression was also taken into account.
Conclusions:
Our data identify MYO5B as a powerful prognostic biomarker in CRC, especially in early stages (stages I and II), which might help stratifying patients with stage II for adjuvant chemotherapy.
Similar content being viewed by others
Main
Colorectal cancer (CRC) is one of the most frequent and deadly cancers in the western world with more than 1.2 million yearly diagnoses and ∼600 000 deaths each year (Torre et al, 2015). Patient survival is largely dependent on early diagnosis and intervention. Accordingly, there is an urgent need for novel diagnostic parameters as well as molecular determinants of clinical outcome, which would allow for the targeted treatment of patients at risk of relapse. Especially in stage II patients, the identification of biomarkers predicting the recurrence of the disease is an unmet clinical need. By using a variety of resources, we previously demonstrated that the suppressor of cytokine signalling 2 serves as a promising marker with a strong prognostic value for early CRC stages (Letellier et al, 2014).
Myosins constitute a superfamily of actin-dependent molecular motors that are involved in a large number of essential cellular functions, such as cell motility, endocytosis, vesicle trafficking, and protein/RNA localisation (Ouderkirk and Krendel, 2014). In the intestine, myosins are responsible for supporting the apical brush in enterocytes, leading to the emergence of polarised differentiated epithelial cells that possess an apical and a basolateral domain. Myosin Vb (MYO5B) is a class V member of non-conventional myosins, which provide a molecular basis for the transport of actin-dependent organelles and regulate membrane traffic in polarised epithelial cells (Trybus, 2008). Loss-of-function mutations in the MYO5B gene lead to a disruption in cell polarity and the absence of microvilli at the enterocyte surface. This condition, which is known as microvillus inclusion disease (MVID), is a rare disorder that affects newborn infants during their first months, leading to malabsorption and potentially life-threatening marked diarrhoea (Müller et al, 2008). Some of the associated mutations have been proposed to affect MYO5B expression levels via nonsense-mediated RNA decay (Szperl et al, 2011).
Members of the Rab family small GTPases, consisting of RAB8 and RAB11, have recently been suggested to act as potential mediators in MYO5B-mediated vesicle transport (Fan et al, 2004; Ishikura and Klip, 2008; Roland et al, 2011). In mammalian cells, the RAB11 family consists of three proteins: RAB11A (Chavrier et al, 1990; Goldenring et al, 1994), RAB11B (Lai et al, 1994), and RAB25 (Goldenring et al, 1993). Rab GTPases function as molecular switches and the GTP-bound active conformation recruits effector proteins, such as MYO5B, that operate specifically for different membrane traffic steps (Roland et al, 2011). Whereas some RABs bind indirectly through adapters, others, such as RAB8A and RAB11A, interact directly with MYO5B: MYO5B has specific binding regions in its tail domain and interacts with RAB8A via exon C (exon 30), and with RAB11A via the globular tail (Hales et al, 2001; Lapierre et al, 2001; Roland et al, 2007). MYO5B thereby functions as a dynamic tether for both RAB8A and RAB11A, and normally maintains these proteins at their appropriate subapical membrane localisation.
Loss of epithelial architecture and cell differentiation/polarity have been shown to be intimately linked to cancer (Royer and Lu, 2011). Accordingly, emerging evidence suggests that myosins have a crucial role and are differentially expressed in several cancer types (Ouderkirk and Krendel, 2014; Li and Yang, 2015). On the basis of its role in cell polarisation (Müller et al, 2008; Kravtsov et al, 2014), we hypothesised that MYO5B might be involved in CRC. Along this line, decreased expression of MYO5B was found in gastric cancer (Dong et al, 2013). In the present study, we examined the expression, methylation, and mutational status of MYO5B and its adapter proteins in primary colorectal tumours, precursor lesions, and distant normal tissue. We show that MYO5B expression is lost during tumour progression and that a gene expression signature composed of either MYO5B alone or in combination with its adapter RAB8A displays very strong prognostic value, thereby demonstrating its relevance as a potential prognostic marker in CRC.
Materials and methods
Patients and samples
All human tissue samples used in this study were donated freely, and written informed consent as well as ethical approval from the ‘Comité National d’Ethique de Recherche du Luxembourg’ (Reference 201009/09) and from the Institutional Ethics Review Panel (ERP-16-032) were obtained. Primary CRC and matched distant non-neoplastic colorectal tissue samples (at the furthest longitudinal surgical margin) were collected at the Hospital Emile Mayrisch by the Integrated Biobank of Luxembourg (IBBL, www.ibbl.lu) following the standard preanalytical protocol for biospecimens (Betsou et al, 2010). Samples were immediately stored in liquid nitrogen after surgical excision. Our CRC cohort was composed of 74 CRC patients and is described in Supplementary Table 1. The histopathological data was provided by a pathologist in a blinded way and further confirmed by an independent second pathologist from the ‘Laboratoire national de santé’. Out of the 74 samples analysed in this study, 16 were of rectum origin, of which 10 corresponding patients had received radiotherapy alone or in combination with chemotherapy before surgical removal of the tumour. The other remaining 58 patients presented a tumour of the colon, none of them had a treatment before tumour resection. As we did not detect any differences in MYO5B expression between colon- and rectum-derived tumours, we decided to assess them altogether as colorectal tumours. The cohort includes stage I (n=10), II (n=34), III (n=24), and IV (n=6) tumour samples, classified according to the TNM Classification of Malignant Tumours (TNM system, American Joint Committee on Cancer) (Hari et al, 2013) staging system, as well as 74 normal tissue samples matching the corresponding tumour (Supplementary Table 1). We complemented our CRC collection with 46 tumour specimens from the Ontario Tumour Bank (Ontario Institute for Cancer Research).
Materials
All CRC cell lines were obtained from the American Type Culture Collection (ATCC, Rockville, MD, USA) and maintained in recommended culture conditions. For methylation analysis, cells were treated with 5 μ m 5-Aza-2'-deoxycytidine (5-aza-dC/DAC, Sigma-Aldrich, Overijse, Belgium) or vehicle (DMSO) for 4 days followed by RNA extraction.
Tissue preparation and lysis, RNA/DNA extraction, and RT–qPCR
Tissue processing and laser microdissection (LMD) were performed as previously described (Letellier et al, 2014). AllPrep extraction kit (Qiagen, Venlo, Netherlands) was used to extract RNA and DNA from microdissected samples. cDNA was obtained via reverse transcription by using a high-capacity cDNA reverse transcription kit (Applied-Biosystems, Thermo-Fisher Scientific, Gent, Belgium). The Experion automated electrophoresis system (Bio-Rad Laboratories, Temse, Belgium) was used to check for RNA quality, which was of acceptable quality for all primary samples. The expression of MYO5B was investigated using TaqMan chemistry-based primer/probe sets that are recommended for the use of RNA from microdissected samples (Erickson et al, 2009) (Supplementary Table 2 for reference numbers). PCR conditions were used as previously described (Letellier et al, 2014) and the expression levels of the gene of interest were normalised against the housekeeping gene HPRT1 (Erickson et al, 2009). For bulk tissue and CRC cell lines, the RNA was extracted with the miRNeasy kit from Qiagen and cDNA was obtained with the miScript II (Qiagen). PCR cycling conditions as well as quality control and normalisation steps were done in qBase+ (Biogazelle, Gent, Belgium), using two reference genes as previously described (Qureshi-Baig et al, 2016; (Ullmann et al, 2016). Primer sequences are listed in Supplementary Table 2.
Tissue microarrays, immunohistochemical, and western blot analysis
First, the specificity of a selected MYO5B antibody (HPA040902, Sigma-Aldrich) was verified by a knockdown experiment. Briefly, HCT116 and HCT15 cells were transduced (MOI=5) with a pool of three target-specific short hairpin RNA constructs against MYO5B (sh MYO5B sc-75853-V, sh scramble sc-108080, Santa Cruz Biotechnology, Heidelberg, Germany), which resulted in a marked reduction in the expression of MYO5B in both cell lines on mRNA and protein levels (Supplementary Figure 1). For immunoblotting, proteins were subjected to SDS–PAGE (6%) and probed with anti-Stat3 (sc-482, Santa Cruz Biotechnology) as loading control as previously described (Ullmann et al (2016). Next, to mimic the condition of a tissue microarray (TMA), cells were clotted with a mix of thrombin/plasma and embedded in paraffin. Sections were cut and stained with the MYO5B antibody used in the TMA. The MYO5B signal intensity was highly reduced in cells after stable knockdown of MYO5B (Supplementary Figure 1).
