Abstract
Background:
The Breast and Ovarian Analysis of Disease Incidence and Carrier Estimation Algorithm (BOADICEA) is a risk prediction model that is used to compute probabilities of carrying mutations in the high-risk breast and ovarian cancer susceptibility genes BRCA1 and BRCA2, and to estimate the future risks of developing breast or ovarian cancer. In this paper, we describe updates to the BOADICEA model that extend its capabilities, make it easier to use in a clinical setting and yield more accurate predictions.
Methods:
We describe: (1) updates to the statistical model to include cancer incidences from multiple populations; (2) updates to the distributions of tumour pathology characteristics using new data on BRCA1 and BRCA2 mutation carriers and women with breast cancer from the general population; (3) improvements to the computational efficiency of the algorithm so that risk calculations now run substantially faster; and (4) updates to the model’s web interface to accommodate these new features and to make it easier to use in a clinical setting.
Results:
We present results derived using the updated model, and demonstrate that the changes have a significant impact on risk predictions.
Conclusion:
All updates have been implemented in a new version of the BOADICEA web interface that is now available for general use: http://ccge.medschl.cam.ac.uk/boadicea/.
Similar content being viewed by others
BOADICEA model
The Breast and Ovarian Analysis of Disease Incidence and Carrier Estimation Algorithm (BOADICEA) (Antoniou et al, 2004, 2008a) is a risk prediction model for familial breast and ovarian cancer. The model is used to compute BRCA1 and BRCA2 mutation carrier probabilities and age-specific risks of breast and ovarian cancer. It was developed using complex segregation analysis of breast and ovarian cancer based on a combination of families identified through population-based studies of breast cancer, and families with multiple affected individuals who had been screened for BRCA1 and BRCA2 mutations. The latest version of the model was based on 2785 families, of which 537 segregated BRCA1 and/or BRCA2 mutations. BOADICEA models the simultaneous effects of BRCA1 and BRCA2 mutations and assumes that the residual familial clustering of breast cancer is explained by a polygenic component (a large number of genes each of small effect on risk) with a variance that decreases linearly with age.
BOADICEA has been validated in a large series of families from UK genetics clinics (Antoniou et al, 2008b). In the United Kingdom, it is recommended as a risk assessment tool in the National Institute for Health and Care Excellence clinical guideline CG164 (National Institute for Health and Care Excellence, 2013) and has been incorporated in the guidelines of several countries for the management of familial breast cancer (Ontario Breast Screening Program, 2012; Riley et al, 2012; Smith et al, 2012).
Cancer incidences
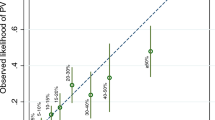
To obtain risk predictions, BOADICEA considers the occurrence of breast, ovarian, pancreatic and prostate cancers in families (Antoniou et al, 2008a). To provide a consistent model, the breast and ovarian cancer incidences over all assumed genetic effects are constrained to agree with population incidences (Antoniou et al, 2001). The old implementation of the model assumes calendar period- and cohort-specific incidences for the United Kingdom that span the period 1960–1997, taken from the Cancer Incidence in Five Continents (CI5) publications (Ferlay et al, 2010). These incidences were the most up-to-date and relevant cancer incidences available to us when the BOADICEA model was initially developed. However, breast, ovarian and prostate cancer incidences have increased over time (Hayat et al, 2007). For example, UK age-specific breast cancer incidences for females shown in Figure 1A (Ferlay et al, 2010) show that breast cancer incidence has increased over the time period between 1960 and 2010. As BOADICEA is used to predict future risks of developing breast or ovarian cancer, it is essential to consider the most up-to-date cancer incidences. Cancer incidences from recent calendar periods (1992–2010) are now available and have been included in this version of BOADICEA.
(A) The age-specific female breast cancer incidences per 100 000 for the United Kingdom for the periods 1960–1963 and 1973–1977, and the years 1993 and 2010. (B) The age-specific female breast cancer incidences per 100 000 for 2007 for the United Kingdom, Australia, Canada, Sweden and the United States. (C) The old, non-smoothed and smoothed updated incidence per 100 000 for females born in 1975 used in the BOADICEA code. (D) The BOADICEA predicted risk for a 30-year-old female born in 1975 with no family history information for the United Kingdom, updated and old, and other countries.
Furthermore, incidences of breast, ovarian, pancreatic and prostate cancers vary widely by geographical region (Parkin et al, 2005). Figure 1B demonstrates the differences in age-specific female breast cancer incidence between various countries in the year 2007. As BOADICEA is now used in over 45 countries, in the new version of BOADICEA we have included incidences specific to other countries.
Breast tumour pathology
The BOADICEA has recently been extended to incorporate breast tumour pathology information, where breast cancer subtypes are treated as distinct disease end points (Mavaddat et al, 2010). In particular, oestrogen receptor (ER) status, triple-negative (TN) status (ER, progesterone (PR) and HER2 negative) and expression of basal markers (CK5/6 and CK14) are taken into account. Initially, tumour subtype distributions for BRCA1 and BRCA2 mutation carriers were based on data from the Breast Cancer Linkage Consortium (BCLC) (Lakhani et al, 2002), and data from the Surveillance, Epidemiology and End Results (SEER) Program (Surveillance Epidemiology and End Results (SEER) Program, 2006) were used to obtain the ER distribution in the general population. However, because of the relatively small number of mutation carriers in the BCLC study (182 BRCA1 and 64 BRCA2), these distributions were imprecise, particularly for BRCA2 tumours. Recent analyses based on the much larger collaborative data set from the Consortium of Investigators of Modifiers of BRCA1/2 (CIMBA) demonstrated additional differences in the characteristics of BRCA1 and BRCA2 tumours, such as an increasing proportion of ER-negative tumours with age among BRCA2 carriers (Mavaddat et al, 2012).Up-to-date data from CIMBA, in combination with data from the Breast Cancer Association Consortium (BCAC) on the age-specific frequencies in the general population, by tumour type, allow us to incorporate more accurate distributions of ER, PR and HER2 status into BOADICEA.
Computational requirements
In order to exploit fully the predictive potential of genetic models in the future, it will be necessary to add the explicit effects of known breast cancer susceptibility variants such as common SNPs (Michailidou et al, 2013) and rare moderate-risk variants, such as in ATM, CHEK2, PALB2 and BRIP1 (Meijers-Heijboer et al, 2002; Renwick et al, 2006; Seal et al, 2006; Rahman et al, 2007). However, this presents a substantial problem in terms of the future computational requirements of the BOADICEA program. As the old implementation of the model uses the Elston–Stewart algorithm (Elston and Stewart, 1971) to compute pedigree likelihoods, incorporating additional genetic effects will result in an exponential increase in runtime. This problem is exacerbated by the fact that we may not be able to rely on future increases in single-processor performance (Fuller and Millett, 2011). In order to use these more sophisticated models in a clinical setting, they must run faster.
BOADICEA web application
We have implemented the BOADICEA model as a standalone FORTRAN program, termed here the BOADICEA FORTRAN program (BFP). In recent years, scientists have used the BFP as a research tool. However, in practice, it is time consuming to set up and run calculations using the BFP alone, which makes it inappropriate for use in a clinical setting. To address this problem, we developed the BOADICEA web application (BWA; http://ccge.medschl.cam.ac.uk/boadicea/), (Cunningham et al, 2012), a user-friendly web interface to the BFP that makes it much easier for health-care professionals and members of the public to run BOADICEA calculations. The first version of the BWA was released for general use in November 2007. Since then, the number of BWA users has grown, and the purposes for which they use it have diversified. The BWA is now widely used for genetic counselling purposes with more than 3000 registered users based in >50 countries.
