Abstract
Background
Predictive biomarkers for the recurrence of hepatocellular carcinoma (HCC) have great benefit in the selection of treatment options, including liver transplantation (LT), for HCC. The purpose of this study was to identify specific microRNAs (miRs) in exosomes from the serum of patients with recurrent HCC and to validate these molecules as novel biomarkers for HCC recurrence.
Methods
We employed microarray-based expression profiling of miRs derived from exosomes in the serum of HCC patients to identify a biomarker that distinguishes between patients with and without HCC recurrence after LT. This was followed by the validation in a separate cohort of 59 HCC patients who underwent living related LT. The functions and potential gene targets of the recurrence-specific miRs were analysed using a database, clinical samples and HCC cell lines.
Results
We found that miR-718 showed significantly different expression in the serum exosomes of HCC cases with recurrence after LT compared with those without recurrence. Decreased expression of miR-718 was associated with HCC tumour aggressiveness in the validated cohort series. We identified HOXB8 as a potential target gene of miR-718, and its upregulation was associated with poor prognosis.
Conclusion
Circulating miRs in serum exosomes have potential as novel biomarkers for predicting HCC recurrence.
Similar content being viewed by others
Main
Hepatocellular carcinoma (HCC) is the fourth most common malignancy in Japan and the fifth worldwide, and the mainstay of its treatment is hepatic resection (Llovet, 2005; Okita, 2006). Many HCC patients do not meet the indications for hepatectomy due to the underlying liver cirrhosis; therefore, liver transplantation (LT) for HCC has been introduced and improved with restrictive selection criteria such as the Milan criteria, based on the report by Mazzaferro et al (1996). The size and number of HCCs have been reported as essential factors for consideration in any of the LT criteria for HCC (Mazzaferro et al, 2008; Taketomi et al, 2009; Sugimachi et al, 2011). However, there are no established molecular-based standards regarding the preoperative evaluation and selection of HCC patients for LT.
Exosomes are small membrane vesicles (30–100 nm) derived from the luminal membranes of multivesicular bodies and are constitutively released by fusion with the cell membrane (Simons and Raposo, 2009). Exosomes transfer not only membrane components but also nucleic acids to other cells; therefore, cell-derived exosomes have recently been described as a new mode of cell-to-cell communication (Peinado et al, 2012). According to the current version of the exosome database ExoCarta (http://www.exocarta.org) (Mathivanan et al, 2012), 764 microRNAs (miRs) have been identified in exosomes derived from several different cell types and from multiple organisms. Exosomal transport of miRs has led to an increased interest in cancer research in exosomes as carriers of genetic information (Valadi et al, 2007; Hannafon and Ding, 2013).
miRs are small (17–21 nt), non-coding RNAs sthat regulate gene expression at the post-transcriptional level through the RNA interference pathway. Currently, ∼2000 miRs have been described in humans, and a single miR may regulate many mRNAs. Through this mechanism, miRs are essential components in the regulation of many cellular and developmental processes, including developmental timing, organ development, differentiation, proliferation, immune regulation and cancer development and progression. Depending on their target gene(s) and level of expression, miRs may function as either tumour suppressors or oncogenes and assist in the promotion or suppression of cancer growth and progression.
Because they are resistant to RNases, exosomal miRs are stable in the blood and, thus, are considered to be promising candidates as novel biomarkers of various cancers (Katsuda et al, 2013). Exosomal miRs extracted from various body fluids, such as blood, urine and saliva, have been investigated as novel diagnostic tools for early cancer detection or malignant potential (Taylor and Gercel-Taylor, 2008; Murakami et al, 2012; Lau et al, 2013) Unlike conventional protein-based cancer biomarkers, such as carcinoembryonic antigen and α-fetoprotein (AFP), exosomal miRs possibly have biological and physiological functions in cancer progression, as well as serve as simple biomarkers (Hagiwara et al, 2014). Several serum miRs have been reported to be biomarkers for HCC (Yamamoto et al, 2009; Qi et al, 2011; Li et al, 2012; Liu et al, 2012; Hu et al, 2013) but were not specifically measured in serum exosomes. Thus, their functions in the progression and recurrence of the disease have not been clear.