The TMA blocks were prepared using 74 primary CRC formalin-fixed paraffin-embedded tumour samples as well as their paired normal colon counterparts. Two punches, 1 mm in diameter, were taken from each donor block providing two spots of both tumour and normal control tissue. The punching of tissue cores and their transfer to the receiver block were done using a 3D Histech TMA arrayer (Sysmex, Budapest, Hungari). Immunohistochemical staining was done on an automated Benchmark XT device (Ventana, Tucson, Arizona, USA) using the CC1M antigen retrieval protocol (Cell conditioning 1 buffer, basic pH and M=30 min). Primary antibody against MYO5B was used at a 1 : 50 dilution, with an incubation time of 32 min. The secondary antibody was retrieved from an UltraView DAB detection kit (Ventana). Tissue sections were analysed and a pathologist blindly scored MYO5B staining as follows: 0 (no signal), 1 (low signal), 2 (moderate signal) or 3 (strong signal). Two sections of cancerous as well as normal counterpart tissue were scored per patient and the mean value was considered.
Bioinformatic meta-analysis
We have set up a meta-analysis and used it as previously described (Letellier et al, 2014). Briefly, we integrated all the individual CEL files from selected data sets profiled on HG-U133 plus 2.0 (Affymetrix, Santa Clara, CA, USA), retrieved from GEO (GSE14333, GSE17538, GSE21510, GSE8671, GSE9254, GSE20916, GSE10714, GSE15960, GSE4183, and GSE10961) and corresponding to different studies (Sabates-Bellver et al, 2007; Galamb et al, 2008a, 2008b, 2010; LaPointe et al, 2008; Pantaleo et al, 2008; Jorissen et al, 2009; Skrzypczak et al, 2010; Smith et al, 2010; Tsukamoto et al, 2011) into one single global analysis covering expression data on 829 patients. The suitability of potential markers to discriminate between CRC and normal colorectal samples was assessed by ROC curves as previously described (Letellier et al, 2014).
Methylation analysis
For monitoring of the methylation pattern of the entire MYO5B promoter, methylation analysis using MassARRAY technology (Sequenom) was performed at Varionostics GmbH, Ulm, Germany as previously described (Letellier et al, 2014). To further investigate the methylation of MYO5B, we used the TCGA data set for CRC (COAD project) with the R package TCGAbiolinks (Colaprico et al, 2016). Among the 456 patients with expression data (‘Gene Expression Quantification’ as data.type), 295 patients were investigated for DNA methylation (with the platform ‘Illumina Human Methylation 450’). RNA expression and DNA methylation were joined according to the patient ID, tissue origin, tumour stage, and gene symbol.
Identification of somatic mutations
Data on somatic mutations were retrieved for 399 patients using TCGAbiolinks with the MUTECT pipeline (function: GDCquery_Maf (‘COAD’, pipelines=‘mutect’)). One patient had 5 mutations, one 4 mutations, two 2 mutations, and fourteen 1 mutation. Even if the COSMIC IDs are present in the TCGA data set, the COSMIC database (Forbes et al, 2017) contains additional somatic mutations. Version 81 of COSMIC was used to download mutations in MYO5B. Five additional somatic mutations are referenced for TCGA patients, however the predicted consequences are missing. In total, 33 somatic MYO5B mutations were found in a total of 20 patients. All mutations are displayed on the lolliplot (Figure 3D) and their potential implications highlighted in Supplementary Table 3. The evaluation of potential structural and/or functional effects is based on existing structures of MYO5A and MYO5B (PBD entry codes: 4LNZ, 4LX0, 1OE9, and 2DFS) (Coureux et al, 2003; Liu et al, 2006; Pylypenko et al, 2013; Velvarska and Niessing, 2013).
Survival analyses
Normalised gene expression values from three different microarray studies with the accession number GSE39582 (Marisa et al, 2013), GSE24551 (Agesen et al, 2012), and GSE2881 (Loboda et al, 2011), and containing clinical data on the outcome of CRC patients were retrieved from GEO using the R package GEOquery (v2.40 (Davis and Meltzer, 2007)). For gene symbols with multiple probe set assignments, we selected the probe set with the largest interquartile range per gene, as suggested in Shi and He (2016). The retained probe sets for the genes of interest were as follows: MYO5B, 225299_at; RAB8B, 222846_at; RAB10, 222981_s_at; RAB11A, 234998_at; RAB11B, 34478_at; CDC42, 208727_s_at; RAC1, 1567458_s_at; and RHOA, 1555814_a_at. RAB25 only had one probe set assignment. Survival curves were generated using the R package survival (v2.41-2 (Therneau, 2015)) and plotted with the R package survminer (v0.3.1 (Kassambara and Kosinski, 2017)). The continuous log2 expression was separated into two discrete categories (‘high’ and ‘low’), separated according to the median MYO5B expression value. Of note, the median value was assigned to the ‘high’ category for odd-numbered data sets.
Combined gene signatures
To combine several genes, the log2 expression of the six retained genes (MYO5B, RAB8A, RAB9A, RAB10, RAB11A, and RAB25) or nine retained genes (the latter six combined with CDC42, RAC1, and RHOA) was mean-centred and scaled using the scale() function to make them comparable. Then, for each of the 63 or 511 combinations, respectively, scaled expressions were combined following the PROGgene procedure (Goswami and Nakshatri, 2013). Briefly, the six or nine scaled values were averaged to obtain only one gene expression signature value for each patient. The gene signature values were divided into ‘high’ and ‘low’ groups using the median as described above. Hazard ratios were computed using the coxph() function and ranked by the inferred estimates. Differences between Kaplan–Meier curves were assessed using the surdiff() function and P-values are reported on each plot.
Data analysis
All analyses and plots were generated using the R environment (v3.3.3 (https://www.R-project.org/), the R package ggplot2 (v2.2.1 (Wickham, 2009)), and other packages from the tidyverse idiom (Wickham, 2017) as well as GraphPad Prism software, version 5 (GraphPad Software, Inc, La Jolla, CA, USA). We used the student t-test to compare expression levels between tumour and normal tissue. Kaplan–Meier plots were analysed using the Cox proportional hazards model via the survival R package. Unless otherwise stated, results are shown as mean±s.d. and P-values <0.05 are considered as statistically significant.
Results
A bioinformatic study identifies MYO5B as a potential marker for CRC
Recently, myosins have gained much interest in cancer research (Ouderkirk and Krendel, 2014; Li and Yang, 2015). Accordingly, we assessed the relevance of Myosin V family members as novel CRC markers. First, we used a previously established meta-analysis (Letellier et al, 2014) of several human CRC data sets (n=829 patients) to examine the gene expression of different members of the Myosin V family, namely MYO5A, MYO5B, and MYO5C, in CRC. We found that compared to samples obtained from normal colorectal mucosa, MYO5A levels were decreased in adenoma samples but did not vary significantly between cancerous and normal tissue (Figure 1A). Lan et al (2010) have previously reported that MYO5A expression is increased in metastatic CRC tissues. In that study, the epithelial–mesenchymal transition (EMT) inducer Snail acted as a transcriptional activator of MYO5A, positively affecting cell migration and, subsequently, metastasis dissemination. However, using our meta-analysis on 829 patients, we were not able to observe a significant alteration in MYO5A expression in metastatic vs non-metastatic tissues (Supplementary Figure 2A). Further studies are needed to shed light on the expression of MYO5A in CRC. Interestingly, MYO5B expression was reduced by ∼52% in tumour tissue compared to normal colorectal mucosa samples (Figure 1A). MYO5C showed a similar decrease in expression (Figure 1A). In addition, a detailed analysis of MYO5B expression during CRC progression suggested that MYO5B is downregulated in a stage-dependent manner (Figure 1B).