This report
In this paper, we first describe the update to the UK cancer incidences and the extensions to the BOADICEA model to include cancer incidences from other regions. We then describe updates to the distributions of tumour pathology characteristics using new data on BRCA1 and BRCA2 mutation carriers and women with breast cancer from the general population obtained from the largest data sets currently available. We also describe improvements to the computational efficiency of the algorithm so that risk calculations now run substantially faster, and a new version of the BWA (termed here BWA v3) that incorporates these new features and additional updates to make it easier to use in a clinical setting. We discuss cancer risk predictions and mutation carrier probabilities generated using the updated model. Finally, we summarise conclusions and prospects for future work.
Materials and methods
The underlying genetic model
Details of the underlying statistical model in BOADICEA have been described previously (Antoniou et al, 2008a; Mavaddat et al, 2010). Briefly, the breast cancer incidence for individual i at age t was assumed to be birth cohort specific and to depend on the underlying BRCA1 or BRCA2 genotype and phenotype through a model in the form  , where
, where  is the baseline incidence for the cohort,
is the baseline incidence for the cohort,  represents the major gene effect at age t (i.e., the age-specific log-relative risks associated with BRCA1 and BRAC2 mutations, with
represents the major gene effect at age t (i.e., the age-specific log-relative risks associated with BRCA1 and BRAC2 mutations, with  for non-mutation carriers), and where
for non-mutation carriers), and where  is the polygenic effect assumed to be normally distributed with mean 0 and variance
is the polygenic effect assumed to be normally distributed with mean 0 and variance  . The polygenic variance and BRCA1 and BRCA2 log-relative risks were estimated previously (Antoniou et al, 2008a) and are assumed to remain the same in all the extensions presented. To obtain the baseline incidences, the cohort-specific breast cancer incidences over all assumed genetic effects are constrained to agree with the assumed population incidences (Antoniou et al, 2001) that have been updated as described below. In all our extensions we assumed that the BRCA1 and BRCA2 mutation frequencies remain as previously estimated (Antoniou et al, 2008a). The genetic model can therefore be fully specified. BOADICEA incorporates tumour phenotypes by treating breast cancer subtypes as different disease end points (Mavaddat et al, 2010). For example, in the case of ER status, breast cancer is divided into ER-negative and ER-positive disease with incidences that depend on the underlying genetic effects. To obtain the ER-specific incidences for each genotype in the model an extra constraint is imposed such that the overall incidence over ER status, major gene (BRCA1 carriers, BRCA2 carriers and noncarriers) and polygenic effects agrees with the population breast cancer incidences (Mavaddat et al, 2010). This requires knowledge of the age-specific distributions of breast cancer subtypes in BRCA1 and BRCA2 mutation carriers and the general population.
. The polygenic variance and BRCA1 and BRCA2 log-relative risks were estimated previously (Antoniou et al, 2008a) and are assumed to remain the same in all the extensions presented. To obtain the baseline incidences, the cohort-specific breast cancer incidences over all assumed genetic effects are constrained to agree with the assumed population incidences (Antoniou et al, 2001) that have been updated as described below. In all our extensions we assumed that the BRCA1 and BRCA2 mutation frequencies remain as previously estimated (Antoniou et al, 2008a). The genetic model can therefore be fully specified. BOADICEA incorporates tumour phenotypes by treating breast cancer subtypes as different disease end points (Mavaddat et al, 2010). For example, in the case of ER status, breast cancer is divided into ER-negative and ER-positive disease with incidences that depend on the underlying genetic effects. To obtain the ER-specific incidences for each genotype in the model an extra constraint is imposed such that the overall incidence over ER status, major gene (BRCA1 carriers, BRCA2 carriers and noncarriers) and polygenic effects agrees with the population breast cancer incidences (Mavaddat et al, 2010). This requires knowledge of the age-specific distributions of breast cancer subtypes in BRCA1 and BRCA2 mutation carriers and the general population.
Incidence update
We have updated the BOADICEA model to include UK population cancer incidences for more recent calendar periods using data from two sources: (1) data for the period 1960–1992 were obtained from CI5 (Ferlay et al, 2010) (reported for periods: 1960–1962, 1963–1967, 1968–1972, 1973–1977, 1978–1982, 1983–1987 and 1988–1992); and (2) data for the years 1992–2010 were obtained from the Office for National Statistics (2011). Both sets of incidences were reported in 5-year age intervals up to the age of 84 years (0–4, 5–9, 10–14 and so on). As in Antoniou et al (2008a), the cancer incidences were assumed to be calendar period and cohort specific. In previous versions of the BOADICEA model, we assumed only five birth cohorts. We have now extended this to eight birth cohorts in order to capture the incidences more accurately for those born more recently. For each birth cohort (<1919, 1920–1929, 1930–1939, 1940–1949, 1950–1959, 1960–1969, 1970–1979 and >1980) we derived lifetime incidences by assuming that each individual was born at the midpoint of the relevant birth cohort (1915 for the first cohort and 1985 for the last cohort) and that at each age they experience the relevant calendar period incidences. Incidences before 1960 were assumed to be the same as for the period 1960–1962. Incidences post 2010 were assumed to be the same as for 2010. As in the original BOADICEA model (Antoniou et al, 2008a), we assumed that the relative risks associated with BRCA1 and BRCA2 mutations and the polygenic variances were the same for each birth cohort. However, the absolute risks of disease for each underlying genotype in the model were not the same because the incidences over all genetic effects were constrained to agree with the population incidences.
Published incidences are reported in 5-year age intervals that can result in large variations between adjacent age intervals. As it is plausible that incidences vary continuously with age, we smoothed the population incidences using a locally weighted regression technique (LOWESS) (Cleveland, 1979) with a bandwidth of 0.2. We smoothed the population incidences using the STATA statistical software (StataCorp, College Station, TX, USA, Stata Statistical Software: Release 11, 2009), as described by Antoniou et al (2008a).
To make the model more specific for populations outside the United Kingdom, we compiled incidences for other geographical regions, including Australia, Canada, Denmark, Finland, Iceland, New Zealand, Norway, Sweden and the United States. Data for these regions were obtained from a combination of CI5 and CI5 plus (Ferlay et al, 2010), NORDCAN (Engholm et al, 2010, 2013) as well as national bodies (Australian Institute of Health and Welfare (AIHW), 2011; Surveillance Epidemiology and End Results (SEER) Program, 2011a, 2011b, 2011c; Statistics Canada, 2012; Lewis, 2013). The BOADICEA model was originally developed using data reported for families of European ancestry. As cancer incidences vary by ethnic background, we derived incidences for the United States using only data reported for US whites, and for New Zealand using data from the non-Maori population. In all cases incidences were reported in 5-year age intervals.
We derived smoothed calendar period- and cohort-specific incidences for these regions in the same manner as for the United Kingdom, for the same 8 birth cohorts. We assumed that relative risks of all cancers conferred by BRCA1 and BRCA2, relative to the population incidences, was the same for each country. However, this still allowed for the absolute cancer risks conferred by BRCA1 and BRCA2 to vary between countries. We also assumed that the polygenic variance was the same for each country. As part of these updates, we also modified the BFP so that we can now easily extend the BOADICEA model to include cancer incidences from other countries at the request of users, if appropriate data are available.
Pathology proportions
We obtained age-specific proportions of ER and TN tumour status for BRCA1 and BRCA2 mutation carriers from CIMBA data previously described by Mavaddat et al (2012). For this purpose, we used an updated data set from CIMBA that included ER status information on 3832 BRCA1 mutation carriers and 2399 BRCA2 mutation carriers. Of those who had ER-negative tumours, a total of 1582 BRCA1 mutation carriers and 231 BRCA2 mutation carriers, had information on breast cancer TN status.
All mutation carriers were of self-reported European ancestry. We derived the corresponding distributions of tumour characteristics in the general population using country-matched data from the Breast Cancer Association (Blows et al, 2010; Broeks et al, 2011; Garcia-Closas et al, 2013), based on participants of European ancestry. We used the distributions of the CK5/6 and CK14 tumour markers among TN tumours reported by Mavaddat et al (2010) because of the lack of information on these markers in both CIMBA and BCAC. We used 5-year age intervals for ER status in the general population. However, we used longer age intervals for BRCA1 and BRCA2 mutation carriers and for the TN distributions to ensure that we obtained robust estimates of these proportions. As it is reasonable to assume that the age-specific proportions of ER-positive and TN tumours vary continuously with age, we smoothed these parameters using LOWESS (Cleveland, 1979) with a bandwidth of 0.2.