In this study, we sought to identify specific biomarkers in serum exosomes collected at the time of surgery that reliably define patients with recurrent HCC after LT. We believe that specific miRs in exosomes derived from recurrent HCC could be used as novel biomarkers. Furthermore, such markers could help elucidate the molecular mechanisms underlying HCC recurrence.
Materials and methods
Clinical samples
Six cases that underwent living donor LT (LDLT) were selected from records of the Department of Surgery and Science, Kyushu University. All cases were routinely followed for more than 5 years after surgery, and tumour recurrence was found in four cases but not in the other two cases. Serum samples from these six cases were obtained before surgery for miR microarray analysis for recurrence-specific miR profiling. For the validation analysis, serum from 59 consecutive patients with HCC who underwent LDLT from 2002 to 2011 at Kyushu University was obtained before surgery. The inclusion criteria consisted of (1) no modality except LDLT available to treat the HCC patients with end-stage liver disease; (2) no extrahepatic metastasis; and (3) no major vascular infiltration. There was no restriction on the tumour size or the number of tumours. Since 2009, we have been applying new selection criteria for HCC patients (a tumour size of <5 cm or a serum des-gamma-carboxy prothrombin (DCP) level of <300 mAU ml−1) because these two factors were found to be favourable independent factors for the recurrence of HCC after LDLT (Taketomi et al, 2009). The study was approved by the institutional review board (the Ethics and Indications Committee of Kyushu University, 24–160). Written informed consent for the study was obtained from each patient. The median age of the 59 patients was 57.7 years (range 21–72 years). The aetiologies were hepatitis C infection in 39 patients (66%), hepatitis B infection in 12 (20%) and other causes in eight (14%).
Purification of exosomes from serum and cultured cell supernatant
Peripheral blood was collected and centrifuged at 3000 r.p.m. for 10 min at 4 °C to collect serum. Supernatant of HuH7 cells were collected after incubation with exosome-depleted FBS for 3 days. Supernatant was collected and centrifuged at 2000 g for 10 min at room temperature, and at 12 000 g for 30 min followed by filtration through 0.22-μm filter to remove cell debris. We purified exosome using the ultracentrifuge method. Serum of 1.5 ml was used for western blotting and microarray analysis, and that of 300 μl was used for RT–PCR. The blood serum samples or cell supernatant were ultracentrifuged once at 100 000 g for 70 min at 4 °C to collect the pellet containing exosome. The whole amount of acquired pellet was washed with 11 ml of PBS once, and used for the further experiment in each sample.
Transmission electron microscopy
The pellets were washed once with 11 ml phosphate-buffered saline. The samples were dissolved in HEPES (4-[2-hydroxyethyl]-1-piperazine ethanesulfonic acid) buffer, and a drop of the suspension was placed on a sheet of parafilm. A carbon-coated copper grid was floated on the drop for 10 s. The grid was then removed and excess liquid was drained by touching the edge of the grid against a piece of clean filter paper. The grid was placed onto a drop of 2% uranyl acetate or phosphotungstic acid, pH 7.0, for ∼5 s, and the excess liquid was drained off. The grid was allowed to dry for several minutes and then examined using a JEM-1200 EX microscope (JEOL, Akishima, Japan) at 80 K electron volts.
Western blotting
Proteins were extracted from the ultracentrifugation pellets and electrophoresed in 10% concentrated polyacrylamide gel and then electrophoresed and then electroblotted. The primary mouse monoclonal antibody against TSG101 (Abcam, Cambridge UK) was used at a dilution of 1 : 200. TSG101 protein were detected using horseradish peroxidase-conjugated secondary antibodies (GE Healthcare Japan, Tokyo, Japan) and the ImageQuant LAS 4000 mini system (GE Healthcare).