Bioinformatic analysis reveals the potential use of MYO5B as a diagnostic marker for CRC. (A) Dot plot showing the log2 FC values of the Myosin V family genes in adenoma (beige) and CRC (blue) samples compared to healthy colorectal mucosa samples (grey) in a meta-analysis of different CRC data sets, including 829 patients. Bar represents the mean expression intensity (***P<0.001, t-test corrected for multiple testing). (B) MYO5B expression according to the different TNM stages in the meta-analysis. Bar represents the mean expression intensity (*P<0.05, ***P<0.001, t-test corrected for multiple testing). (C) ROC curves with corresponding AUC values for MYO5A, MYO5B, and MYO5C when comparing CRC and healthy samples in the meta-analysis. Distribution of gene expression values for healthy and CRC samples are shown in the insets.
Next, we generated ROC curves to determine the predictive power of MYO5B expression levels to distinguish cancerous from normal tissue. We observed AUC values of 0.9697 and 0.9284 for MYO5B and MYO5C, respectively (Figure 1C). In contrast, and in accordance with the expression data in Figure 1A, the levels of MYO5A did not allow for a clear separation of CRC and normal colorectal mucosa samples (AUC=0.5334; Figure 1C). The distribution of MYO5B gene expression levels shows two distinct, almost non-overlapping, peaks, whereas a certain degree of overlap was still observable for the MYO5C peaks (Figure 1C, inset). In conclusion, its high AUC values as well as its well-separated distribution profile support the use of MYO5B as a diagnostic marker. We therefore decided to analyse the clinical relevance of MYO5B downregulation in CRC.
MYO5B is downregulated on mRNA and protein level in primary human CRC samples
To validate the findings of our bioinformatic analysis, we assessed the expression of MYO5B in snap-frozen human CRC samples from a distinct patient cohort. First, we analysed the gene expression of MYO5B in 50 matching CRC and non-tumour bulk tissue samples and found that MYO5B expression was significantly lower in the tumour samples (Figure 2A). To ensure that the expression of MYO5B was derived from epithelial cells and not from infiltrated immune or residing stromal cells, we decided to specifically select for a highly pure malignant or normal epithelial cell population by LMD. In a set of 21 laser-microdissected paired samples, which we had already analysed as bulk tissue (Figure 2A) and which was further complemented by a set of 46 CRC laser-microdissected tumour samples, we could observe that MYO5B expression was again significantly decreased in tumour samples, compared to normal colon tissue (Figure 2B). While examining the paired samples selected by LMD, we could observe a higher expression of MYO5B in adjacent non-tumour tissue than in the corresponding tumour samples in 20 out of 21 tested patients (Figure 2C). Interestingly, laser-microdissected samples allowed for a better separation between CRC and non-tumour samples, suggesting that tumour-microenvironment-derived cells interfere with signals emanating from epithelial cells. To further test the potential applicability of MYO5B as a marker in CRC, we used TMAs to determine its expression on the protein level. First, the specificity of the antibody was verified using cells in which the expression of MYO5B was reduced by short-hairpin-mediated knockdown (Supplementary Figure 1 and Materials and Methods section). In normal healthy colon epithelium, MYO5B staining is known to be very strong at the apical brush border of the colonic mucosa (Müller et al, 2008). Likewise, we observed a cytoplasmic signal for MYO5B in normal and tumourigenic epithelial cells and identified an increasing gradient ranging from the basal pole to the apex (Figure 2D, left panel). We then evaluated the staining intensity of MYO5B in 74 paired CRC and non-tumour samples and found that normal counterpart tissue shows a significantly stronger signal when compared to matching tumour samples (Figure 2D, right panel). To determine whether MYO5B expression correlates with disease progression, we divided our sample set into early (I and II) and late (III and IV) stage groups based on their TNM staging. Late-stage samples showed a dramatic loss of MYO5B expression when compared to both normal epithelial tissue and to samples from early-stage tumours (Figure 2E), confirming the gradual loss of MYO5B observed in the meta-analysis. As the loss of MYO5B is known to contribute to the disruption of cell polarity (Müller et al, 2008), we further investigated its expression in tumours with varying differentiation grades, that is, ranging from well- to low-differentiated tumours. Interestingly, we observed a stepwise reduction in signal intensity from well (histological grade I) to moderately (histological grade II) and poorly (histological grade III) differentiated adenocarcinomas (Figure 2F). Overall, our results show that MYO5B is stage-dependently downregulated and suggest that it might have a role in CRC progression. Future studies on large independent CRC patient cohorts will help proving the robustness of MYO5B as a prognostic marker in CRC.
Downregulation of MYO5B on mRNA and protein levels in human CRC. (A) MYO5B mRNA levels in bulk tissue of matched tumour (CRC, n=50) and non-tumour (normal, n=50) samples from CRC patients. Data are presented as mean±s.d. (***P<0.001, paired t-test). (B) MYO5B mRNA levels in laser-microdissected tumour (CRC, n=67) and non-tumour (normal, n=21) samples. Data are presented as mean±s.d. (***P<0.001, unpaired t-test). (C) In 20 out of 21 paired microdissected tumour samples, MYO5B mRNA levels were higher in non-tumour tissue compared to the matched tumour sample (***P<0.001, paired t-test). (D) Immunohistochemical staining of 74 paired TMAs of CRC patients. Left panel: representative image of MYO5B immunostaining on a section that contains normal (bottom) and cancerous tissue (top). Staining was mainly cytoplasmic and showed the expected accumulation of MYO5B at the apical brush border of normal enterocytes, as well as in tumoural cells in which differentiation was still preserved. Scale bar corresponds to 200 μm. Right panel: quantification of MYO5B staining of matched tumour (CRC; n=74) and non-tumour (normal; n=74) samples scored according to the staining intensity of MYO5B-positive cells (0=no signal, 1=mild signal, 2=moderate signal, and 3=strong signal). Two sections of cancerous as well as normal counterpart tissue were scored per patient and the mean value was considered. Data are presented as mean±s.d. (***P<0.001, paired t-test). (E) Representative pictures (left panel) and quantification (right panel) of immunohistochemical staining of MYO5B in patient samples of different TNM stages. Data are presented as mean±s.d. (**P<0.01, ***P<0.001, unpaired t-test). Scale bar corresponds to 200 μm. (F) MYO5B scoring of TMA samples after classification according to their histological grade. The information was missing for one patient. Data are presented as mean±s.d.; unpaired t-test. *P<0.05, **P<0.01, and ***P<0.001.
Loss of MYO5B expression in CRC is not due to promoter hypermethylation
Over the last years, hypermethylation of tumour suppressor genes has been reported in different cancer types (Biswas and Rao, 2017). In a similar manner, MYO5B is methylated in both gastric tumours (Dong et al, 2013) and leukaemias (Kuang et al, 2008). Thus, we decided to analyse whether promoter methylation could explain the observed downregulation of MYO5B in CRC. To this end, we first investigated the expression of MYO5B in CRC cell lines after treatment with DAC, a known demethylating agent. However, DAC treatment resulted in no or only minor changes compared to MYO5B basal expression levels (Figure 3A), suggesting that methylation of its promoter may not be responsible for the observed reduction in MYO5B mRNA and protein expression. However, as DAC unspecifically demethylates the entire DNA, indirect effects may have influenced MYO5B expression. Thus, we decided to investigate the methylation status of the CpG sites within the entire promoter sequence via mass array technology. The methylation profile analysis of eight CRC cell lines, eight CRC patients (S1–S8) as well as four matching normal non-tumour samples (SN1–SN4) thereby did not show any tumour-specific methylation (background signal is defined by a methylation signal of <5%), neither in CRC cell lines nor in primary human tumour samples (Figure 3B). We completed our study by analysing the methylation status of MYO5B in a larger CRC cohort using the TCGA database, which includes methylation data on 295 patients. Eight cytosines in the promoter were assessed for methylation, however and in agreement with our previous data (Figure 3A and B), we did not observe any difference in methylation between healthy and cancerous tissue (Figure 3C).