Optimisation
Previous versions of the BOADICEA model have used the MENDEL pedigree analysis software package (Lange et al, 1988) to calculate likelihoods on pedigrees. In order to exploit fully the predictive potential of genetic models, it is necessary to add the explicit effects of known breast cancer susceptibility variants. However, if we were to use MENDEL to compute the pedigree likelihoods under these circumstances, it would result in an exponential increase in runtime. This presents a substantial problem in terms of the future computational requirements of the BOADICEA program. In addition, the MENDEL FORTRAN source codes used in previous versions of the BOADICEA model were implemented in accordance with the FORTRAN 77 standard that has now been superseded by more recent standards.
In order to improve the computational efficiency of the BFP and thereby facilitate future extensions of the model, we have optimised the algorithm in a number of ways:
(1) We have converted the source code from FORTRAN 77 to FORTRAN 95, and restructured it into modules. The restructured code is easier to read, and simpler to maintain and extend. It has also been designed to allow easy parallelisation of the code in future versions. The FORTRAN 95 module structure allows data abstraction and enforces automatic interfacing, reducing the possibility of programming errors. The conversion of the source codes to FORTRAN 95 has made dynamic memory allocation possible that has enabled us to implement the algorithm in a more resource-efficient manner.
(2) We profiled the code and found that the majority of the runtime was consumed by array multiplication. To address this, where possible we now perform array multiplication using the Basic Linear Algebra Subprograms (BLAS) libraries (Lawson et al, 1979; Dongarra et al, 1990a, 1990b). This has reduced the runtime of the program by a factor of 3.5. We also noted that a significant proportion of the runtime of the code was spent repeatedly calculating a small number of variables. As a result, we modified the code so that these variables were calculated once at the beginning of the program, stored in a lookup table and retrieved when required. This further reduced the runtime of the code by a factor of 1.7, giving an overall decrease by a factor of approximately 6. The relative change in runtime is dependent on the specific computer being used and the pedigree being processed. Here we used a desktop computer with an Intel Xeon E5630 processor and a test pedigree consisting of a dendritic three generational family with 12 people.
(3) Furthermore, we found that these modifications to the FORTRAN source code also resulted in more effective compiler optimisation. In recent years, we have compiled previous versions of the BOADICEA program with the GNU FORTRAN compiler (http://www.gnu.org) (version 4.4.3). However, as part of this study, we conducted benchmark tests to compare the performance of BOADICEA program executables compiled with the GNU FORTRAN compiler and Intel FORTRAN compiler (http://www.intel.co.uk) (version 11.1). These tests showed that when we compiled the previous version of the program with the Intel FORTRAN compiler, the runtime was reduced by a factor of approximately 1.05 (relative to that of an equivalent executable built by compiling the same code with the GNU FORTRAN compiler). However, when we compiled the latest version of the program (with the modifications described above) using the Intel FORTRAN compiler, the runtime of the code was reduced by a factor of approximately 1.37 (relative to that of an equivalent executable built by compiling the same code with the GNU FORTRAN compiler). Relative runtime changes are for the same pedigree and computer as in (2) above.
Web interface updates
We have now implemented BWA v3 in order to accommodate extensions to the BOADICEA model presented here, and to implement modifications requested by users to make the program easier to use in a clinical setting. BWA v3 makes the latest version of the BOADICEA model easily accessible to health-care professionals and members of the public.
Use of pathology and cancer incidence data
BWA v3 now enables users to include pathology information in risk calculations. The BWA enables users to either build a pedigree online for processing or to upload a text file containing one or more pedigrees for processing. When users build an input pedigree online, the program now prompts for details of ER, PR, HER2, CK14 and CK5/6 status. Similarly, we have extended the BOADICEA import/export format (described in Appendix A of the BWA v3 user guide: https://pluto.srl.cam.ac.uk/bd3/v3/docs/BWA_v3_user_guide.pdf) so that users can include the new pathology parameters in the data files that they upload for processing.
BWA v3 allows users to select cancer incidences from a number of countries. The program also includes the most up-to-date UK incidences, but users can still select UK cancer incidences used in previous versions of the BWA for backward compatibility.
Batch processing
Previous versions of the BWA enabled users to either build or upload a single input pedigree. BWA v3 now includes a batch processing module that enables users to upload and process multiple pedigree data sets in a single processing run. When the user uploads a text file that contains multiple pedigrees, BWA v3 now initiates a batch processing job to validate and process the pedigrees sequentially. When the processing job has run to completion, the user can download the computed results across a secure web connection.
Displaying equivalent baseline predicted risks
In recent years, clinical geneticists have requested a means to plot BOADICEA breast and ovarian cancer risks with equivalent baseline cancer risks (i.e., the equivalent predicted cancer risks for a random female of the same age from the general population) to help them communicate the significance of the computed risks to patients. To meet this requirement, BWA v3 now plots BOADICEA breast and ovarian cancer risks and equivalent baseline cancer risks in graphs. BWA v3 also includes additional updates requested by clinical geneticists to make the program easier to use in a clinical setting (e.g., improved pedigree building and pedigree data validation functions).
Checking for breaks in pedigree trees
The BWA allows users to upload input pedigrees for processing. However, in the past, we have encountered problems when a single pedigree has included multiple disjoint family trees or disconnected individuals. As a result, BWA v3 now checks that all family members within a single pedigree are genealogically connected to the index before it is processed.
Results
Incidences
We derived up-to-date calendar- and cohort-specific incidences for breast, ovarian, prostate and pancreatic cancer for the United Kingdom and a number of other regions. For each cancer site, gender and region, we derived eight sets of incidences corresponding to different birth cohort periods, which were then smoothed. The effect of smoothing on breast cancer incidence for a UK female born in 1975 is demonstrated in Figure 1C. The resulting smoothed incidences capture the locality of age-specific incidences before smoothing. Figure 1C also shows a comparison of the updated incidences and previous BOADICEA incidences. The corresponding absolute breast cancer risks for a 30-year-old UK female born in 1975 are shown in Figure 1D. The updated breast cancer incidences are higher at all ages. The remaining risk by age 80 years is 11.4% under the updated model as compared with 9.2% based on the previous implementation.
As expected, differences in the population-specific cancer incidences result in differences in the absolute risks of developing breast or ovarian cancer predicted by the BOADICEA model. This is demonstrated in Figure 1D that shows how the risk of breast cancer predicted by BOADICEA varies by country (for a 30-year-old female born in 1975). Table 1 shows the population-based lifetime risks for all cancer sites considered in BOADICEA for some of the regions now available in BWA v3.
Pathology proportions
We derived age-specific pathology proportions for ER and TN status for the general population, BRCA1 mutation carriers and BRCA2 mutation carriers (Tables 2 and 3). The proportion of ER-negative tumours among BRCA1 mutation carriers decreased with increasing age at diagnosis, whereas the proportion of ER-negative tumours in BRCA2 mutation carriers increased with increasing age at diagnosis, in contrast to the decreasing trend in tumours in the general population seen in the BCAC data Table 1. The proportions of TN tumours among those individuals with an ER-negative tumour were estimated to be 88% and 75.8% in BRCA1 and BRCA2 mutation carriers, respectively. The corresponding proportion of TN tumours in the general population decreased with increasing age at diagnosis from 65.8% for ages <30 years to 57.8% for those diagnosed with ER-negative breast cancer at age ⩾40 years. In Figure 2, the updated distributions of ER and TN status for BRCA2 mutation carriers and the general population are compared with those described in the original publication.