Isolation of miRs
RNAs were isolated using the miRNeasy Mini Kit (Qiagen, Venlo, Netherlands) according to the manufacturer’s protocol. Exosomes purified from specific volumes of serum were diluted in 1 ml Qiazol Lysis Reagent. After incubation for 5 min at room temperature, 10 μl 0.1 nM cel-miR-39 mimic was added to each sample, followed by vortexing for 30 s. Subsequent extraction and cartridge work were performed according to the manufacturer’s protocol.
miR microarray analysis
Extracted total RNA was labelled with Hy5 using the miRCURY LNA microRNA Array labeling kit (Exiqon, Vedbaek, Denmark). Labelled RNAs were hybridised onto 3D-Gene Human miRNA Oligo chips (Toray, Kamakura, Japan). Fluorescent signals were scanned and analysed using the 3D-Gene Scanner (Toray). The raw data from each spot were normalised by subtraction of the background signal mean intensity, determined by the 95% confidence intervals of the signal intensities of all blank spots. Valid measurements were considered those in which the signal intensity of both duplicate spots was >2 s.d. of the background signal intensity.
Quantitative real time-PCR for miRs
miR expression in serum samples from HCC/CRC patients was assessed using quantitative real time-PCR (qRT–PCR). cDNA was synthesised from total RNA using TaqMan microRNA hsa-miR-718 and hsa-miR-1246 specific primers (Applied Biosystems, Carlsbad, CA, USA) and a TaqMan Micro-RNA Reverse Transcription Kit (Applied Biosystems). PCR amplification was performed using a LightCycler PCR instrument (Roche, Basel, Switzerland) and the LightCycler-FastStart DNA Master SYBR Green I Kit (Roche) as described previously (Ogawa et al, 2005). Relative quantification of miR expression was calculated using the 2−ΔΔCt method. The raw data were presented as the relative target miR amount, normalised to cel-miR-39 as the invariant control for the samples, and compared with a reference sample.
Cell lines, pre-miR and MTT cell proliferation assays
The human HCC cell lines HuH7 and PLC/PRF/5 were obtained from the American Type Culture Collection and the Japanese Collection of Research Bioresources. Huh7 cells, an HCC cell line that expresses low levels of miR-718, were transfected with either pre-miR-718 or pre-miR negative control (Ambion Pre-miR miRNA Precursors, Applied Biosystems) at a 30-nmol l−1 final concentration using Lipofectamine RNAiMAX (Life Technologies, Carlsbad, CA, USA), according to the manufacturer’s instructions. Cells were seeded at 4.0 × 103 cells per well in 96-well flat-bottomed microtiter plates in a final volume of 100 μl culture medium per well. Cells were incubated in a humidified atmosphere (37 °C and 5% CO2) for 24, 48 or 72 h following transfection. 3-(4,5-Dimethylthiazole-2-yl)-2,5-diphenyl tetrazolium bromide assays were used to measure cell proliferation during each time period, as described previously (Nishida et al, 2011). Three independent replicate assays were performed.
Prognostic analysis of public gene expression data
The HCC gene expression data set GSE14520, submitted by Roessler et al (2012), was downloaded from the NCBI Gene Expression Omnibus. The prognostic relevance of gene expression in GSE14520 was calculated using the log-rank test and Cox proportional hazards model.
Validation of HOXB8 expression in HCC patients
A total of 58 patients with HCC who underwent hepatic resection at Kyushu University Beppu Hospital and its affiliated hospitals from 2001 to 2003 were enroled in this study. The resected tumour and paired non-tumourous tissue specimens were immediately frozen in liquid nitrogen and kept at −80 °C until analysis. Written informed consent was obtained from all patients. Total RNA was extracted from frozen tissue specimens using ISOGEN (Nippon Gene, Tokyo, Japan) following the manufacturer’s protocol. cDNAs were reverse transcribed from all RNA samples. HOXB8 expression levels were quantified using the LightCycler 480 Probes Master kit (Roche), following the manufacturer’s protocol, and specific HOXB8 primers. GAPDH was used as the internal control.
Luciferase assay
A luciferase reporter plasmid was constructed by subcloning the HOXB8 3′-untranslated region (UTR) fragment length, which contained the miR-718 binding sites, into the pmirGlo Dual-luciferase miRNA Target Expression Vector (Promega, Madison, WI, USA), positioned 5′ to the firefly luciferase-coding region. The nucleotide sequences of the constructed plasmids were confirmed by DNA sequencing analysis. For the luciferase reporter assays, HuH7 and PLC/PRL/5 cells were seeded in 96-well plates and then co-transfected with the pmirGlo-HOXB8 3′-UTR construct and pre-miR-718 or the pre-miR negative control (Ambion). Assays were conducted 48 h after transfection using the Dual-Luciferase Reporter Assay System (Promega). Firefly luciferase signals were normalised to Renilla luciferase signals. Transfections were conducted three times in independent experiments.