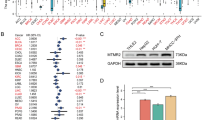
Frequently reduced expression of MYO5B is neither due to methylation nor mutations in CRC. (A) 5-aza-dC (DAC) treatment does not increase basal MYO5B expression in colon cancer cell lines. Data are representative of 2–4 independent experiments and presented as mean±s.d. (B) Methylation pattern analysed by mass array technology of eight CRC cell lines as well as eight tumour samples and four matching non-tumour counterparts. Each analysed cytosine is represented by a dot; cytosines with values below 5% were considered as non-methylated. (C) Analysis of MYO5B methylation in the TCGA data set covering methylation data on 295 tumour and 19 non-tumour samples. (D) MYO5B mutations in CRC patients. The protein domains and motifs are presented according to Qiu et al (2017), and the RAB8A- and RAB11A-binding sites according to Roland et al (2011). The dilute domain is also called globular tail domain. IQ=IQ motif, BD=binding domain.
MYO5B mutations in CRC patients
Another reason for reduced expression can be ascribed to the presence of mutations, which can lead to degradation at either RNA or protein level; MYO5B mutations have mostly been studied in MVID where inactivating mutations induce defects in the polarisation of cells leading to an intractable diarrhoea and malabsorption (Müller et al, 2008; van der Velde et al, 2013; Knowles et al, 2014; Qiu et al, 2017). Recently, a comprehensive human genetic study has further linked MYO5B mutations to 20% of previously undiagnosed low CGT cholestasis, an atypical form of MVID (Qiu et al, 2017). Most importantly, mutations in MYO5B have been identified in the malignant transformation of pheochromocytoma and paraganglioma (PCC/PGL) (Comino-Méndez et al, 2011; Fishbein et al, 2015; Wilzén et al, 2016), unravelling MYO5B as a putative player in PCC/PGL tumourigenesis and potential driver of malignancy. Four mutations in MYO5B have been identified: the first one is within the myosin head domain (also called motor domain) and is responsible for actin binding (p.L578P); the second is located in the coiled coil motif of MYO5B (p.V1261G); the two other mutations, p.R1641C and p.G1611S, are found within the globular tail domain of MYO5B, which is mediating RAB11A binding. Along this line, we have performed a mutational analysis of the MYO5B gene in CRC patients (Figure 3D). Supplementary Table 3 highlights the mutations identified in MYO5B among CRC patients and describes their potential functional implication: whereas a number of mutations are silent, some missense and nonsense mutations could clearly affect the stability and function of MYO5B. None of the identified mutations were common to the ones that are associated with the progression from benign to malignant PCC/PGL. Three identified nonsense mutations (p.X253_splice, p.Y287Lfs*17, and p.S608*) are located in the motor domain and may affect RNA expression. Similarily to the p.Gln1456X and c.4460-1G>C mutations that were described by Szperl et al (2011), these mutations may result in nonsense-mediated RNA decay leading to a reduction in the expression of MYO5B. A number of other mutations (Supplementary Table 3) have a high potential to affect the structure and stability of the protein or even specifically affect the function by interfering with RAB11A interaction. These mutations will generally not affect RNA expression. Overall, only a small subset of patients harbours such mutations (20 of 399 patients), which cannot explain the decreased expression of MYO5B in the majority of CRC patient samples. We therefore concluded that the widespread downregulation of MYO5B mRNA and protein expression in our patient collection is most likely neither due to MYO5B promoter methylation nor to inactivating mutations, suggesting that other factors, such as histone modifications, might be responsible for the generic loss of MYO5B expression among all CRC patients. We hope that future studies will allow to elucidate the mechanism underpinning MYO5B downregulation in CRC.
MYO5B expression has prognostic value in CRC
To evaluate whether MYO5B has prognostic value in CRC, we used publicly available gene expression data of 585 CRC patients (GSE39582) (Marisa et al, 2013). We confirmed the downregulation of MYO5B in this independent data set, which was not contained in our meta-analysis (data not shown). Next, patients were divided into two groups (‘high’ or ‘low’), based on their MYO5B expression, followed by Kaplan–Meier survival analysis. We detected a significant correlation between low MYO5B expression and shorter overall (Figure 4A) and relapse-free (Figure 4B) survival. Using the same data set GSE39582, we extended our analysis to include CRC staging data and found that in stage I and II (lymph node-negative cancer) patients, both overall (Figure 4C) and relapse-free (Figure 4D) survival time were associated with MYO5B expression. The strong increase in the statistical power of MYO5B to predict relapse-free survival in stage I/II patients compared to all stages confounded (P=0.003895 in stage I/II vs P=0.014 in stages I–IV altogether) highlights the clinical relevance of MYO5B expression in early CRC stages. Indeed, identifying patients at risk for recurrence during the early course of the disease might help clinicians to choose the best treatment plan. It is well known that stage II patients do not all benefit from chemotherapy and that markers able to successfully identify ‘high-risk’ patients are urgently needed. In addition, we further confirmed the prognostic value of MYO5B using a second independent patient cohort (GSE24551 (Sveen et al, 2011)) that included clinical data from 160 CRC patients (Figure 4E). Loss of MYO5B has been linked to invasion and motility in gastric cancer cells (Dong et al, 2012), which will ultimately lead to the dissemination of the disease by metastases spreading. Accordingly, we investigated in a data set that contained clinical information on metastasis from 125 CRC patients (GSE28814 (Loboda et al, 2011)) whether MYO5B expression could predict the occurrence of metastasis and, by extension, disease relapse. We found that a patient classification according to their MYO5B expression allows for the prediction of metastasis development (Figure 4F). In conclusion, our data suggest that MYO5B expression has a strong prognostic value in CRC patients.
MYO5B is a prognostic marker in CRC patients. (A and B) Overall (A), n=585 and relapse-free (B), n=574 survival in CRC patients stratified according to MYO5B expression in the GSE39582 data set. (C and D) Overall (C) and relapse-free (D) survival in early stage (stages I and II) CRC patients stratified according to MYO5B expression in the GSE39582 data set. (E) Disease-free survival curves for patients with ‘high’ and ‘low’ expression levels of MYO5B in an independent data set (GSE24551) covering 160 patients. (F) Metastasis-free survival based on ‘high’ or ‘low’ expression of MYO5B in a data set (GSE28814) containing clinical data on metastasis from 125 CRC patients. Patient numbers and associated P-values are indicated in the figures.
RAB family members RAB8A and RAB25 are downregulated in CRC
MYO5B is known to interact with different members of the RAB family, such as RAB8A, RAB9A, RAB10, RAB11A, and RAB25 (Fan et al, 2004; Ishikura and Klip, 2008; Roland et al, 2011) that are essential components of the vesicle trafficking machinery (Kelly et al, 2012a; Zhen and Stenmark, 2015). Several of these proteins have recently been identified to exhibit a tumour suppressor role in cancer (Nam et al, 2010; Mitra et al, 2012; Tzeng and Wang, 2016). To select the most relevant RAB family members in CRC, we analysed the correlation between the expression of MYO5B and different RAB family members by performing linear regression analyses using our meta-analysis. Five RAB genes, namely RAB8A, RAB9A, RAB10, RAB11A, and RAB25, which are known to interact with MYO5B and for which the expression correlated with that of MYO5B, were selected for further expression analysis (Supplementary Figure 2B and data not shown). Interestingly, all examined RAB family members were significantly downregulated in CRC (Figure 5A). Furthermore, RAB8A and RAB9A levels were far lower in adenoma samples than in normal healthy samples (Figure 5A). This finding may potentially be used as an indicator for pre-malignant tumours, as RAB8A and RAB9A expression is further reduced over the course of tumour progression (Figure 5A). Following this, we aimed at determining the suitability of our examined RAB members as potential prognostic markers for CRC. The results obtained were further strengthened by our correlation data, as RAB8A and RAB25 showed the highest AUC values, indicating their suitability as CRC markers (Figure 5B and Supplementary Figure 2C). Finally, we assessed the expression of these adapter molecules in CRC and found that RAB8A and RAB25 were significantly downregulated in our paired tumour/non-tumour data set (Figure 5C). We may hypothesise that the expression of MYO5B and its adapters, RAB8A and RAB25, is reduced in CRC, potentially contributing to the loss of polarity in epithelial cells and ultimately leading to tumour invasion and disease progression. Interestingly, RAB25 is methylated in two CRC cell lines (SW480 and SW620) and in patients (TCGA dataset, n=295, Supplementary Figure 3), which is in line with the epigenetic silencing of RAB25 observed in other cancer types (Wrzeszczynski et al, 2011; Clausen et al, 2016) and the poor prognosis that is associated with low RAB25 expression in CRC (Nam et al, 2010; Goldenring and Nam, 2011).
