Abstract
Förster resonance energy transfer (FRET) has been widely used in biological and biomedical research because it can determine molecule or particle interactions within a range of 1–10 nm. The sensitivity and efficiency of FRET strongly depend on the distance between the FRET donor and acceptor. Historically, FRET assays have been used to quantitatively deduce molecular distances. However, another major potential application of the FRET assay has not been fully exploited, that is, the use of FRET signals to quantitatively describe molecular interactive events. In this review, we discuss the use of quantitative FRET assays for the determination of biochemical parameters, such as the protein interaction dissociation constant (Kd), enzymatic velocity (kcat) and Km. We also describe fluorescent microscopy-based quantitative FRET assays for protein interaction affinity determination in cells as well as fluorimeter-based quantitative FRET assays for protein interaction and enzymatic parameter determination in solution.
Similar content being viewed by others
Quantitative FRET analysis
The phenomena and principles of FRET were first discovered in 1948. In groundbreaking studies in the early 1970s, the quantitative relationship between FRET efficiency and the orientation and overlapping spectrum of the two fluorophores was described in biological macromolecules. This finding established the use of FRET as a spectroscopic ruler and bio-analytic tool1,2,3,4,5,6. Since then, FRET-based techniques have been extensively used in biological research in various settings to identify molecular interactive events in vitro and in vivo7,8,9.
The FRET signal strength is generally determined by two major factors: the intrinsic FRET efficiency (E) of the donor and acceptor and the amounts of the interactive donor and acceptor. Moreover, the FRET efficiency depends strongly on the distance between the two fluorescent moieties, and this property is often used to measure the distance between donor and acceptor. The FRET efficiency is represented by the following:

where R0 is the Förster radius, the distance between donor and acceptor chromophores, when the efficiency of energy transfer is 50%, and

where Q0 is the quantum yield of the donor chromophore in the absence of acceptor and n is the refractive index of the medium. The orientation factor (κ2) depends on the relative orientation of the donor and acceptor dipoles and is given by the following:

where α is the angle between the donor and acceptor transition moments, β is the angle between the donor moment and the line joining the centers of the donor and acceptor, and γ is the angle between the acceptor moment and the line joining the centers of the donor and acceptor. J is the spectral overlap integral, which is calculated as follows:

where F(λ) is the fluorescent intensity of the donor chromophore at wavelength λ and (λ) is the molar extinction coefficient of the acceptor chromophore.
For a specific FRET pair, the FRET efficiency is fixed in a static state, and the FRET signal is proportional to the amount of interactive donor and acceptor complex. Therefore, it can be used to quantitatively describe the protein interaction. Specifically, the assessment of the absolute FRET signal is critical for such a determination. However, for most FRET pairs, the spectral overlap between the donor and acceptor that allows FRET to occur is also the cause of FRET signal contamination: the direct excitation of the acceptor by donor excitation wavelength and the direct emission of the donor at the acceptor emission wavelength.
In recent decades, several methods have been developed to quantify FRET signals, including the “three-cube” FRET measurement for fluorescence microscopy/spectroscopy10, signal bleed-through (SBT)11, the stepwise algorithm to remove SBT contamination12, photobleaching13, and the more common ratiometric method14. Among these approaches, the three-cube FRET measurement is considered a general approach and has consequently attracted the most attention.
Determining the protein Kd by fluorescent microscopy and “three-cube” FRET measurements
The first effort to determine the protein interaction dissociation constant, Kd, in cells using the quantitative FRET assay was carried out by Prof David YUE with a fluorescent microscope imaging approach and three-cube FRET measurements15,16. This approach is based on the biophysical analysis of the fluorescent excitation and emission properties of the individual free donor, free acceptor, and bound donor-acceptor complex. The three-cube method for FRET signal analysis was first used to estimate the total amount of bound donor-acceptor complex. A FRET ratio, FR (the total emission of acceptor divided by direct emission at FRET wavelength), was introduced as a variable in the FRET signal that corresponds to the amount of bound complex. The predicted FR (FRpredicted) was estimated from the total amount of bound complex.
In a multiple variable regression process to minimize the squared error (FRexp-FRpredicted)2, a relative dissociation constant, Kd,EFF, was estimated as a fluorescent unit of the free donor and acceptor divided by total amount of bound complex. The Kd,EFF can be converted to a Kd when the cell volume and fluorescent signals of individual labeled fluorescent protein are considered. However, this methodology requires that several parameters be determined by measuring the purified donor and acceptor fluorescent proteins and assumes that the spectra of purified proteins and proteins in cells are equivalent. Detailed knowledge of the optical filter characteristics, eg, the average molar extinction coefficient of donor and acceptor fluorophores over the bandwidth of the FRET cube excitation filter, is also required.
Subsequently, several studies have been conducted to improve the process of imaging-based FRET assay for Kd determination17,18,19. These improvements mainly focus on the mathematic algorithms, and all of them require that the FRET efficiency is first determined. From the FRET efficiency, multi-variable mathematic models were developed to determine the fraction of bound complex and free fractions of donor and acceptor to determine the dissociation constant, Kd. These methods employed different approaches to measure the FRET efficiency. For example, Chen et al directly measured the FRET efficiency by photobleaching imaging, which did not require recombinant proteins to obtain standard curves18. Mehta et al developed a computational imaging method to eliminate imaging noise and include endogenous unlabeled proteins in the process of Kd determination by FRET imaging19. However, the accurate determination of FRET efficiency is challenging because the outcome may depend on the method used to determine the FRET efficiency. As such, the resultant Kd may depend on the measurement strategy20.
The cross-wavelength co-efficiency method
Martin and colleagues were the first to attempt to measure a protein interaction dissociation constant using a biochemistry approach21. Specifically, they conducted an in vitro experiment at a fixed concentration of donor, CFP-SUMO1, with increasing amounts of acceptor, YFP-Ubc9, to obtain titrated signals. Both the steady-state binding curves in the FRET signal and protein concentration were determined. To eliminate signal contamination from the direct emission of donor and acceptor, they established external standard curves of mixtures of fluorescent proteins alone or fluorescent proteins with only one interactive partner. Although this approach is straightforward in concept, it requires multiple experiments to correct for the direct emission of donor and acceptor and is consequently tedious. Because the fluorescent proteins used to determine the standard curve differed from those used in Kd measurement, the purity and fluorescent emission spectra may differ by experiment.
The spectrum crosstalk between the donor and acceptor is a major challenge for quantitative FRET analysis. The donor emission spectrum can overlap with the acceptor emission spectrum, and the acceptor may also be directly excited at the donor excitation wavelength. Over the years, many experimental efforts have attempted to circumvent these challenges. For example, fluorophores exhibiting a large Stokes shift, such as quantum dots, have been used to reduce the crosstalk22,23,24. Alternatively, time-resolved FRET assays with fluorophores that exhibit long half-lives, such as Terbium-based dyes, have proven effective in eliminating interfering fluorescence from unwanted sources25,26,27. Although these methods greatly reduce spectrum crosstalk in the FRET assay and simplify FRET signal analysis, more generalized approaches are needed when fluorophores with significant spectrum crosstalk are used, such as for intracellularly expressed recombinant proteins labeled with fluorescent proteins.