(A) The age-specific proportions of ER-negative tumours used in BOADICEA for BRCA2 mutation carriers. The previous version used data from SEER for the general population, whereas the new version uses data from CIMBA, specific to BRCA2 mutation carriers. (B) The age-specific proportions for TN tumours among ER-negative tumours for the general population. The previous version of BOADICEA used an age constant proportion derived from BCLC data, whereas the new version uses an age-specific proportion derived from BCAC data.
Discussion
In this study, we have described updates and further extensions to the BOADICEA model for genetic susceptibility to breast cancer. These included extensions to the input parameters of the underlying genetic model, to the algorithm and to the BWA. Changes to the model input parameters included the use of up-to-date cancer incidences, new population-specific incidences and updates to the distributions of tumour pathology characteristics using large data sets from CIMBA and BCAC. We have improved the computational efficiency of the algorithm so that BOADICEA risk calculations now run substantially faster. These updates also make possible future extensions of BOADICEA to model more complex genetic effects. In addition, we have implemented BWA v3 so that the latest version of the BOADICEA model is easily accessible to health-care professionals and members of the public. BWA v3 also includes several modifications requested by clinical geneticists to make the software easier to use in a clinical setting.
A consequence of including more up-to-date cancer incidences is that, given the same input, cancer risks predicted by the new BOADICEA model are higher than those predicted by previous versions. This is to be expected because population cancer incidences have increased over time (in particular, the incidence of breast cancer). The observed increase in breast cancer incidences over time is known to be driven by the increased prevalence of screening. These changes in incidences could explain results from two recently published studies that evaluated BOADICEA. A Swedish study reported a slight underestimation (which was not statistically significant) in the predicted number of invasive breast cancers for the first version of BOADICEA (Stahlbom et al, 2012). However, a more recent prospective study that evaluated the BOADICEA model with the updated cancer incidences presented here has demonstrated that the updated version is well calibrated for predicting first invasive breast cancers overall and for most age and family history subgroups (Macinnis et al, 2013). Furthermore, the same study suggested that the updated model has good discriminatory accuracy with an estimated area under the receiver-operating characteristic curve of 0.70 (95% CI: 0.66–0.75).
In extending the model, we are implicitly assuming that the genetic loci (both BRCA1/2 and the variants comprising the polygenic component) continue to confer the same relative risk in a screened population. There are no direct data to substantiate this, but there is evidence that the common SNPs associated with invasive breast cancer are also associated with DCIS (Easton et al, 2007; Michailidou et al, 2013). In the future, we hope to incorporate the effect of screening into BOADICEA more explicitly. Similarly, we assumed that the BRCA1 and BRCA2 relative risks associated with different cancer sites relative to the population incidences remained the same across birth cohorts and across populations. These assumptions are consistent with the observation that the absolute risk of breast cancer in carriers increases with more recent year of birth, in line with the increased risk observed for breast cancer in the general population, and with evidence of variation in the absolute cancer risks by country (Antoniou et al, 2003; Simchoni et al, 2006; Milne et al, 2008). The assumptions are also consistent with a model in which the risks of cancer in carriers are modified by lifestyle risk factors in an approximately multiplicative model. A multiplicative model for the combined effects of lifestyle risk factors and the established common variants has been shown to fit well (i.e., no evidence for GxE interaction), but has not been shown directly for most breast cancer risk factors, although we have shown elsewhere that oral contraceptive use confers a similar (protective) association for ovarian cancer risk in BRCA1/2 mutation carriers as in the general population (Antoniou et al, 2009). Moreover, the absolute risks of breast cancer in BRCA1/2 mutation carriers have been shown to increase with more recent year of birth (Antoniou et al, 2003).
By incorporating cancer incidences from different countries, we also assumed that the distribution of the breast cancer polygenic component remains the same across populations. This assumption would be violated where polygenic effects exist because of loci conferring small effects on risk (e.g., the common breast cancer susceptibility alleles) were different across populations. However, recent results from large genome-wide association studies (based on data from international consortia) did not demonstrate any significant heterogeneity in the associations of common breast cancer susceptibility alleles across populations of European ancestry (Michailidou et al, 2013). However, it is possible that the polygenic distribution differs between populations of different ethnic ancestry; it is known that some loci confer different risks in women of East Asian ancestry, and the frequencies of many of the risk SNPs differ markedly by ethnicity. In addition, BRCA1 and BRCA2 mutation frequencies also vary between populations, and previous studies have shown that BOADICEA may not be well calibrated in populations of non-European ancestry (Thirthagiri et al, 2008). Extending the model to non-European populations, particularly from Asia and Africa, may therefore require additional modifications and perhaps re-estimation of the model parameters. Although not currently implemented, the use of different baseline incidences also provides a basis for incorporating the effect of nongenetic risk factors, such as reproductive history, HRT use and mammographic density into the model.
A previous version of the BOADICEA model incorporated tumour pathology information based primarily on data from BCLC that included only 182 BRCA1, 62 BRCA2 mutation carriers and 109 controls (Mavaddat et al, 2010). Because of limited data, accurate estimates of the age-specific distributions of tumour marker information could not be obtained. Further information on the distribution of ER tumour status in BRCA2 mutation carriers had been scarcer. We had therefore assumed that the distribution of ER status for BRCA2 mutation carriers was the same as in the general population. Recent analyses based on CIMBA data (Mavaddat et al, 2012) however suggest that the proportion of ER-negative tumours in BRCA2 mutation carriers increases with increasing age at diagnosis. Such differences could potentially improve the predictive ability of the model. Using data from CIMBA and BCAC, we were able to include age-specific distributions of ER and TN status for mutation carriers and the general population. A recent evaluation of BOADICEA for its ability to predict BRCA1 and BRCA2 carrier status showed that the updated BOADICEA model incorporating tumour pathology information, based on the CIMBA/BCAC data distributions, provided a significant improvement in discrimination and re-classification over the previous BOADICEA model without pathology, and that the updated BOADICEA model also performed better than the previous BOADICEA model that incorporated tumour pathology information based primarily on the BCLC data (Fischer et al, 2013). These results suggest that the updated BOADICEA model is a valid tool for use in genetic counselling.
The inclusion of tumour pathology in BOADICEA affects both the predicted mutation carrier probabilities, as demonstrated by Mavaddat et al (2010), and the predicted risks of developing breast or ovarian cancer. Figure 3A shows the predicted remaining lifetime risks of breast cancer by age 80 years for a healthy female aged 30 years, born in 1975, depending on the age at breast cancer diagnosis and tumour characteristics of her mother. BOADICEA is the only model to: (1) use tumour pathology information from affected relatives of the proband and (2) use tumour pathology information for cancer risk predictions. This is a unique feature among current breast cancer risk models. (Tai et al, 2008; Evans et al, 2009).
As part of this work, we have implemented BWA v3 to make the updated BOADICEA model easily accessible to health-care professionals and members of the public. During the course of the BOADICEA project, clinical geneticists have provided important feedback on the usability of the program. As a result, BWA v3 also includes additional modifications to make the software easier to use in a clinical setting.
In summary, we present updates to the BOADICEA breast and ovarian cancer risk prediction model that result in more accurate cancer risk predictions and that make the software easier to use in a clinical setting. However, it will be important to evaluate the updated BOADICEA model in prospective studies for its ability to predict future cancer risks. Our current research focusses on extending BOADICEA to include the explicit effects of SNPs known to be associated with breast or ovarian cancer risk, the effects of other breast or ovarian cancer risk factors and modelling the residual genetic susceptibility to ovarian cancer not due to BRCA1 or BRCA2.
Change history
21 January 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Antoniou A, Pharoah PD, Narod S, Risch HA, Eyfjord JE, Hopper JL, Loman N, Olsson H, Johannsson O, Borg A, Pasini B, Radice P, Manoukian S, Eccles DM, Tang N, Olah E, Anton-Culver H, Warner E, Lubinski J, Gronwald J, Gorski B, Tulinius H, Thorlacius S, Eerola H, Nevanlinna H, Syrjakoski K, Kallioniemi OP, Thompson D, Evans C, Peto J, Lalloo F, Evans DG, Easton DF (2003) Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case Series unselected for family history: a combined analysis of 22 studies. Am J Hum Genet 72: 1117–1130.