RAB family members RAB8A and RAB25 are downregulated in CRC. (A) Dot plot showing the log2 expression intensities of different RAB genes in adenoma (beige) and CRC (blue) samples compared to unmatched healthy colorectal mucosa (grey) samples in a meta-analysis of different CRC data sets, including 829 patients. Bar represents the mean expression intensity (*P<0.05, ***P<0.001, t-test corrected for multiple testing). (B) ROC curves with corresponding AUC values for RAB genes when classifying CRC and healthy patients in the meta-analysis. Distribution of gene expression values for normal and CRC samples are shown in the insets. (C) RAB8A and RAB25 expression levels in bulk tissue of matched tumour (CRC, n=50) and non-tumour (normal, n=50) samples from CRC patients. Data are presented as mean±s.d.; *P<0.05, ***P<0.001, paired t-test.
The combinatorial signature of MYO5B with its adapter RAB8A has strong prognostic value in CRC
As we identified a correlation between the expression of certain adapters and MYO5B, we hypothesised that a combinatorial signature with one or more adapters might further increase the prognostic power, compared to MYO5B alone. To analyse this further, we tested a multitude of different combinatorial signatures (n=63), carrying out our analysis on the GSE39582 data set (n=585 CRC patients). Strikingly, among the 63 different possibilities, the combination of the expression of MYO5B and its adapter RAB8A showed the highest response (Figure 6A) and markedly increased the prognostic power compared to MYO5B alone, both on overall and on relapse-free survival (Figure 6B). As MYO5B is also associated with the Rac/Rho pathway (Wilzén et al, 2016), we analysed whether a combinatorial signature of MYO5B together with CDC42, RAC1, or RHOA could outperform the single gene expression of MYO5B alone or the combinatorial signature, including MYO5B and RAB8A as well as the other RAB members. While the combinatorial signature of RAB8A, RAB11A, and RHOA outperformed the prognostic value of MYO5B expression alone, the combination of MYO5B together with the small GTPase RAB8A demonstrated the best prognostic value for relapse-free survival (Figure 6 and Supplementary Figure 4B).
The combination of MYO5B expression with its adapter protein RAB8A improves the prognostic power of MYO5B in CRC. (A) Top 20 significant combinatorial signatures between MYO5B and different genes of the RAB family for overall (OS, left) and relapse-free (RFS, right) survival in 585 CRC patients of the GSE39582 data set. Because of space issues, MYO as well as RAB have been omitted from the name description and only the last letters are shown. (B) Overall (left) and relapse-free (right) survival in 585 CRC patients stratified according to their combined RAB8A and MYO5B expression signature in the GSE39582 data set. The dotted line represents the MYO5B signature alone, whereas continuous lines represent the combinatorial signature. Patient numbers as well as their P-values are indicated in the figure.
Discussion
There is a lack of reliable prognostic markers in the current treatment of CRC. Up to now, the prognosis of CRC patients highly depends on tumour staging, which is defined by the degree of tumour penetration through the intestinal wall and the presence of distant metastasis. Nevertheless, patients with an identical TNM stage often differ in terms of treatment response and survival outcome. Consequently, a better classification of the patients is urgently needed to advise clinicians in choosing the best treatment options ultimately leading to increased patient survival. Recently, a combined effort of several different groups has led to a consensus on CRC classification, resulting in the identification of four distinct molecular CRC subtypes with clear biological interpretability (Guinney et al, 2015). However, this classification is based on large-scale gene expression analysis and could therefore potentially be difficult to translate into clinical use due to cost issues as well as limitations in sufficient quantity and quality from formalin-fixed paraffin-embedded tissue. Thus, it is vital to identify markers that can easily be used in clinics and that help to stratify patients with different prognosis, thereby identifying ‘high-risk’ patients. In this study, we examined the expression of Myosin V family members in CRC; MYO5B mRNA and protein expression is reduced in CRC samples and loss of its expression correlates with TNM stage and tumour differentiation grade. Most importantly, we were able to show that MYO5B downregulation, alone or in combination with a lower expression of its adapter RAB8A, is an independent predictor of shorter overall and relapse-free survival, which may guide oncologists in choosing the best available treatment option for a given patient. In a recent study, the Vermeulen group has used paraffin-embedded primary CRC tissues to classify CRC patients based on the expression of five immunohistochemical markers (CDX2, FRMD6, HTR2B, ZEB1, and KER) and showed an 87% concordance with the transcriptome-based four molecular subtype classification of CRC (Trinh et al, 2016). Keeping this in mind, it would be interesting to see whether analysing MYO5B expression could further improve the robustness of immunohistochemical assays.
Increasing evidence suggests that myosins have multiple crucial roles during tumourigenesis (Ouderkirk and Krendel, 2014; Li and Yang, 2015). Nevertheless, MYO5B, a member of the non-conventional myosins, remains poorly studied in the context of cancer. Dong and colleagues have shown that inactivation of MYO5B in gastric cancer leads to c-Met signalling, thereby enhancing tumour invasion and progression. In addition, they reported a reduced expression of MYO5B and found its promoter to be hypermethylated in 93% of gastric carcinoma samples (Dong et al, 2012, 2013). In our study, mass array analysis, a highly quantitative method, was used to assess the methylation pattern of all cytosines within the MYO5B promoter. We could not detect any significant methylation, neither in the CRC cell lines nor in the analysed CRC tissue samples, nor could we find any evidence for significant methylation in the TCGA data set. Thus, alternative mechanisms other than promoter methylation seem to be responsible for the downregulation of MYO5B in CRC. Interestingly, MYO1A has been shown to be inactivated by genetic mutations in CRC, though restricted to MSI-positive patients (Mazzolini et al, 2012). Similarly, mutations in MYO5B were exclusively found in the malignant form of PCC/PGL (Wilzén et al, 2016). In our study however, as most CRC patients do not harbour MYO5B mutations, inactivating mutations only allow to explain low expression of MYO5B in a small subset of CRC patients, suggesting that other factors, such as histone modifications, might be responsible for the loss of MYO5B expression in cancer.
During tumour progression, the structural proteins of the intestinal brush border lose their expression (Chantret et al, 1988; West et al, 1988), which correlates with the loss of epithelial architecture and cell differentiation/polarity. Accordingly, EMT, a pre-requisite to tumour invasion and metastasis dissemination, is characterised by the downregulation of epithelial differentiation markers and the loss of cell polarity and tissue organisation (Nieto et al, 2016). In agreement with this, we could show that MYO5B expression is decreased in tumours during progression of the disease (Figure 2E) and upon loss of tissue organisation (Figure 2F). In addition, our own unpublished data suggest that the induction of EMT correlates with the loss of MYO5B expression, a further possible explanation for the downregulation of MYO5B in CRC. As EMT correlates with the acquisition of cells with stem cell properties (Mani et al, 2008), we may further hypothesise that loss of MYO5B expression is involved in tumour initiation and progression by maintaining cells in an undifferentiated stem-cell-like state. Accordingly, we observed that the differentiation grade of the tumour was dependent on MYO5B expression. Along similar lines, a recent study by Mazzolini et al (2012) uncovered MYO1A to be essential for the polarisation and differentiation of tumourigenic colon cells, demonstrating its tumour suppressor activity in the intestine.