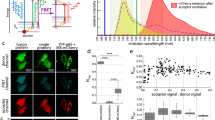
Our group developed a method to mathematically analyze the spectrum crosstalk between the FRET donor and acceptor28. This method is based on a FRET system of the fluorescent proteins CyPet and YPet, a pair that was specifically optimized for FRET efficiencies greater than those of traditional fluorescent protein pairs used in FRET studies, such as ECFP/EYFP29. CyPet and YPet have excitation/emission peak wavelengths of 414 nm/475 nm and 515 nm/530 nm, respectively. When CyPet and YPet are conjugated to two interacting molecules and the mixture is excited at 414 nm, the interaction quenches CyPet emission at 475 nm and increases YPet emission at 530 nm. However, the emission intensity at 530 nm does not linearly correlate with the FRET-excited YPet because CyPet has significant emission at 530 nm when excited at 414 nm, and YPet can be directly excited at 414 nm and emit at 530 nm, independently of FRET. To quantitatively analyze the FRET signal, the FRET emission of YPet needs to be determined, and the interference from the direct emission of unquenched CyPet and total YPet needs to be eliminated. To this end, we introduced a second excitation wavelength of 475 nm, at which only YPet, but not CyPet, is excited (Figure 1).
Quantitative analysis of fluorescence signals of FRET emission. Fluorescent emission at 530 nm at the excitation wavelength of 414 nm (FLDA) can be divided into three components: FRET emission from YPet-Ubc9, direct emission of unquenched CyPet-SUMO1, and direct emission of YPet-Ubc9. Fluorescent emission at 530 nm at the excitation wavelength of 475 nm (FLAA) consists of the direct emission of YPet-Ubc9.
To determine the direct emission components, two cross-wavelength ratiometric factors are defined: the ratio of CyPet emission at 475 nm and 530 nm when excited at 414 nm and the ratio of YPet emission at 530 nm when excited at 414 nm and 475 nm. Because these factors are constants for a given FRET donor and acceptor and a particular imaging setup, they can be determined using purified CyPet or YPet proteins. With these factors, the direct emission of unquenched CyPet at 530 nm can be calculated from the emission at 475 nm when excited at 414 nm, and the direct emission of YPet at 530 nm can be calculated from the emission intensity at 530 nm when excited at 475 nm. The FRET emission intensity of CyPet at 530 nm when excited at 414 nm can then be determined by subtracting the direction emission intensities from the total emission and used in subsequent analyses.
The versatile cross-wavelength co-efficiency method that we have developed for quantitative FRET signal analysis can be applied to many FRET applications for which the donor and acceptor spectra significantly overlap. In this analysis, an emission wavelength at which only the donor emits is used to calculate donor emission at the acceptor emission wavelength, and a second excitation wavelength at which only the acceptor emits is used to calculate the acceptor direction emission. For cases in which these conditions cannot be met, Chen et al developed a similar method to quantitatively analyze the FRET emission18. In their analysis, three filter tubes are used to collect fluorescence images of the donor emission at the donor excitation wavelength, the acceptor emission at the donor excitation wavelength, and the acceptor emission at the acceptor excitation wavelength. These fluorescent intensities are then expressed as functions of four crosstalk parameters and three fluorescence components: donor fluorescence, sensitized acceptor FRET emission, and the direct emission of acceptor. The crosstalk parameters can be determined from samples with only donor or acceptor, and the three fluorescence components can be calculated by a three-variable linear equation group.
The Kd is an important parameter to quantitatively assess the binding affinity of protein-protein interactions. Traditional methods to measure Kd include surface plasmon resonance (SPR) and isothermal titration calorimetry (ITC). Compared with these methods, FRET-based Kd measurement does not require specific instrumentation and can be applied to complex biological environments. Using CFP and YFP as a FRET donor and acceptor, Martin et al developed a FRET-based assay to quantitatively determine the dissociation constant of SUMO1-Ubc9 proteins21. In their assay, CFP and YFP were covalently conjugated to SUMO1 and Ubc9. Recombinant fusion proteins were purified and mixed in vitro. Their fluorescence spectrum was then compared with those from non-interacting control fluorescent protein pairs to determine the sensitized FRET emission from YFP. The FRET emission intensities were plotted against YFP concentrations to calculate the maximal FRET emission, which was then used to convert FRET emission intensities into concentrations of bound YFP-Ubc9 protein. Finally, the concentrations of bound protein were plotted against those of free protein to calculate the value of Kd. This method is effective but requires the use of control proteins and multiple rounds of fitting of raw data.
Our group developed a novel FRET-based assay with a new mathematical formula that correlates the FRET signal with Kd to quantitatively determine the Kd of protein-protein interactions. In this assay, CyPet and YPet were conjugated to SUMO1 and Ubc9, respectively, and the fusion proteins were expressed and purified. Different concentrations of YPet-Ubc9 were titrated into a fixed concentration of CyPet in a 384-well plate, and the fluorescence spectra at the excitation wavelengths of 414 nm or 475 nm was measured by a fluorescence plate reader. The FRET-sensitized emission of YPet-Ubc9 was analyzed using the method we described previously28. We expressed the FRET emission of YPet-Ubc9 as a function of the maximal FRET emission, Kd and concentration of free YPet-Ubc9. Because the free YPet-Ubc9 concentration can also be expressed as a function of Kd based on the definition of Kd, the formula was finally transformed into the following version:

Here, X represents the concentration of total YPet-Ubc9 at each data point, a represents the concentration of total CyPet-SUMO1, and EmFRET and EmFRETmax represent the intensity of FRET sensitized emission at experimental and theoretical maximal concentration of bound YPet-Ubc9, respectively. Because a is a constant in our assay, the FRET-sensitized emission of YPet-Ubc9 can be fitted to the YPet-Ubc9 concentration to derive the value of Kd in one step (Figure 2). We used this method to determine the Kd of SUMO1 and Ubc9 for different CyPet-SUMO1 concentrations and obtained consistent results. We also determined the Kd between CyPet-SUMO1 and YPet-Ubc9 as well as untagged SUMO1 and Ubc9 using SPR. We then compared these Kd values with those obtained from a previously documented ITC assay30 (Table 1). The conjugation of CyPet and YPet does not significantly alter the Kd between SUMO1 and Ubc9, and the FRET-based Kd measurement assay produces results comparable with those obtained using traditional methods. Thus, our assay is an accurate method to measure the Kd of protein-protein interactions. In addition, our FRET-based assay can be conducted in a 384-well plate format, thus making it suitable for high-throughput applications.
Kd determination by FRET-based assay. FRET emission of YPet-Ubc9 was fitted against protein concentration to derive Kd of CyPet-SUMO1 and YPet-Ubc9 in one step.
Quantitative FRET assay of protease kinetics parameters
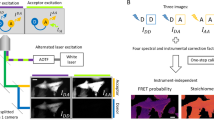
The general strategy for a FRET-based protease assay is based on a fluorescent protein-tagged substrate (Figure 3). The SENP substrate, pre-SUMOs, is flanked by a FRET pair, CyPet and YPet. The fused recombinant protein is cleaved by the protease SENPs to release two products: the CyPet-SUMO and the SUMO tail with YPet, which leads to the disruption of FRET and an increase in the emission of CyPet while dramatical decrease in the emission of YPet after the excitation of CyPet. The decreased fluorescent emission of YPet after cleavage can be used to characterize the kinetic properties of SENPs in real time.
Design of FRET-based protease assay. Schematic depicting the CyPet-(pre-SUMO)-YPet substrate indicating the principle of FRET from CyPet (donor) to YPet (acceptor).
To determine the kinetic parameters of SENP1 with a FRET assay, two challenges must be overcome: determining the absolute FRET signal that corresponds to the concentration of digested substrates and converting the absolute FRET signal into related protein concentrations. For the first challenge, the fluorescence signal of FRET must be distinguished from the direct fluorescence signals of the donor and acceptor at the emission wavelength of the acceptor. The absolute FRET signal will determine the amount of the undigested substrate, excluding the interference of the donor and acceptor direct emissions from both digested and undigested substrates. For the second challenge, standard curves are needed to convert FRET signals to the concentrations of the corresponding proteins.
Before being digested by SENP1c, the total fluorescent emission of CyPet-(pre-SUMO1)-YPet at 530 nm in response to excitation at 414 nm can be divided into three parts: the FRET-induced emission of the acceptor (Ida), the direct emission of the donor (Id530/414) and the direct emission of the acceptor (Ia530/414), as shown in Figure 4:
Quantitative analysis of fluorescent signals.