Antoniou AC, Cunningham AP, Peto J, Evans DG, Lalloo F, Narod SA, Risch HA, Eyfjord JE, Hopper JL, Southey MC, Olsson H, Johannsson O, Borg A, Pasini B, Radice P, Manoukian S, Eccles DM, Tang N, Olah E, Anton-Culver H, Warner E, Lubinski J, Gronwald J, Gorski B, Tryggvadottir L, Syrjakoski K, Kallioniemi OP, Eerola H, Nevanlinna H, Pharoah PD, Easton DF (2008a) The BOADICEA model of genetic susceptibility to breast and ovarian cancers: updates and extensions. Br J Cancer 98: 1457–1466.
Antoniou AC, Hardy R, Walker L, Evans DG, Shenton A, Eeles R, Shanley S, Pichert G, Izatt L, Rose S, Douglas F, Eccles D, Morrison PJ, Scott J, Zimmern RL, Easton DF, Pharoah PD (2008b) Predicting the likelihood of carrying a BRCA1 or BRCA2 mutation: validation of BOADICEA, BRCAPRO, IBIS, Myriad and the Manchester scoring system using data from UK genetics clinics. J Med Genet 45: 425–431.
Antoniou AC, Pharoah PD, McMullan G, Day NE, Ponder BA, Easton D (2001) Evidence for further breast cancer susceptibility genes in addition to BRCA1 and BRCA2 in a population-based study. Genet Epidemiol 21: 1–18.
Antoniou AC, Pharoah PP, Smith P, Easton DF (2004) The BOADICEA model of genetic susceptibility to breast and ovarian cancer. Br J Cancer 91: 1580–1590.
Antoniou AC, Rookus M, Andrieu N, Brohet R, Chang-Claude J, Peock S, Cook M, Evans DG, Eeles R, Nogues C, Faivre L, Gesta P, van Leeuwen FE, Ausems MG, Osorio A, Caldes T, Simard J, Lubinski J, Gerdes AM, Olah E, Furhauser C, Olsson H, Arver B, Radice P, Easton DF, Goldgar DE (2009) Reproductive and hormonal factors, and ovarian cancer risk for BRCA1 and BRCA2 mutation carriers: results from the International BRCA1/2 Carrier Cohort Study. Cancer Epidemiol Biomarkers Prev 18: 601–610.
Australian Institute of Health and Welfare (AIHW) (2011) Australian Cancer Incidence and Mortality (ACIM) Books. AIHW: Canberra.
Blows FM, Driver KE, Schmidt MK, Broeks A, van Leeuwen FE, Wesseling J, Cheang MC, Gelmon K, Nielsen TO, Blomqvist C, Heikkila P, Heikkinen T, Nevanlinna H, Akslen LA, Begin LR, Foulkes WD, Couch FJ, Wang X, Cafourek V, Olson JE, Baglietto L, Giles GG, Severi G, McLean CA, Southey MC, Rakha E, Green AR, Ellis IO, Sherman ME, Lissowska J, Anderson WF, Cox A, Cross SS, Reed MW, Provenzano E, Dawson SJ, Dunning AM, Humphreys M, Easton DF, Garcia-Closas M, Caldas C, Pharoah PD, Huntsman D (2010) Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival: a collaborative analysis of data for 10,159 cases from 12 studies. PLoS Med 7: e1000279.
Broeks A, Schmidt MK, Sherman ME, Couch FJ, Hopper JL, Dite GS, Apicella C, Smith LD, Hammet F, Southey MC, Van 't Veer LJ, de GR, Smit VT, Fasching PA, Beckmann MW, Jud S, Ekici AB, Hartmann A, Hein A, Schulz-Wendtland R, Burwinkel B, Marme F, Schneeweiss A, Sinn HP, Sohn C, Tchatchou S, Bojesen SE, Nordestgaard BG, Flyger H, Orsted DD, Kaur-Knudsen D, Milne RL, Perez JI, Zamora P, Rodriguez PM, Benitez J, Brauch H, Justenhoven C, Ko YD, Hamann U, Fischer HP, Bruning T, Pesch B, Chang-Claude J, Wang-Gohrke S, Bremer M, Karstens JH, Hillemanns P, Dork T, Nevanlinna HA, Heikkinen T, Heikkila P, Blomqvist C, Aittomaki K, Aaltonen K, Lindblom A, Margolin S, Mannermaa A, Kosma VM, Kauppinen JM, Kataja V, Auvinen P, Eskelinen M, Soini Y, Chenevix-Trench G, Spurdle AB, Beesley J, Chen X, Holland H, Lambrechts D, Claes B, Vandorpe T, Neven P, Wildiers H, Flesch-Janys D, Hein R, Loning T, Kosel M, Fredericksen ZS, Wang X, Giles GG, Baglietto L, Severi G, McLean C, Haiman CA, Henderson BE, Le ML, Kolonel LN, Alnaes GG, Kristensen V, Borresen-Dale AL, Hunter DJ, Hankinson SE, Andrulis IL, Mulligan AM, O’Malley FP, Devilee P, Huijts PE, Tollenaar RA, van Asperen CJ, Seynaeve CS, Chanock SJ, Lissowska J, Brinton L, Peplonska B, Figueroa J, Yang XR, Hooning MJ, Hollestelle A, Oldenburg RA, Jager A, Kriege M, Ozturk B, van Leenders GJ, Hall P, Czene K, Humphreys K, Liu J, Cox A, Connley D, Cramp HE, Cross SS, Balasubramanian SP, Reed MW, Dunning AM, Easton DF, Humphreys MK, Caldas C, Blows F, Driver K, Provenzano E, Lubinski J, Jakubowska A, Huzarski T, Byrski T, Cybulski C, Gorski B, Gronwald J, Brennan P, Sangrajrang S, Gaborieau V, Shen CY, Hsiung CN, Yu JC, Chen ST, Hsu GC, Hou MF, Huang CS, Anton-Culver H, Ziogas A, Pharoah PD, Garcia-Closas M (2011) Low penetrance breast cancer susceptibility loci are associated with specific breast tumor subtypes: findings from the Breast Cancer Association Consortium. Hum Mol Genet 20: 3289–3303.
Cleveland WS (1979) Robust locally weighted regression and smoothing scatterplots. J Am Stat Assoc 74: 829–836.
Cunningham AP, Antoniou AC, Easton DF (2012) Clinical software development for the Web: lessons learned from the BOADICEA project. BMC Med Inform Decis Mak 12: 30.
Dongarra JJ, Cruz JD, Hammarling S, Duff IS (1990a) Algorithm 679: a set of level 3 basic linear algebra subprograms: model implementation and test programs. ACM Trans Math Softw 16: 18–28.
Dongarra JJ, Du Croz J, Hammarling S, Duff IS (1990b) A set of level 3 basic linear algebra subprograms. ACM Trans Math Softw 16: 1–17.