Studies on MVID have demonstrated that MYO5B regulates global intestinal enterocyte polarity (Müller et al, 2008), which is dependent on RAB8A and RAB11A (Knowles et al, 2014). Interestingly, Rab GTPases have host protective roles (Kelly et al, 2012a), including the establishment of endothelial and epithelial barrier integrity (Kelly et al, 2012b), and bacterial and viral pathogens often hijack the membrane trafficking machinery to facilitate their replication within host cells (Guichard et al, 2014). One may therefore speculate that the loss of junctional integrity in enterocytes, which is dependent on MYO5B (Knowles et al, 2014), might lead to pathogen infection and to chronic inflammation, one of the main causes of CRC (Grivennikov et al, 2010). Strikingly, we observed a reduced expression of MYO5B in inflammatory bowel disease (P=1.66 × 10−3, own unpublished observation), a chronic inflammatory condition with major defects in epithelial barrier integrity that is associated with an increased risk of CRC (Neurath, 2014). Future studies will focus on unravelling the mechanism by which loss of MYO5B leads to tumour progression. While the precise molecular mechanisms remain ellusive, our data clearly identify MYO5B, alone or together with its adapter protein RAB8A, as a strong prognostic marker in CRC.
Change history
21 November 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Agesen TH, Sveen A, Merok MA, Lind GE, Nesbakken A, Skotheim RI, Lothe RA (2012) ColoGuideEx: a robust gene classifier specific for stage II colorectal cancer prognosis. Gut 61: 1560–1567.
Betsou F, Lehmann S, Ashton G, Barnes M, Benson EE, Coppola D, DeSouza Y, Eliason J, Glazer B, Guadagni F, Harding K, Horsfall DJ, Kleeberger C, Nanni U, Prasad A, Shea K, Skubitz A, Somiari S, Gunter E (2010) Standard preanalytical coding for biospecimens: defining the sample PREanalytical code. Cancer Epidemiol Biomarkers Prev 19: 1004–1011.
Biswas S, Rao CM (2017) Epigenetics in cancer: fundamentals and beyond. Pharmacol Ther 173: 118–134.
Chantret I, Barbat A, Dussaulx E, Brattain MG, Zweibaum A (1988) Epithelial polarity, villin expression, and enterocytic differentiation of cultured human colon carcinoma cells: a survey of twenty cell lines. Cancer Res 48: 1936–1942.
Chavrier P, Vingron M, Sander C, Simons K, Zerial M (1990) Molecular cloning of YPT1/SEC4-related cDNAs from an epithelial cell line. Mol Cell Biol 10: 6578–6585.
Clausen MJAM, Melchers LJ, Mastik MF, Slagter-Menkema L, Groen HJM, van der Laan BFAM, van Criekinge W, de Meyer T, Denil S, van der Vegt B, Wisman GBA, Roodenburg JLN, Schuuring E (2016) RAB25 expression is epigenetically downregulated in oral and oropharyngeal squamous cell carcinoma with lymph node metastasis. Epigenetics 11: 653–663.
Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D, Sabedot TS, Malta TM, Pagnotta SM, Castiglioni I, Ceccarelli M, Bontempi G, Noushmehr H (2016) TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res 44: e71.
Comino-Méndez I, Gracia-Aznárez FJ, Schiavi F, Landa I, Leandro-García LJ, Letón R, Honrado E, Ramos-Medina R, Caronia D, Pita G, Gómez-Graña A, de Cubas AA, Inglada-Pérez L, Maliszewska A, Taschin E, Bobisse S, Pica G, Loli P, Hernández-Lavado R, Díaz JA, Gómez-Morales M, González-Neira A, Roncador G, Rodríguez-Antona C, Benítez J, Mannelli M, Opocher G, Robledo M, Cascón A (2011) Exome sequencing identifies MAX mutations as a cause of hereditary pheochromocytoma. Nat Genet 43: 663–667.
Coureux P-D, Wells AL, Ménétrey J, Yengo CM, Morris CA, Sweeney HL, Houdusse A (2003) A structural state of the myosin V motor without bound nucleotide. Nature 425: 419–423.
Davis S, Meltzer PS (2007) GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23: 1846–1847.
Dong W, Chen X, Chen P, Yue D, Zhu L, Fan Q (2012) Inactivation of MYO5B promotes invasion and motility in gastric cancer cells. Dig Dis Sci 57: 1247–1252.
Dong W, Wang L, Shen R (2013) MYO5B is epigenetically silenced and associated with MET signaling in human gastric cancer. Dig Dis Sci 58: 2038–2045.
Erickson HS, Albert PS, Gillespie JW, Rodriguez-Canales J, Marston Linehan W, Pinto PA, Chuaqui RF, Emmert-Buck MR (2009) Quantitative RT-PCR gene expression analysis of laser microdissected tissue samples. Nat Protoc 4: 902–922.
Fan G-H, Lapierre LA, Goldenring JR, Sai J, Richmond A (2004) Rab11-family interacting protein 2 and myosin Vb are required for CXCR2 recycling and receptor-mediated chemotaxis. Mol Biol Cell 15: 2456–2469.
Fishbein L, Khare S, Wubbenhorst B, DeSloover D, D’Andrea K, Merrill S, Cho NW, Greenberg RA, Else T, Montone K, LiVolsi V, Fraker D, Daber R, Cohen DL, Nathanson KL (2015) Whole-exome sequencing identifies somatic ATRX mutations in pheochromocytomas and paragangliomas. Nat Commun 6: 6140.
Forbes SA, Beare D, Boutselakis H, Bamford S, Bindal N, Tate J, Cole CG, Ward S, Dawson E, Ponting L, Stefancsik R, Harsha B, Kok CY, Jia M, Jubb H, Sondka Z, Thompson S, De T, Campbell PJ (2017) COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res 45: D777–D783.
Galamb O, Györffy B, Sipos F, Spisák S, Németh AM, Miheller P, Tulassay Z, Dinya E, Molnár B (2008a) Inflammation, adenoma and cancer: objective classification of colon biopsy specimens with gene expression signature. Dis Markers 25: 1–16.
Galamb O, Sipos F, Solymosi N, Spisák S, Krenács T, Tóth K, Tulassay Z, Molnár B (2008b) Diagnostic mRNA expression patterns of inflamed, benign, and malignant colorectal biopsy specimen and their correlation with peripheral blood results. Cancer Epidemiol Biomarkers Prev 17: 2835–2845.
Galamb O, Spisák S, Sipos F, Tóth K, Solymosi N, Wichmann B, Krenács T, Valcz G, Tulassay Z, Molnár B (2010) Reversal of gene expression changes in the colorectal normal-adenoma pathway by NS398 selective COX2 inhibitor. Br J Cancer 102: 765–773.
Goldenring JR, Nam KT (2011) Rab25 as a tumour suppressor in colon carcinogenesis. Br J Cancer 104: 33–36.
Goldenring JR, Shen KR, Vaughan HD, Modlin IM (1993) Identification of a small GTP-binding protein, Rab25, expressed in the gastrointestinal mucosa, kidney, and lung. J Biol Chem 268: 18419–18422.
Goldenring JR, Soroka CJ, Shen KR, Tang LH, Rodriguez W, Vaughan HD, Stoch SA, Modlin IM (1994) Enrichment of rab11, a small GTP-binding protein, in gastric parietal cells. Am J Physiol 267: G187–94.
Goswami CP, Nakshatri H (2013) PROGgene: gene expression based survival analysis web application for multiple cancers. J Clin Bioinformatics 3: 22.
Grivennikov SI, Greten FR, Karin M (2010) Immunity, inflammation, and cancer. Cell 140: 883–899.
Guichard A, Nizet V, Bier E (2014) RAB11-mediated trafficking in host-pathogen interactions. Nat Rev Microbiol 12: 624–634.
Guinney J, Dienstmann R, Wang X, de Reyniès A, Schlicker A, Soneson C, Marisa L, Roepman P, Nyamundanda G, Angelino P, Bot BM, Morris JS, Simon IM, Gerster S, Fessler E, De Sousa E Melo F, Missiaglia E, Ramay H, Barras D, Homicsko K, Maru D, Manyam GC, Broom B, Boige V, Perez-Villamil B, Laderas T, Salazar R, Gray JW, Hanahan D, Tabernero J, Bernards R, Friend SH, Laurent-Puig P, Medema JP, Sadanandam A, Wessels L, Delorenzi M, Kopetz S, Vermeulen L, Tejpar S (2015) The consensus molecular subtypes of colorectal cancer. Nat Med 21: 1350–1356.
Hales CM, Griner R, Hobdy-Henderson KC, Dorn MC, Hardy D, Kumar R, Navarre J, Chan EKL, Lapierre LA, Goldenring JR (2001) Identification and characterization of a family of Rab11-interacting proteins. J Biol Chem 276: 39067–39075.