Statistical analysis
For continuous variables, the data were expressed as means±s.d. The relationships among miR expression, clinicopathological factors and the in vitro assay data were analysed using Student’s t-tests, χ2-tests and ANOVA. Post-operative survival curves, measured from the day of surgery, were plotted using the Kaplan–Meier method, and the log-rank test was applied for comparison analysis. All differences were considered statistically significant at the level of P<0.05. Statistical analysis was performed using the JMP software package (SAS Institute, Cary, NC, USA).
Results
Identification of exosome in serum of HCC patients
We purified round-shaped vesicles with a diameter of about 25–75 nm from serum of HCC patients using ultracentrifuge method (Figure 1A and B). Western blot analysis demonstrated enrichment of an exosomal marker (TSG101; Figure 1C). These results confirmed the efficacy of our protocol for isolating exosomes from serum.
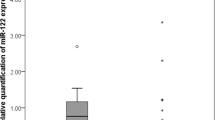
Exosomal miR in the serum was associated with malignant progression of HCC. (A, B) Exosomes were purified from serum of HCC patients using ultracentrifuge method and examined by electron microscopy; (A) scale bar, 200 nm and (B) scale bar, 100 nm. (C) The exosome extract from serum after an ultracentrifuge method was analysed by western blotting. The data demonstrated that extracts were enriched with exosomal marker protein TSG101. Supernatant of cultured cancer cell lines were used as control. (D–G) miR-718 expression levels in the exosomes from serum of HCC patients who underwent LT were examined according to clinicopathological parameters. (D) Tumour diameter, (E) histological differentiation, (F) vascular invasion, (G) recurrence of the disease. Data represent the means±s.d.
miR profiling and identification of recurrence-specific exosomal miRs
To identify recurrence-specific exosomal miRs in serum of HCC patients, we performed miRNA microarray analyses of samples from four patients with recurrence after LDLT and from two patients without recurrence. Two miRs, hsa-miR-718 and miR-1246, were significantly downregulated and upregulated, respectively, in HCC cases with recurrence compared with those without recurrence (fold change >1.5 and Q<0.05).
Validation of the clinicopathologic significance of miR-718 expression in serum exosomes
We validated the clinicopathological significance of miR-718 and miR-1246 expression in serum exosomes by qRT–PCR. In the clinicopathologic analyses, miR-718 expression was significantly lower in those cases with larger tumour diameters and recurrence after LDLT (Figures 1D–G). We then categorised 59 HCC cases with LDLT into two groups according to the mean miR-718 expression level in preoperative serum exosomes. Compared with the high miR-718 expression group, the low expression group showed poorer histological differentiation (P=0.026), a higher incidence of tumours beyond the Milan criteria (P=0.04), and stronger tendencies of association with larger (≥3 cm) tumour size (P=0.09) and tumour number (P=0.07; Table 1). On the other hand, no correlations between exosomal miR-1246 expression and the clinicopathological factors were found. A recurrence-free survival curve according to the miR-718 expression level showed that six cases with recurrent disease after LDLT were included in the low miR-718 expression group, whereas no cases showed recurrence in the high miR-718 expression group (Supplementary Figure 1). However, this difference was not significant due to the low incidence of recurrence. In the subgroup analysis, patients who showed low miR-718 expression and tumour diameter ⩾3 cm (n=15) showed lower recurrence-free survival rate than other patients (n=44) with statistically significant difference (P=0.0002 by log-rank test, Supplementary Figure 2)
miR-718 suppresses cell proliferation in HCC cells
Since low miR-718 expression was associated with the malignant potential of HCC, the function of miR-718 was assessed using a MTT proliferation assay. Transfection with pre-miR-718 significantly suppressed proliferation of Huh7 cells compared with the control (P=0.01, Figure 2A and B). These experiments were performed using two different HCC cell lines (Huh7 and PLC/PRF/5), which showed similar results (data not shown). These results support the suggestion that low miR-718 expression is associated with the malignant potential of HCC.