Easton DF, Pooley KA, Dunning AM, Pharoah PD, Thompson D, Ballinger DG, Struewing JP, Morrison J, Field H, Luben R, Wareham N, Ahmed S, Healey CS, Bowman R, Meyer KB, Haiman CA, Kolonel LK, Henderson BE, Le ML, Brennan P, Sangrajrang S, Gaborieau V, Odefrey F, Shen CY, Wu PE, Wang HC, Eccles D, Evans DG, Peto J, Fletcher O, Johnson N, Seal S, Stratton MR, Rahman N, Chenevix-Trench G, Bojesen SE, Nordestgaard BG, Axelsson CK, Garcia-Closas M, Brinton L, Chanock S, Lissowska J, Peplonska B, Nevanlinna H, Fagerholm R, Eerola H, Kang D, Yoo KY, Noh DY, Ahn SH, Hunter DJ, Hankinson SE, Cox DG, Hall P, Wedren S, Liu J, Low YL, Bogdanova N, Schurmann P, Dork T, Tollenaar RA, Jacobi CE, Devilee P, Klijn JG, Sigurdson AJ, Doody MM, Alexander BH, Zhang J, Cox A, Brock IW, MacPherson G, Reed MW, Couch FJ, Goode EL, Olson JE, Meijers-Heijboer H, van den Ouweland A, Uitterlinden A, Rivadeneira F, Milne RL, Ribas G, Gonzalez-Neira A, Benitez J, Hopper JL, McCredie M, Southey M, Giles GG, Schroen C, Justenhoven C, Brauch H, Hamann U, Ko YD, Spurdle AB, Beesley J, Chen X, Mannermaa A, Kosma VM, Kataja V, Hartikainen J, Day NE, Cox DR, Ponder BA (2007) Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447: 1087–1093.
Elston RC, Stewart J (1971) A general model for the genetic analysis of pedigree data. Hum Hered 21: 523–542.
Engholm, Ferlay, Christensen, Johannesen, Klint, Køtlum, Milter, Ólafsdóttir, Pukkala, Storm (2013) NORDCAN: Cancer Incidence, Mortality, Prevalence and Survival in the Nordic Countries, Version 5.3 (25.04.2013). Association of the Nordic Cancer Registries, Danish Cancer Society.
Engholm G, Ferlay J, Christensen N, Bray F, Gjerstorff ML, Klint A, Kotlum JE, Olafsdottir E, Pukkala E, Storm HH (2010) NORDCAN—a Nordic tool for cancer information, planning, quality control and research. Acta Oncol 49: 725–736.
Evans DG, Lalloo F, Cramer A, Jones EA, Knox F, Amir E, Howell A (2009) Addition of pathology and biomarker information significantly improves the performance of the Manchester scoring system for BRCA1 and BRCA2 testing. J Med Genet 46: 811–817.
Ferlay, Parkin, Curado, Bray, Edwards, Shin, Forman (2010) Cancer Incidence in Five Continents, Volumes I to IX: IARC CancerBase No. 9 [Internet]. International Agency for Research on Cancer: Lyon, France.
Fischer C, Kuchenbacker K, Engel, Zachariae S, Rhiem K, Meindl A, Rahner N, Dikow N, Plendl H, Debatin I, Grimm T, Gadzicki D, Flottmann R, Horvath J, Schrock E, Stock F, Schafer D, Schwaab I, Kartsonaki C, Mavaddat N, Schlegelberger B, Antoniou AC, Schmutzler R (2013) Evaluating the performance of the breast cancer genetic risk models BOADICEA, IBIS, BRCAPRO and Claus for predicting BRCA1/2 mutation carrier probabilities: a study based on 7352 families from the German Hereditary Breast and Ovarian Cancer Consortium. J Med Genet 50: 360–367.
Fuller SH, Millett LI (2011) Computing performance: game over or next level? IEEE Comp 44: 31–38.
Garcia-Closas M, Couch FJ, Lindstrom S, Michailidou K, Schmidt MK, Brook MN, Orr N, Rhie SK, Riboli E, Feigelson HS, Le ML, Buring JE, Eccles D, Miron P, Fasching PA, Brauch H, Chang-Claude J, Carpenter J, Godwin AK, Nevanlinna H, Giles GG, Cox A, Hopper JL, Bolla MK, Wang Q, Dennis J, Dicks E, Howat WJ, Schoof N, Bojesen SE, Lambrechts D, Broeks A, Andrulis IL, Guenel P, Burwinkel B, Sawyer EJ, Hollestelle A, Fletcher O, Winqvist R, Brenner H, Mannermaa A, Hamann U, Meindl A, Lindblom A, Zheng W, Devillee P, Goldberg MS, Lubinski J, Kristensen V, Swerdlow A, Anton-Culver H, Dork T, Muir K, Matsuo K, Wu AH, Radice P, Teo SH, Shu XO, Blot W, Kang D, Hartman M, Sangrajrang S, Shen CY, Southey MC, Park DJ, Hammet F, Stone J, Veer LJ, Rutgers EJ, Lophatananon A, Stewart-Brown S, Siriwanarangsan P, Peto J, Schrauder MG, Ekici AB, Beckmann MW, dos SS, I, Johnson N, Warren H, Tomlinson I, Kerin MJ, Miller N, Marme F, Schneeweiss A, Sohn C, Truong T, Laurent-Puig P, Kerbrat P, Nordestgaard BG, Nielsen SF, Flyger H, Milne RL, Perez JI, Menendez P, Muller H, Arndt V, Stegmaier C, Lichtner P, Lochmann M, Justenhoven C, Ko YD, Muranen TA, Aittomaki K, Blomqvist C, Greco D, Heikkinen T, Ito H, Iwata H, Yatabe Y, Antonenkova NN, Margolin S, Kataja V, Kosma VM, Hartikainen JM, Balleine R, Tseng CC, Berg DV, Stram DO, Neven P, Dieudonne AS, Leunen K, Rudolph A, Nickels S, Flesch-Janys D, Peterlongo P, Peissel B, Bernard L, Olson JE, Wang X, Stevens K, Severi G, Baglietto L, McLean C, Coetzee GA, Feng Y, Henderson BE, Schumacher F, Bogdanova NV, Labreche F, Dumont M, Yip CH, Taib NA, Cheng CY, Shrubsole M, Long J, Pylkas K, Jukkola-Vuorinen A, Kauppila S, Knight JA, Glendon G, Mulligan AM, Tollenaar RA, Seynaeve CM, Kriege M, Hooning MJ, van den Ouweland AM, van Deurzen CH, Lu W, Gao YT, Cai H, Balasubramanian SP, Cross SS, Reed MW, Signorello L, Cai Q, Shah M, Miao H, Chan CW, Chia KS, Jakubowska A, Jaworska K, Durda K, Hsiung CN, Wu PE, Yu JC, Ashworth A, Jones M, Tessier DC, Gonzalez-Neira A, Pita G, Alonso MR, Vincent D, Bacot F, Ambrosone CB, Bandera EV, John EM, Chen GK, Hu JJ, Rodriguez-Gil JL, Bernstein L, Press MF, Ziegler RG, Millikan RM, Deming-Halverson SL, Nyante S, Ingles SA, Waisfisz Q, Tsimiklis H, Makalic E, Schmidt D, Bui M, Gibson L, Muller-Myhsok B, Schmutzler RK, Hein R, Dahmen N, Beckmann L, Aaltonen K, Czene K, Irwanto A, Liu J, Turnbull C, Rahman N, Meijers-Heijboer H, Uitterlinden AG, Rivadeneira F, Olswold C, Slager S, Pilarski R, Ademuyiwa F, Konstantopoulou I, Martin NG, Montgomery GW, Slamon DJ, Rauh C, Lux MP, Jud SM, Bruning T, Weaver J, Sharma P, Pathak H, Tapper W, Gerty S, Durcan L, Trichopoulos D, Tumino R, Peeters PH, Kaaks R, Campa D, Canzian F, Weiderpass E, Johansson M, Khaw KT, Travis R, Clavel-Chapelon F, Kolonel LN, Chen C, Beck A, Hankinson SE, Berg CD, Hoover RN, Lissowska J, Figueroa JD, Chasman DI, Gaudet MM (2013) Genome-wide association studies identify four ER negative-specific breast cancer risk loci. Nat Genet 45: 392.
Hayat MJ, Howlader N, Reichman ME, Edwards BK (2007) Cancer statistics, trends, and multiple primary cancer analyses from the Surveillance, Epidemiology, and End Results (SEER) Program. Oncologist 12: 20–37.
Lakhani SR, Van De Vijver MJ, Jacquemier J, Anderson TJ, Osin PP, McGuffog L, Easton DF (2002) The pathology of familial breast cancer: predictive value of immunohistochemical markers estrogen receptor, progesterone receptor, HER-2, and p53 in patients with mutations in BRCA1 and BRCA2. J Clin Oncol 20: 2310–2318.