Hari DM, Leung AM, Lee J-H, Sim M-S, Vuong B, Chiu CG, Bilchik AJ (2013) AJCC Cancer Staging Manual 7th edition criteria for colon cancer: do the complex modifications improve prognostic assessment? J Am Coll Surg 217: 181–190.
Ishikura S, Klip A (2008) Muscle cells engage Rab8A and myosin Vb in insulin-dependent GLUT4 translocation. Am J Physiol Cell Physiol 295: C1016–25.
Jorissen RN, Gibbs P, Christie M, Prakash S, Lipton L, Desai J, Kerr D, Aaltonen LA, Arango D, Kruhøffer M, Orntoft TF, Andersen CL, Gruidl M, Kamath VP, Eschrich S, Yeatman TJ, Sieber OM (2009) Metastasis-associated gene expression changes predict poor outcomes in patients with Dukes stage B and C colorectal cancer. Clin Cancer Res 15: 7642–7651.
Kassambara A, Kosinski M (2017) survminer: Drawing Survival Curves using 'ggplot2'. Available at: https://CRAN.R-project.org/package=survminer (accessed on February 2017).
Kelly EE, Horgan CP, Goud B, McCaffrey MW (2012a) The Rab family of proteins: 25 years on. Biochem Soc Trans 40: 1337–1347.
Kelly EE, Horgan CP, McCaffrey MW (2012b) Rab11 proteins in health and disease. Biochem Soc Trans 40: 1360–1367.
Knowles BC, Roland JT, Krishnan M, Tyska MJ, Lapierre LA, Dickman PS, Goldenring JR, Shub MD (2014) Myosin Vb uncoupling from RAB8A and RAB11A elicits microvillus inclusion disease. J Clin Invest 124: 2947–2962.
Kravtsov D, Mashukova A, Forteza R, Rodriguez MM, Ameen NA, Salas PJ (2014) Myosin 5b loss of function leads to defects in polarized signaling: implication for microvillus inclusion disease pathogenesis and treatment. Am J Physiol Gastrointest Liver Physiol 307: G992–G1001.
Kuang S-Q, Tong W-G, Yang H, Lin W, Lee MK, Fang ZH, Wei Y, Jelinek J, Issa J-P, Garcia-Manero G (2008) Genome-wide identification of aberrantly methylated promoter associated CpG islands in acute lymphocytic leukemia. Leukemia 22: 1529–1538.
Lai F, Stubbs L, Artzt K (1994) Molecular analysis of mouse Rab11b: a new type of mammalian YPT/Rab protein. Genomics 22: 610–616.
Lan L, Han H, Zuo H, Chen Z, Du Y, Zhao W, Gu J, Zhang Z (2010) Upregulation of myosin Va by Snail is involved in cancer cell migration and metastasis. Int J Cancer 126: 53–64.
Lapierre LA, Kumar R, Hales CM, Navarre J, Bhartur SG, Burnette JO, Provance DW, Mercer JA, Bähler M, Goldenring JR (2001) Myosin vb is associated with plasma membrane recycling systems. Mol Biol Cell 12: 1843–1857.
LaPointe LC, Dunne R, Brown GS, Worthley DL, Molloy PL, Wattchow D, Young GP (2008) Map of differential transcript expression in the normal human large intestine. Physiol Genomics 33: 50–64.
Letellier E, Schmitz M, Baig K, Beaume N, Schwartz C, Frasquilho S, Antunes L, Marcon N, Nazarov PV, Vallar L, Even J, Haan S (2014) Identification of SOCS2 and SOCS6 as biomarkers in human colorectal cancer. Br J Cancer 111: 726–735.
Li Y-R, Yang W-X (2015) Myosins as fundamental components during tumourigenesis: diverse and indispensable. Oncotarget 7: 46785–46812.
Liu J, Taylor DW, Krementsova EB, Trybus KM, Taylor KA (2006) Three-dimensional structure of the myosin V inhibited state by cryoelectron tomography. Nature 442: 208–211.
Loboda A, Nebozhyn MV, Watters JW, Buser CA, Shaw PM, Huang PS, Van’t Veer L, Tollenaar RAEM, Jackson DB, Agrawal D, Dai H, Yeatman TJ (2011) EMT is the dominant program in human colon cancer. BMC Med Genomics 4: 9.
Mani SA, Guo W, Liao M-J, Eaton EN, Ayyanan A, Zhou AY, Brooks M, Reinhard F, Zhang CC, Shipitsin M, Campbell LL, Polyak K, Brisken C, Yang J, Weinberg RA (2008) The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133: 704–715.
Marisa L, de Reyniès A, Duval A, Selves J, Gaub MP, Vescovo L, Etienne-Grimaldi M-C, Schiappa R, Guenot D, Ayadi M, Kirzin S, Chazal M, Fléjou J-F, Benchimol D, Berger A, Lagarde A, Pencreach E, Piard F, Elias D, Parc Y, Olschwang S, Milano G, Laurent-Puig P, Boige V (2013) Gene expression classification of colon cancer into molecular subtypes: characterization, validation, and prognostic value. PLoS Med 10: e1001453.
Mazzolini R, Dopeso H, Mateo-Lozano S, Chang W, Rodrigues P, Bazzocco S, Alazzouzi H, Landolfi S, Hernandez-Losa J, Andretta E, Alhopuro P, Espin E, Armengol M, Tabernero J, Ramon y Cajal S, Kloor M, Gebert J, Mariadason JM, Schwartz S, Aaltonen LA, Mooseker MS, Arango D (2012) Brush border Myosin Ia has tumor suppressor activity in the intestine. Proc Natl Acad Sci USA 109: 1530–1535.
Mitra S, Cheng KW, Mills GB (2012) Rab25 in cancer: a brief update. Biochem Soc Trans 40: 1404–1408.
Müller T, Hess MW, Schiefermeier N, Pfaller K, Ebner HL, Heinz-Erian P, Ponstingl H, Partsch J, Röllinghoff B, Köhler H, Berger T, Lenhartz H, Schlenck B, Houwen RJ, Taylor CJ, Zoller H, Lechner S, Goulet O, Utermann G, Ruemmele FM, Huber LA, Janecke AR (2008) MYO5B mutations cause microvillus inclusion disease and disrupt epithelial cell polarity. Nat Genet 40: 1163–1165.
Nam KT, Lee H-J, Smith JJ, Lapierre LA, Kamath VP, Chen X, Aronow BJ, Yeatman TJ, Bhartur SG, Calhoun BC, Condie B, Manley NR, Beauchamp RD, Coffey RJ, Goldenring JR (2010) Loss of Rab25 promotes the development of intestinal neoplasia in mice and is associated with human colorectal adenocarcinomas. J Clin Invest 120: 840–849.
Neurath MF (2014) Cytokines in inflammatory bowel disease. Nat Rev Immunol 14: 329–342.
Nieto MA, Huang RY-J, Jackson RA, Thiery JP (2016) EMT: 2016. Cell 166: 21–45.
Ouderkirk JL, Krendel M (2014) Non-muscle myosins in tumor progression, cancer cell invasion, and metastasis. Cytoskeleton 71: 447–463.
Pantaleo MA, Astolfi A, Nannini M, Paterini P, Piazzi G, Ercolani G, Brandi G, Martinelli G, Pession A, Pinna AD, Biasco G (2008) Gene expression profiling of liver metastases from colorectal cancer as potential basis for treatment choice. Br J Cancer 99: 1729–1734.
Pylypenko O, Attanda W, Gauquelin C, Lahmani M, Coulibaly D, Baron B, Hoos S, Titus MA, England P, Houdusse AM (2013) Structural basis of myosin V Rab GTPase-dependent cargo recognition. Proc Natl Acad Sci USA 110: 20443–20448.
Qiu Y-L, Gong J-Y, Feng J-Y, Wang R-X, Han J, Liu T, Lu Y, Li L-T, Zhang M-H, Sheps JA, Wang N-L, Yan Y-Y, Li J-Q, Chen L, Borchers CH, Sipos B, Knisely AS, Ling V, Xing Q-H, Wang J-S (2017) Defects in myosin VB are associated with a spectrum of previously undiagnosed low γ-glutamyltransferase cholestasis. Hepatology 65: 1655–1669.