miR-718 inhibited cell proliferation and targeted HOXB8 gene. (A, B) Induction of miR-718 inhibited the proliferation of HCC cells. (A) miR-718 expression levels in HuH7 cells transfected with pre-miR-718 were significantly higher than those of negative control (nc) cells. (B) The cell proliferation ratio of HuH7 cells transfected with pre-miR-718 were significantly suppressed compared to the negative control (*P<0.01). (C) Overall survival curves of the low and high HOXB8 expression groups for 58 HCC patients who underwent hepatectomy. The patients with high HOXB8 expression showed significantly poorer long-term prognosis after hepatectomy.
HOXB8 as a potential target gene of miR-718 in HCC patients
The TargetScan online database, a collection of information regarding predicted miRNA targets, suggested nine potential target genes of miR-718 (RNF44, HOXB8, TBX1, GORAB, BAD, ADRBK1, OSR1, MPDU1, LRP8; http://www.targetscan.org). Using previously reported expression array data from 242 HCC samples (Roessler et al, 2012), we examined the association between the expression status of the potential miR-718 target genes in the primary HCC and post-operative prognosis (Supplementary Table 1). We demonstrated that the upregulation of HOXB8 expression affected the poor overall and recurrence-free survivals of HCC patients, with statistical significance (Supplementary Figure 3). Next, to validate the results from the public database, we examined the association between HOXB8 expression and HCC patient prognosis using 58 primary HCCs with curative hepatectomy. Patients with expression levels less than the median value were assigned to a low expression group (n=35) and those above the median to a high expression group (n=23). In the validation study, the high HOXB8 expression group had a significantly poorer prognosis than the low expression group (P=0.048, Figure 2C), which was in concordance with the published array data from the 242 HCC cases. Multivariate analysis showed that high HOXB8 expression was an independent prognostic predictor (Table 2). In Huh7 and PLC/PRF/5 HCC cell lines, co-transfected cells expressing both miR-718 and the HOXB8 3′ UTR showed a significant reduction in luciferase activity (P<0.05, Student’s t-test) compared with the control (Figure 3A–C); this suggested that HOXB8 is a target gene of miR-718 in HCC. These results indicated that the suppressed expression of miR-718 results in the upregulation of HOXB8 and contributes to the poor prognosis of HCC patients.
miR-718 inhibited the transcription of HOXB8 gene. (A) Sequences of the miR-718 binding sites in the 3′UTRs of transcripts encoding HOXB8. (B) Luciferase analyses in HuH7 and PLC/PRF/5 cell lines. HOXB8 3′ UTR luciferase vector+miR-718 transfectants showed lower luciferase activities than did control cells. Relative luciferase levels were calculated by the following: (Sample Luc/Sample Renilla)/(Control Luc/Control Renilla). Luc, raw firefly luciferase activity; Renilla, Renilla activity (internal transfection control); Pre-miR n.c., pre-miR negative control.
Discussion
Here we demonstrated that a functional miR in exosomes was associated with the recurrence of HCC after LDLT. We explored preoperative biomarkers that predict HCC recurrence after surgery, which could help select the patients who need LT, resulting in the proper usage of donor organs. Our findings indicate that the specific miRs in serum exosomes are not only biomarkers but also biologically functional molecules in tumour recurrence.
Sensitive biomarkers for early diagnosis and post-operative recurrence of HCC have been explored in the serum and resected tumour specimens. Previous reports showed that the miR profiles from resected tumour tissues could predict the risk of HCC recurrence after LT (Barry et al, 2012; Han et al, 2012). Those results suggested that miRs have potential to be biomarkers; yet, they could not aid in the preoperative selection of LT because the profiles were specific to the extracted liver. Serum AFP and DCP expression levels are useful tumour markers for the detection and monitoring of HCC, but the false negative rates are high and they cannot predict post-operative recurrence (Volk et al, 2007). Serum miRs are considered to be promising biomarkers to improve sensitivity, specificity, early detection and prognostic prediction of HCC (Qu et al, 2011; Wang et al, 2014b).