where Id475/414 is the emission of CyPet at 475 nm in response to excitation at 414 nm and Ia530/475 is the emission of YPet at 530 nm in response to excitation at 475 nm.
After the digestion of the protein by SENP1c, the fluorescent signal at 530 nm decreases and fluorescent signal at 475 nm increases due to the disrupted FRET. The remaining fluorescent emission at 530 nm (FL'530/414) continues to be divisible into the same three parts:

where I'da is the remaining FRET-induced emission of the acceptor and I'd475/414 is the fluorescent emission of CyPet at 475 nm, which can be divided into two parts, the emission from the undigested CyPet-(pre-SUMO1)-YPet and the digested CyPet-SUMO1. I'a530/475 is the fluorescent emission of YPet, which is constant irrespective of substrate digestion. Therefore, the fraction βI'a530/475 remains the same.
After treatment with SENP1c, the remaining FRET-induced emission of the acceptor (I'da) is:

where C is the total concentration of CyPet-(pre-SUMO1)-YPet (before SENP1c is added to the reaction system) and x is the concentration of digested CyPet-(pre-SUMO1)-YPet at different time points.
Standard curves are created by plotting the fluorescent emissions against related protein concentrations. For undigested CyPet-(pre-SUMO1)-YPet, the fluorescence emissions of various concentrations at 475 nm in response to excitation at 414 nm were determined and plotted against the protein concentrations (Figure 5A). For digested CyPet–SUMO1, different concentrations of CyPet–SUMO1 were mixed with YPet at a molar ratio of 1:1, and the emissions at 475 nm with excitation at 414 nm was then determined and plotted against the protein concentrations (Figure 5B). The slopes, k and j, 230800 and 326700 in CyPet and YPet pair, respectively, are two constants that correlate a linear relationship between the detected fluorescent signals and the protein concentrations.
Standard curves of fluorescent signal versus protein concentration. (A) Emission of CyPet-(pre-SUMO1)-YPet at 475 nm under excitation at 414 nm. (B) Emission of CyPet-SUMO1+YPet (1:1 molar ration) under excitation at 414 nm (the x axis is the protein concentration of CyPet-SUMO1).
According to the standard curves,

where Id475/414-csy (y in standard curve) is the emission of CyPet-(pre-SUMO1)-YPet at 475 nm in response to excitation at 414 nm, k is the slope of the standard curve for Id-csy as a function of the concentration of CyPet-(pre-SUMO1)-YPet (μmol/L), Id475/414-cs (z in standard curve) is the emission of CyPet-SUMO1 at 475 nm in response to excitation at 414 nm, and j is the slope of the standard curve for Id-cs as a function of the concentration of CyPet-SUMO1 (μmol/L).
If the above-analyzed items are combined, the detected fluorescent signal at 530 nm in response to excitation at 414 nm for the treatment of the substrate CyPet–(pre- SUMO1)–YPet with protease SENP1c is:

The maturation of the pre-SUMOs by SENP1c can be determined by monitoring the changes in the fluorescent signal at 530 nm during the process. Changes in the absolute FRET signal and other fluorescence components can be analyzed with calculations based on the standard curves. Different concentrations of substrates, ranging from 0.115 to 2.300 μmol/L, were incubated with 0.8 nmol/L of SENP1c. The remaining fluorescence intensity (FL'530/414) was monitored, and the digested substrate (x) was calculated according to the above analysis.

where the digested substrate (or product) concentration increases exponentially from 1 when t=0 to [S]0 at infinite time is:

The original velocities were calculated at different substrate concentrations, as shown in Table 2.
Michaelis–Menten kinetics is one of the simplest and best-known models of enzyme kinetics and is described by the following:

The value of kcat can be directly calculated by dividing the experimentally determined value of vmax by the enzyme concentration, [E]. The Michaelis–Menten graph for the data in Table 3 is plotted in Figure 6.
Michaelis-Menten graphical analysis of CyPet–(pre-SUMO1)–YPet's processing by SENP1c. Data were plotted and analyzed by GraphPad Prism 5 of nonlinear regression.
One-sample measurement of protease kinetics parameters by internal calibration in high-throughput settings
In the standard curve method31, standard curves are generated from the fluorescent reading detected by a fluorometer as a function of the related protein concentration. The slopes are then derived by linearly fitting the fluorescent reading (RFU) to the protein concentration (μmol/L). However, the emission of CyPet at 475 nm can be directly obtained by the fluorometer as I'd475/414. As such, the above equation can be modified as follows31:

During the maturation process of pre-SUMO, the increase in the direct emission of CyPet and the decrease in the emission of the FRET-induced acceptor are due to the disruption of energy transfer. The detected total fluorescent emission at 530 nm, CyPet and YPet direct emission, and the FRET-induced YPet's emission determined by the quantitative analysis in the internal calibration and standard curve-dependent detections are compared in Figure 7. The fluorescent emission detected at 530 nm is not equal to the FRET-induced emission of YPet, which is always considered to be the same in the ratiometric analysis.
Time-course of fluorescence component changes during pre-SUMO maturation. Reactions were monitored within original 5 min and 10-s intervals.
Pre-SUMO1's maturation by SENP1c can be determined by monitoring the changes in the fluorescent signal at 475 nm and 530 nm in response to excitation at 414 nm during the process. Different amounts of the fluorescent substrates were incubated with SENP1c. The concentration of digested substrate, x, was then calculated according to the quantitative FRET analysis in the internal calibration detection (Figure 8).
Concentrations of digested substrate during pre-SUMO maturation process analyzed by the internal calibration method. Reactions were monitored within the original 5 min.
The initial velocity (v0) of the maturation of CyPet-(pre-SUMO1)-YPet by SENP1c was determined using the quantitative FRET analysis for both the internal calibration detection and standard curve-dependent method (Table 4). The results exhibit a good substrate dose-dependent relationship and subtle differences from the above two methods.
The values of KM and Vmax can be obtained from the Michaelis–Menten equation by plotting the various velocities of SENP1c digestion versus different concentrations of the substrates. The initial velocities in Table 4 were plotted using the Michaelis-Menten equation (Figure 9). KM, kcat, and the kcat/KM ratio were then obtained using both the quantitative FRET analysis of the internal calibration and the standard curve-dependent method (Table 5).
Michaelis-Menten graphical analysis of pre-SUMO1's maturation by SENP1c. Data were plotted and analyzed by GraphPad Prism 5 in nonlinear regression (Michaelis-Menten model).
The kcat/KM values from the quantitative FRET analysis are similar to those derived from the ratiometric FRET analysis in one of the previous SENP1 endopeptidase functional studies32. However, the quantitative FRET-derived KM and kcat did not approximate previously reported values. As expected, the kinetic parameters derived from the internal calibration and standard curve-dependent methods are very similar, but the standard errors are larger for the standard curve method.
The fluorescent signals linearly correlate with fluorescent protein concentrations in the standard curves, whereas the slopes are derived to fit a linear relationship of fluorescent readings versus protein concentrations. Variations in the slopes cannot be avoided and consequently produce inaccuracies in the kinetic constant determination. Moreover, the fluorescent emission of proteins used to determine the standard curve varies by batch due to pipetting variations, impure protein samples, maturation level, and variations in determining the protein concentration by Beer's law in the Bradford assay. In addition to these variations, fluorometer-induced variations cannot be ignored in the study of protease kinetics.
According to a review of quantitative FRET analysis10, FRET's efficiency in detecting a FRET signal cannot be repeated in only one channel (the donor with or without the existence of the acceptor), and multiple channels need to be observed to improve the accuracy and robustness of results. Comparisons of the previous standard curve-dependent and the improved internal calibration method (Table 5) indicate that the standard errors of the standard curve-dependent method are larger than those of the internal calibration method.
The improved internal calibration method for quantitative FRET analysis in the protease kinetic study simplifies the operation by omitting the steps to establish a standard curve and also helps to minimize the errors in kinetic constant determination.
Summary