Lange K, Weeks D, Boehnke M (1988) Programs for pedigree analysis: MENDEL, FISHER, and dGENE. Genet Epidemiol 5: 471–472.
Lawson CL, Hanson RJ, Kincaid DR, Krogh FT (1979) Basic linear algebra subprograms for Fortran usage. ACM Trans Math Softw 5: 308–323.
Lewis C New Zealand Cancer Incidences (9-7-2013).
Macinnis RJ, Bickerstaffe A, Apicella C, Dite GS, Dowty JG, Aujard K, Phillips KA, Weideman P, Lee A, Terry MB, Giles GG, Southey MC, Antoniou AC, Hopper JL (2013) Prospective validation of the breast cancer risk prediction model BOADICEA and a batch-mode version BOADICEACentre. Br J Cancer 109: 1296–1301.
Mavaddat N, Barrowdale D, Andrulis IL, Domchek SM, Eccles D, Nevanlinna H, Ramus SJ, Spurdle A, Robson M, Sherman M, Mulligan AM, Couch FJ, Engel C, McGuffog L, Healey S, Sinilnikova OM, Southey MC, Terry MB, Goldgar D, O'Malley F, John EM, Janavicius R, Tihomirova L, Hansen TV, Nielsen FC, Osorio A, Stavropoulou A, Benitez J, Manoukian S, Peissel B, Barile M, Volorio S, Pasini B, Dolcetti R, Putignano AL, Ottini L, Radice P, Hamann U, Rashid MU, Hogervorst FB, Kriege M, van der Luijt RB, Peock S, Frost D, Evans DG, Brewer C, Walker L, Rogers MT, Side LE, Houghton C, Weaver J, Godwin AK, Schmutzler RK, Wappenschmidt B, Meindl A, Kast K, Arnold N, Niederacher D, Sutter C, Deissler H, Gadzicki D, Preisler-Adams S, Varon-Mateeva R, Schonbuchner I, Gevensleben H, Stoppa-Lyonnet D, Belotti M, Barjhoux L, Isaacs C, Peshkin BN, Caldes T, de la Hoya M, Canadas C, Heikkinen T, Heikkila P, Aittomaki K, Blanco I, Lazaro C, Brunet J, Agnarsson BA, Arason A, Barkardottir RB, Dumont M, Simard J, Montagna M, Agata S, D'Andrea E, Yan M, Fox S, Rebbeck TR, Rubinstein W, Tung N, Garber JE, Wang X, Fredericksen Z, Pankratz VS, Lindor NM, Szabo C, Offit K, Sakr R, Gaudet MM, Singer CF, Tea MK, Rappaport C, Mai PL, Greene MH, Sokolenko A, Imyanitov E, Toland AE, Senter L, Sweet K, Thomassen M, Gerdes AM, Kruse T, Caligo M, Aretini P, Rantala J, von WA, Henriksson K, Steele L, Neuhausen SL, Nussbaum R, Beattie M, Odunsi K, Sucheston L, Gayther SA, Nathanson K, Gross J, Walsh C, Karlan B, Chenevix-Trench G, Easton DF, Antoniou AC (2012) Pathology of breast and ovarian cancers among BRCA1 and BRCA2 mutation carriers: results from the Consortium of Investigators of Modifiers of BRCA1/2 (CIMBA). Cancer Epidemiol Biomarkers Prev 21: 134–147.
Mavaddat N, Rebbeck TR, Lakhani SR, Easton DF, Antoniou AC (2010) Incorporating tumour pathology information into breast cancer risk prediction algorithms. Breast Cancer Res 12: R28.
Meijers-Heijboer H, van den Ouweland A, Klijn J, Wasielewski M, de SA, Oldenburg R, Hollestelle A, Houben M, Crepin E, van Veghel-Plandsoen M, Elstrodt F, van DC, Bartels C, Meijers C, Schutte M, McGuffog L, Thompson D, Easton D, Sodha N, Seal S, Barfoot R, Mangion J, Chang-Claude J, Eccles D, Eeles R, Evans DG, Houlston R, Murday V, Narod S, Peretz T, Peto J, Phelan C, Zhang HX, Szabo C, Devilee P, Goldgar D, Futreal PA, Nathanson KL, Weber B, Rahman N, Stratton MR (2002) Low-penetrance susceptibility to breast cancer due to CHEK2(*)1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat Genet 31: 55–59.
Michailidou K, Hall P, Gonzalez-Neira A, Ghoussaini M, Dennis J, Milne RL, Schmidt MK, Chang-Claude J, Bojesen SE, Bolla MK, Wang Q, Dicks E, Lee A, Turnbull C, Rahman N, Fletcher O, Peto J, Gibson L, Dos Santos Silva I, Nevanlinna H, Muranen TA, Aittomäki K, Blomqvist C, Czene K, Irwanto A, Liu J, Waisfisz Q, Meijers-Heijboer H, Adank M, van der Luijt RB, Hein R, Dahmen N, Beckman L, Meindl A, Schmutzler RK, Müller-Myhsok B, Lichtner P, Hopper JL, Southey MC, Makalic E, Schmidt DF, Uitterlinden AG, Hofman A, Hunter DJ, Chanock SJ, Vincent D, Bacot F, Tessier DC, Canisius S, Wessels LF, Haiman CA, Shah M, Luben R, Brown J, Luccarini C, Schoof N, Humphreys K, Li J, Nordestgaard BG, Nielsen SF, Flyger H, Couch FJ, Wang X, Vachon C, Stevens KN, Lambrechts D, Moisse M, Paridaens R, Christiaens MR, Rudolph A, Nickels S, Flesch-Janys D, Johnson N, Aitken Z, Aaltonen K, Heikkinen T, Broeks A, Veer LJ, van der Schoot CE, Guénel P, Truong T, Laurent-Puig P, Menegaux F, Marme F, Schneeweiss A, Sohn C, Burwinkel B, Zamora MP, Perez JI, Pita G, Alonso MR, Cox A, Brock IW, Cross SS, Reed MW, Sawyer EJ, Tomlinson I, Kerin MJ, Miller N, Henderson BE, Schumacher F, Le Marchand L, Andrulis IL, Knight JA, Glendon G, Mulligan AM, Lindblom A, Margolin S, Hooning MJ, Hollestelle A, van den Ouweland AM, Jager A, Bui QM, Stone J, Dite GS, Apicella C, Tsimiklis H, Giles GG, Severi G, Baglietto L, Fasching PA, Haeberle L, Ekici AB, Beckmann MW, Brenner H, Müller H, Arndt V, Stegmaier C, Swerdlow A, Ashworth A, Orr N, Jones M, Figueroa J, Lissowska J, Brinton L, Goldberg MS, Labrèche F, Dumont M, Winqvist R, Pylkäs K, Jukkola-Vuorinen A, Grip M, Brauch H, Hamann U, Brüning T, Radice P, Peterlongo P, Manoukian S, Bonanni B, Devilee P, Tollenaar RA, Seynaeve C, van Asperen CJ, Jakubowska A, Lubinski J, Jaworska K, Durda K, Mannermaa A, Kataja V, Kosma VM, Hartikainen JM, Bogdanova NV, Antonenkova NN, Dörk T, Kristensen VN, Anton-Culver H, Slager S, Toland AE, Edge S, Fostira F, Kang D, Yoo KY, Noh DY, Matsuo K, Ito H, Iwata H, Sueta A, Wu AH, Tseng CC, Van Den Berg D, Stram DO, Shu XO, Lu W, Gao YT, Cai H, Teo SH, Yip CH, Phuah SY, Cornes BK, Hartman M, Miao H, Lim WY, Sng JH, Muir K, Lophatananon A, Stewart-Brown S, Siriwanarangsan P, Shen CY, Hsiung CN, Wu PE, Ding SL, Sangrajrang S, Gaborieau V, Brennan P, McKay J, Blot WJ, Signorello LB, Cai Q, Zheng W, Deming-Halverson S, Shrubsole M, Long J, Simard J, Garcia-Closas M, Pharoah PD, Chenevix-Trench G, Dunning AM, Benitez J, Easton DF (2013) Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat Genet 45: 353–361.