Recent study showed that exosomal miR-21 was enriched in the serum of HCC patients (Wang et al, 2014a), but the present study is the first to explore comprehensive miRs in exosomes, which circulate in the serum of patients. We consider that there are several advantages to using exosomal miRs compared with conventional proteomic methods. Exosomal miRs are stable in the blood and easy to quantitate by qRT–PCR. Moreover, their contents are similar to those in the original tumour (Hu et al, 2012); thus, exosomal miR profiles may be useful for cancer diagnostics by reflecting the clinical behaviour and therapeutic response of HCC. We found that exosomal miR-718 was significantly suppressed in patients with tumour recurrence after LT. These results are in accord with the reports by Shrauder et al (2012), which showed that miR-718 was a significantly suppressed miR in the blood of breast cancer patients.
Considering that miR-718 significantly suppressed proliferation of HCC cells, it may be a tumour-suppressive miR. Our present data showed that the downregulation of miR-718 results in the overexpression of HOXB8 in HCC patients. The aberrant expression of HOX genes, a highly conserved subgroup of the homeobox superfamily, is known to promote tumorigenesis and metastasis of various cancers (Shah and Sukumar, 2010). Our results suggest that the miR-718-HOXB8 axis has an important role in the progression and recurrence of HCC. As the primary goal of our study was to identify the significantly useful biomarkers in serum exosomes, we could not show direct evidence that exosomal miR-718 was secreted from either HCC cells or other host cells, such as bone marrow-derived cells. Functional exosomal miRs both from cancer cells and bone marrow mesenchymal stem cells have been reported (Cereghetti and Lee, 2014; Ono et al, 2014). The role of exosomal miRs in the communication between cells and organs of cancer patients is also of interest, and further studies are definitely needed to clarify the mechanisms.
In conclusion, this study provides new and important information on the functional significance of serum exosomal miRs that could be novel biomarkers for predicting the recurrence and therapeutic targets of HCC.
Change history
03 February 2015
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Barry CT, D’Souza M, McCall M, Safadjou S, Ryan C, Kashyap R, Marroquin C, Orloff M, Almudevar A, Godfrey TE (2012) Micro RNA expression profiles as adjunctive data to assess the risk of hepatocellular carcinoma recurrence after liver transplantation. Am J Transplant 12 (2): 428–437.
Cereghetti DM, Lee PP (2014) Tumor-derived exosomes contain microRNAs with immunological function: implications for a novel immunosuppression mechanism. MicroRNA 2 (3): 194–204.
Hagiwara K, Ochiya T, Kosaka N (2014) A paradigm shift for extracellular vesicles as small RNA carriers: from cellular waste elimination to therapeutic applications. Drug Deliv Transl Res 4: 31–37.
Han ZB, Zhong L, Teng MJ, Fan JW, Tang HM, Wu JY, Chen HY, Wang ZW, Qiu GQ, Peng ZH (2012) Identification of recurrence-related microRNAs in hepatocellular carcinoma following liver transplantation. Mol Oncol 6 (4): 445–457.
Hannafon BN, Ding WQ (2013) Intercellular communication by exosome-derived microRNAs in cancer. Int J Mol Sci 14 (7): 14240–14269.
Hu G, Drescher KM, Chen XM (2012) Exosomal miRNAs: biological properties and therapeutic potential. Front Genet 3: 56.
Hu QY, Jiang H, Su J, Jia YQ (2013) MicroRNAs as biomarkers for hepatocellular carcinoma: a diagnostic meta-analysis. Clin Lab 59 (9-10): 1113–1120.
Katsuda T, Kosaka N, Ochiya T (2013) The roles of extracellular vesicles in cancer biology: towards the development of novel cancer biomarkers. Proteomics 14 (4-5): 412–425.
Lau C, Kim Y, Chia D, Spielmann N, Eibl G, Elashoff D, Wei F, Lin YL, Moro A, Grogan T, Chiang S, Feinstein E, Schafer C, Farrell J, Wong DT (2013) Role of pancreatic cancer-derived exosomes in salivary biomarker development. J Biol Chem 288 (37): 26888–26897.