Qureshi-Baig K, Ullmann P, Rodriguez F, Frasquilho S, Nazarov PV, Haan S, Letellier E (2016) What do we learn from spheroid culture systems? Insights from tumorspheres derived from primary colon cancer tissue. PLoS One 11: e0146052.
Roland JT, Bryant DM, Datta A, Itzen A, Mostov KE, Goldenring JR (2011) Rab GTPase-Myo5B complexes control membrane recycling and epithelial polarization. Proc Natl Acad Sci USA 108: 2789–2794.
Roland JT, Kenworthy AK, Peranen J, Caplan S, Goldenring JR (2007) Myosin Vb interacts with Rab8a on a tubular network containing EHD1 and EHD3. Mol Biol Cell 18: 2828–2837.
Royer C, Lu X (2011) Epithelial cell polarity: a major gatekeeper against cancer? Cell Death Differ 18: 1470–1477.
Sabates-Bellver J, Van der Flier LG, de Palo M, Cattaneo E, Maake C, Rehrauer H, Laczko E, Kurowski MA, Bujnicki JM, Menigatti M, Luz J, Ranalli TV, Gomes V, Pastorelli A, Faggiani R, Anti M, Jiricny J, Clevers H, Marra G (2007) Transcriptome profile of human colorectal adenomas. Mol Cancer Res 5: 1263–1275.
Shi M, He J (2016) ColoFinder: a prognostic 9-gene signature improves prognosis for 871 stage II and III colorectal cancer patients. PeerJ 4: e1804.
Skrzypczak M, Goryca K, Rubel T, Paziewska A, Mikula M, Jarosz D, Pachlewski J, Oledzki J, Ostrowski J, Ostrowsk J (2010) Modeling oncogenic signaling in colon tumors by multidirectional analyses of microarray data directed for maximization of analytical reliability. PLoS One 5: e13091.
Smith JJ, Deane NG, Wu F, Merchant NB, Zhang B, Jiang A, Lu P, Johnson JC, Schmidt C, Bailey CE, Eschrich S, Kis C, Levy S, Washington MK, Heslin MJ, Coffey RJ, Yeatman TJ, Shyr Y, Beauchamp RD (2010) Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology 138: 958–968.
Sveen A, Agesen TH, Nesbakken A, Rognum TO, Lothe RA, Skotheim RI (2011) Transcriptome instability in colorectal cancer identified by exon microarray analyses: associations with splicing factor expression levels and patient survival. Genome Med 3: 32.
Szperl AM, Golachowska MR, Bruinenberg M, Prekeris R, Thunnissen A-MWH, Karrenbeld A, Dijkstra G, Hoekstra D, Mercer D, Ksiazyk J, Wijmenga C, Wapenaar MC, Rings EHHM, van IJzendoorn SCD (2011) Functional characterization of mutations in the myosin Vb gene associated with microvillus inclusion disease. J Pediatr Gastroenterol Nutr 52: 307–313.
Therneau T (2015) A Package for Survival Analysis in S. version 2.38. Available at: https://CRAN.R-project.org/package=survival (accessed on February 2017).
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA Cancer J Clin 65: 87–108.
Trinh A, Trumpi K, De Sousa E, Melo F, Wang X, de Jong JH, Fessler E, Kuppen PJK, Reimers MS, Swets M, Koopman M, Nagtegaal ID, Jansen M, Hooijer GKJ, Offerhaus GJA, Kranenburg O, Punt CJ, Medema JP, Markowetz F, Vermeulen L (2016) Practical and robust identification of molecular subtypes in colorectal cancer by immunohistochemistry. Clin Cancer Res 23: 387–398.
Trybus KM (2008) Myosin V from head to tail. Cell Mol Life Sci 65: 1378–1389.
Tsukamoto S, Ishikawa T, Iida S, Ishiguro M, Mogushi K, Mizushima H, Uetake H, Tanaka H, Sugihara K (2011) Clinical significance of osteoprotegerin expression in human colorectal cancer. Clin Cancer Res 17: 2444–2450.
Tzeng H-T, Wang Y-C (2016) Rab-mediated vesicle trafficking in cancer. J Biomed Sci 23: 70.
Ullmann P, Qureshi-Baig K, Rodriguez F, Ginolhac A, Nonnenmacher Y, Ternes D, Weiler J, Gäbler K, Bahlawane C, Hiller K, Haan S, Letellier E (2016) Hypoxia-responsive miR-210 promotes self-renewal capacity of colon tumor-initiating cells by repressing ISCU and by inducing lactate production. Oncotarget 7: 65454–65470.
van der Velde KJ, Dhekne HS, Swertz MA, Sirigu S, Ropars V, Vinke PC, Rengaw T, van den Akker PC, Rings EHHM, Houdusse A, van Ijzendoorn SCD (2013) An overview and online registry of microvillus inclusion disease patients and their MYO5B mutations. Hum Mutat 34: 1597–1605.
Varrette S, Bouvry P, Cartiaux H, Georgatos F (2014) Management of an academic HPC cluster: The UL experience. Paper presented at 2014 International Conference on High Performance Computing & Simulation (HPCS).
Velvarska H, Niessing D (2013) Structural insights into the globular tails of the human type v myosins Myo5a, Myo5b, And Myo5c. PLoS One 8: e82065.
West AB, Isaac CA, Carboni JM, Morrow JS, Mooseker MS, Barwick KW (1988) Localization of villin, a cytoskeletal protein specific to microvilli, in human ileum and colon and in colonic neoplasms. Gastroenterology 94: 343–352.
Wickham H (2009) ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag: New York, NY, USA, (accessed on February 2017).
Wickham H (2017) tidyverse: Easily Install and Load 'Tidyverse' Packages. R package version 1.1.1. Available at: https://CRAN.R-project.org/package=tidyverse (accessed on February 2017).
Wilzén A, Rehammar A, Muth A, Nilsson O, Tešan Tomić T, Wängberg B, Kristiansson E, Abel F (2016) Malignant pheochromocytomas/paragangliomas harbor mutations in transport and cell adhesion genes. Int J Cancer 138: 2201–2211.
Wrzeszczynski KO, Varadan V, Byrnes J, Lum E, Kamalakaran S, Levine DA, Dimitrova N, Zhang MQ, Lucito R (2011) Identification of tumor suppressors and oncogenes from genomic and epigenetic features in ovarian cancer. PLoS ONE 6: e28503.
Zhen Y, Stenmark H (2015) Cellular functions of Rab GTPases at a glance. J Cell Sci 128: 3171–3176.
Acknowledgements
We thank the Fondation Cancer for funding this study and the Fonds National de la Recherche (FNR) for its support of KQB and PU under the AFR grant scheme. We are grateful to the Fondation Pélican for its support of KQB and PU. We thank the contributing surgeons, in particular Dr Nikolaus Zügel and Dr Marc Boulmont, from the Centre Hospitalier Emile Mayrisch in Esch-sur-Alzette, and the nurses of the Clinical and Epidemiological Investigation Center of the Luxembourg Institute of Health, for their work with the patients. We thank Dr Nikolai Goncharenko, Dr Christelle Bahlawane, and Dr Fay Betsou from the IBBL for their assistance in establishing a sample collection for the study. Some of the computational work was carried out using the HPC facilities of the University of Luxembourg (Varrette et al, 2014). We thank the Laboratoire national de santé for a second blinded assessment and confirmation of the pathological report. Finally, we thank Martin Nurmik for manuscript corrections.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors have a corresponding patent application (LU100371).
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Letellier, E., Schmitz, M., Ginolhac, A. et al. Loss of Myosin Vb in colorectal cancer is a strong prognostic factor for disease recurrence. Br J Cancer 117, 1689–1701 (2017). https://doi.org/10.1038/bjc.2017.352
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.352
Keywords
This article is cited by
-
Myosin Vb as a tumor suppressor gene in intestinal cancer
Oncogene (2022)
-
Targeting the cytoskeleton against metastatic dissemination
Cancer and Metastasis Reviews (2021)