Milne RL, Osorio A, Cajal TR, Vega A, Llort G, de la Hoya M, Diez O, Alonso MC, Lazaro C, Blanco I, Sanchez-de-Abajo A, Caldes T, Blanco A, Grana B, Duran M, Velasco E, Chirivella I, Cardenosa EE, Tejada MI, Beristain E, Miramar MD, Calvo MT, Martinez E, Guillen C, Salazar R, San RC, Antoniou AC, Urioste M, Benitez J (2008) The average cumulative risks of breast and ovarian cancer for carriers of mutations in BRCA1 and BRCA2 attending genetic counseling units in Spain. Clin Cancer Res 14: 2861–2869.
National Institute for Health and Care Excellence (2013) Familial breast cancer: classification and care of people at risk of familial breast cancer and management of breast cancer and related risks in people with a family history of breast cancer (CG164).
Quinn M, Babb P, Brock A, Kirby L, Jones J (2001) Cancer Trends in England and Wales 1950-1999: Studies on Medical and Population Subjects No. 66. The Office for National Statistics, The Stationary Office: London, UK.
Office for National Statistics Cancer Statistics Registrations, England, Series MB1 (2011).
Ontario Breast Screening Program Ontario Breast Screening Program (Canada) (2012).
Parkin DM, Bray F, Ferlay J, Pisani P (2005) Global cancer statistics, 2002. CA Cancer J Clin 55: 74–108.
Rahman N, Seal S, Thompson D, Kelly P, Renwick A, Elliott A, Reid S, Spanova K, Barfoot R, Chagtai T, Jayatilake H, McGuffog L, Hanks S, Evans DG, Eccles D, Easton DF, Stratton MR (2007) PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nat Genet 39: 165–167.
Renwick A, Thompson D, Seal S, Kelly P, Chagtai T, Ahmed M, North B, Jayatilake H, Barfoot R, Spanova K, McGuffog L, Evans DG, Eccles D, Easton DF, Stratton MR, Rahman N (2006) ATM mutations that cause ataxia-telangiectasia are breast cancer susceptibility alleles. Nat Genet 38: 873–875.
Riley BD, Culver JO, Skrzynia C, Senter LA, Peters JA, Costalas JW, Callif-Daley F, Grumet SC, Hunt KS, Nagy RS, McKinnon WC, Petrucelli NM, Bennett RL, Trepanier AM (2012) Essential elements of genetic cancer risk assessment, counseling, and testing: updated recommendations of the National Society of Genetic Counselors. J Genet Couns 21: 151–161.
Seal S, Thompson D, Renwick A, Elliott A, Kelly P, Barfoot R, Chagtai T, Jayatilake H, Ahmed M, Spanova K, North B, McGuffog L, Evans DG, Eccles D, Easton DF, Stratton MR, Rahman N (2006) Truncating mutations in the Fanconi anemia J gene BRIP1 are low-penetrance breast cancer susceptibility alleles. Nat Genet 38: 1239–1241.
Simchoni S, Friedman E, Kaufman B, Gershoni-Baruch R, Orr-Urtreger A, Kedar-Barnes I, Shiri-Sverdlov R, Dagan E, Tsabari S, Shohat M, Catane R, King MC, Lahad A, Levy-Lahad E (2006) Familial clustering of site-specific cancer risks associated with BRCA1 and BRCA2 mutations in the Ashkenazi Jewish population. Proc Natl Acad Sci USA 103: 3770–3774.
Smith RA, Cokkinides V, Brawley OW (2012) Cancer screening in the United States, 2012: a review of current American Cancer Society guidelines and current issues in cancer screening. CA Cancer J Clin 62 (2): 129–142.
Stahlbom AK, Johansson H, Liljegren A, von WA, Arver B (2012) Evaluation of the BOADICEA risk assessment model in women with a family history of breast cancer. Fam Cancer 11: 33–40.
Statistics Canada Table 103-0550 - New cases for ICD-O-3 primary sites of cancer (based on the July 2011 CCR tabulation file), by age group and sex, Canada, provinces and territories, CANSIM (database) (2012).
Surveillance Epidemiology and End Results (SEER) Program Limited Use Data (1973-2006) www.seer.cancer.gov (2006).
Surveillance Epidemiology and End Results (SEER) Program SEER*Stat Database: Incidence - SEER 13 Regs Research Data, Nov 2011 Sub (1992-2009) <Katrina/Rita Population Adjustment> - Linked To County Attributes - Total U.S (2011a).
Surveillance Epidemiology and End Results (SEER) Program SEER*Stat Database: Incidence - SEER 18 Regs Research Data, Nov 2011 Sub (2000-2009) <Katrina/Rita Population Adjustment> - Linked To County Attributes - Total U.S (2011b).
Surveillance Epidemiology and End Results (SEER) Program SEER*Stat Database: Incidence - SEER 9 Regs Research Data, Nov 2011 Sub (1973-2009) <Katrina/Rita Population Adjustment> - Linked To County Attributes - Total U.S (2011c).
Tai YC, Chen S, Parmigiani G, Klein AP (2008) Incorporating tumor immunohistochemical markers in BRCA1 and BRCA2 carrier prediction. Breast Cancer Res 10: 401.
Thirthagiri E, Lee SY, Kang P, Lee DS, Toh GT, Selamat S, Yoon SY, Taib NA, Thong MK, Yip CH, Teo SH (2008) Evaluation of BRCA1 and BRCA2 mutations and risk-prediction models in a typical Asian country (Malaysia) with a relatively low incidence of breast cancer. Breast Cancer Res 10: R59.
Acknowledgements
This work was funded by Cancer Research UK Grants C12292/A11174 and C1287/A10118. ACA is a Cancer Research – UK Senior Cancer Research Fellow. CIMBA and BCAC are funded by Cancer Research UK Grants (C12292/A11174, C1287/A10118, C1287/A12014) and by the European Community’s Seventh Framework Programme under grant agreement no. 223175 (HEALTH-F2-2009-223175) (COGS) and NIH 1R01CA128978-01A2. NM was funded by a scholarship from the Medical Research Council. This work was supported by the Governement of Canada through Genome Canada and the Canadian Institutes of Health Research, and the Ministère de l'enseignement supérieur, de la recherche, de la science et de la technologie du Québec through Génome Québec.We thank Robert MacInnis, Per Hall and Mikael Eriksson for their helpful conversations and CIMBA and BCAC for access to the data.
Author information
Authors and Affiliations
Consortia
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Lee, A., Cunningham, A., Kuchenbaecker, K. et al. BOADICEA breast cancer risk prediction model: updates to cancer incidences, tumour pathology and web interface. Br J Cancer 110, 535–545 (2014). https://doi.org/10.1038/bjc.2013.730
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.730
Keywords
This article is cited by
-
Validation of the BOADICEA model for predicting the likelihood of carrying pathogenic variants in eight breast and ovarian cancer susceptibility genes
Scientific Reports (2023)
-
Regulating cancer risk prediction: legal considerations and stakeholder perspectives on the Canadian context
Human Genetics (2023)
-
A pilot study investigating feasibility of mainstreaming germline BRCA1 and BRCA2 testing in high-risk patients with breast and/or ovarian cancer in three tertiary Cancer Centres in Ireland
Familial Cancer (2023)
-
Randomized trial of surveillance with abbreviated MRI in women with a personal history of breast cancer– impact on patient anxiety and cancer detection
BMC Cancer (2022)
-
Familial history and prevalence of BRCA1, BRCA2 and TP53 pathogenic variants in HBOC Brazilian patients from a public healthcare service
Scientific Reports (2022)