Li L, Guo Z, Wang J, Mao Y, Gao Q (2012) Serum miR-18a: a potential marker for hepatitis B virus-related hepatocellular carcinoma screening. Dig Dis Sci 57 (11): 2910–2916.
Liu AM, Yao TJ, Wang W, Wong KF, Lee NP, Fan ST, Poon RT, Gao C, Luk JM (2012) Circulating miR-15b and miR-130b in serum as potential markers for detecting hepatocellular carcinoma: a retrospective cohort study. BMJ open 2 (2): e000825.
Llovet JM (2005) Updated treatment approach to hepatocellular carcinoma. J Gastroenterol 40 (3): 225–235.
Mathivanan S, Fahner CJ, Reid GE, Simpson RJ (2012) ExoCarta 2012: database of exosomal proteins, RNA and lipids. Nucleic Acids Res 40 (Database issue): D1241–D1244.
Mazzaferro V, Chun YS, Poon RT, Schwartz ME, Yao FY, Marsh JW, Bhoori S, Lee SG (2008) Liver transplantation for hepatocellular carcinoma. Ann Surg Oncol 15 (4): 1001–1007.
Mazzaferro V, Regalia E, Doci R, Andreola S, Pulvirenti A, Bozzetti F, Montalto F, Ammatuna M, Morabito A, Gennari L (1996) Liver transplantation for the treatment of small hepatocellular carcinomas in patients with cirrhosis. N Engl J Med 334 (11): 693–699.
Murakami Y, Toyoda H, Tanahashi T, Tanaka J, Kumada T, Yoshioka Y, Kosaka N, Ochiya T, Taguchi YH (2012) Comprehensive miRNA expression analysis in peripheral blood can diagnose liver disease. PLoS One 7 (10): e48366.
Nishida N, Mimori K, Fabbri M, Yokobori T, Sudo T, Tanaka F, Shibata K, Ishii H, Doki Y, Mori M (2011) MicroRNA-125a-5p is an independent prognostic factor in gastric cancer and inhibits the proliferation of human gastric cancer cells in combination with trastuzumab. Clin Cancer Res 17 (9): 2725–2733.
Ogawa K, Utsunomiya T, Mimori K, Tanaka F, Inoue H, Nagahara H, Murayama S, Mori M (2005) Clinical significance of human kallikrein gene 6 messenger RNA expression in colorectal cancer. Clin Cancer Res 11 (8): 2889–2893.
Okita K (2006) Clinical aspects of hepatocellular carcinoma in Japan. Intern Med 45 (5): 229–233.
Ono M, Kosaka N, Tominaga N, Yoshioka Y, Takeshita F, Takahashi RU, Yoshida M, Tsuda H, Tamura K, Ochiya T (2014) Exosomes from bone marrow mesenchymal stem cells contain a microRNA that promotes dormancy in metastatic breast cancer cells. Sci Signal 7 (332): ra63.
Peinado H, Aleckovic M, Lavotshkin S, Matei I, Costa-Silva B, Moreno-Bueno G, Hergueta-Redondo M, Williams C, Garcia-Santos G, Ghajar C, Nitadori-Hoshino A, Hoffman C, Badal K, Garcia BA, Callahan MK, Yuan J, Martins VR, Skog J, Kaplan RN, Brady MS, Wolchok JD, Chapman PB, Kang Y, Bromberg J, Lyden D (2012) Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med 18 (6): 883–891.
Qi P, Cheng SQ, Wang H, Li N, Chen YF, Gao CF (2011) Serum microRNAs as biomarkers for hepatocellular carcinoma in Chinese patients with chronic hepatitis B virus infection. PLoS One 6 (12): e28486.
Qu KZ, Zhang K, Li H, Afdhal NH, Albitar M (2011) Circulating microRNAs as biomarkers for hepatocellular carcinoma. J Clin Gastroenterol 45 (4): 355–360.
Roessler S, Long EL, Budhu A, Chen Y, Zhao X, Ji J, Walker R, Jia HL, Ye QH, Qin LX, Tang ZY, He P, Hunter KW, Thorgeirsson SS, Meltzer PS, Wang XW (2012) Integrative genomic identification of genes on 8p associated with hepatocellular carcinoma progression and patient survival. Gastroenterology 142 (4): 957–966 e12.
Schrauder MG, Strick R, Schulz-Wendtland R, Strissel PL, Kahmann L, Loehberg CR, Lux MP, Jud SM, Hartmann A, Hein A, Bayer CM, Bani MR, Richter S, Adamietz BR, Wenkel E, Rauh C, Beckmann MW, Fasching PA (2012) Circulating micro-RNAs as potential blood-based markers for early stage breast cancer detection. PLoS One 7 (1): e29770.
Shah N, Sukumar S (2010) The Hox genes and their roles in oncogenesis. Nat Rev Cancer 10 (5): 361–371.
Simons M, Raposo G (2009) Exosomes—vesicular carriers for intercellular communication. Curr Opin Cell Biol 21 (4): 575–581.
Sugimachi K, Shirabe K, Taketomi A, Soejima Y, Iguchi T, Takeishi K, Toshima T, Aishima S, Tajima T, Maehara Y (2011) Prognostic significance of preoperative imaging in recipients of living donor liver transplantation for hepatocellular carcinoma. Transplantation 91 (5): 570–574.
Taketomi A, Sanefuji K, Soejima Y, Yoshizumi T, Uhciyama H, Ikegami T, Harada N, Yamashita Y, Sugimachi K, Kayashima H, Iguchi T, Maehara Y (2009) Impact of des-gamma-carboxy prothrombin and tumor size on the recurrence of hepatocellular carcinoma after living donor liver transplantation. Transplantation 87 (4): 531–537.
Taylor DD, Gercel-Taylor C (2008) MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol 110 (1): 13–21.
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9 (6): 654–659.
Volk ML, Hernandez JC, Su GL, Lok AS, Marrero JA (2007) Risk factors for hepatocellular carcinoma may impair the performance of biomarkers: a comparison of AFP, DCP, and AFP-L3. Cancer Biomark 3 (2): 79–87.
Wang H, Hou L, Li A, Duan Y, Gao H, Song X (2014a) Expression of serum exosomal microRNA-21 in human hepatocellular carcinoma. Biomed Res Int 2014: 864894.
Wang L, Yao M, Dong Z, Zhang Y, Yao D (2014b) Circulating specific biomarkers in diagnosis of hepatocellular carcinoma and its metastasis monitoring. Tumour Biol 35 (1): 9–20.
Yamamoto Y, Kosaka N, Tanaka M, Koizumi F, Kanai Y, Mizutani T, Murakami Y, Kuroda M, Miyajima A, Kato T, Ochiya T (2009) MicroRNA-500 as a potential diagnostic marker for hepatocellular carcinoma. Biomarkers 14 (7): 529–538.
Acknowledgements
We thank K Oda, M Kasagi, S Kohno, T Kawano and M Aoyagi for their technical assistance. This work was supported in part by the following grants and foundations: Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Scientific Research, grant number 24592005; the Sagawa Foundation for Promotion of Cancer Research, Japan; and the Novartis Foundation for the Promotion of Science, Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Sugimachi, K., Matsumura, T., Hirata, H. et al. Identification of a bona fide microRNA biomarker in serum exosomes that predicts hepatocellular carcinoma recurrence after liver transplantation. Br J Cancer 112, 532–538 (2015). https://doi.org/10.1038/bjc.2014.621
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2014.621
Keywords
This article is cited by
-
A state-of-the-art review of the recent advances in exosome isolation and detection methods in viral infection
Virology Journal (2024)
-
Circulating exosomal microRNAs as potential prognostic biomarkers in gastrointestinal cancers: a systematic review and meta-analysis
Cancer Cell International (2023)
-
Migrasomes, a new mode of intercellular communication
Cell Communication and Signaling (2023)
-
The role of Exosomal miRNAs in cancer
Journal of Translational Medicine (2022)
-
A functional screen with metformin identifies microRNAs that regulate metabolism in colorectal cancer cells
Scientific Reports (2022)