Quantitative FRET assays have been improved to determine a series of biochemical parameters in real time. These new developments have several advantages over other traditional biochemical and biophysical approaches for these biochemical parameter determinations. First, these approaches can accurately determine biochemical parameters. The quantifications of FRET signals and their associated molecular events are very precise. As verified in several cases, the Kd values determined by quantitative FRET are comparable to those determined by traditional approaches, such as SPR or ITC. In addition, FRET signal changes can be monitored in real time – a considerable advantage over other methods, such as radioactive labeling and Western blotting. Second, these approaches are straightforward and easily adapted by laboratories in both academia and industry. Fluorescent proteins have been widely used in biological and biomedical research, and the instrumentation for FRET assays is available to most researchers. The fluorescent protein expression techniques are also easily adapted to most laboratories. Third, these methods are environmentally friendly. Because only fluorescent proteins are used, other environmental hazards, such as radioisotopes, are not needed. Fourth, quantitative FRET-based technologies for biomedical parameter determination can be easily converted to a high-throughput format. All of our assays are conducted in 384-well plates, and the throughput can easily be increased further. Moreover, these quantitative FRET assays can replace traditional approaches that require complex instrumentation. Overall, these methodologies can be readily used for quantitative systems biology research and pharmacological applications.
References
Steinberg IZ . Long–range nonradiative transfer of electronic excitation energy in proteins and polypeptides. Annu Rev Biochem 1971; 40: 83–114.
Stryer L . Fluorescence energy transfer as a spectroscopic ruler. Annu Rev Biochem 1978; 47: 819–46.
Giepmans BN, Adams SR, Ellisman MH, Tsien RY . The fluorescent toolbox for assessing protein location and function. Science 2006; 312: 217–24.
Li IT, Pham E, Truong K . Protein biosensors based on the principle of fluorescence resonance energy transfer for monitoring cellular dynamics. Biotechnol Lett 2006; 28: 1971–82.
Wu P, Brand L . Resonance energy transfer: methods and applications. Anal Biochem 1994; 218: 1–13.
Hillisch A, Lorenz M, Diekmann S . Recent advances in FRET: distance determination in protein–DNA complexes. Curr Opin Struct Biol 2001; 11: 201–7.
Miyawaki A . Development of probes for cellular functions using fluorescent proteins and fluorescence resonance energy transfer. Annu Rev Biochem 2011; 80: 357–73.
Grecco HE, Verveer PJ . FRET in cell biology: still shining in the age of super–resolution? Chemphyschem 2011; 12: 484–90.
Jares–Erijman EA, Jovin TM . FRET imaging. Nat Biotechnol 2003; 21: 1387–95.
Gordon GW, Berry G, Liang XH, Levine B, Herman B . Quantitative fluorescence resonance energy transfer measurements using fluorescence microscopy. Biophys J 1998; 74: 2702–13.
Ruiz–Velasco V, Ikeda SR . Functional expression and FRET analysis of green fluorescent proteins fused to G–protein subunits in rat sympathetic neurons. J Physiol 2001; 537: 679–92.
Wallrabe H, Elangovan M, Burchard A, Periasamy A, Barroso M . Confocal FRET microscopy to measure clustering of ligand–receptor complexes in endocytic membranes. Biophys J 2003; 85: 559–71.
Wallrabe H, Periasamy A . Imaging protein molecules using FRET and FLIM microscopy. Curr Opin Biotechnol 2005; 16: 19–27.
Horton RA, Strachan EA, Vogel KW, Riddle SM . A substrate for deubiquitinating enzymes based on time–resolved fluorescence resonance energy transfer between terbium and yellow fluorescent protein. Anal Biochem 2007; 360: 138–43.
Erickson MG, Alseikhan BA, Peterson BZ, Yue DT . Preassociation of calmodulin with voltage–gated Ca2+ channels revealed by FRET in single living cells. Neuron 2001; 31: 973–85.
Erickson MG, Liang H, Mori MX, Yue DT . FRET two–hybrid mapping reveals function and location of L–type Ca2+ channel CaM preassociation. Neuron 2003; 39: 97–107.
Raicu V, Jansma DB, Miller RJ, Friesen JD . Protein interaction quantified in vivo by spectrally resolved fluorescence resonance energy transfer. Biochem J 2005; 385: 265–77.
Chen H, Puhl HL 3rd, Ikeda SR . Estimating protein–protein interaction affinity in living cells using quantitative Forster resonance energy transfer measurements. J Biomed Opt 2007; 12: 054011.
Mehta K, Hoppe AD, Kainkaryam R, Woolf PJ, Linderman JJ . A computational approach to inferring cellular protein–binding affinities from quantitative fluorescence resonance energy transfer imaging. Proteomics 2009; 9: 5371–83.
Berney C, Danuser G . FRET or no FRET: a quantitative comparison. Biophys J 2003; 84: 3992–4010.
Martin SF, Tatham MH, Hay RT, Samuel ID . Quantitative analysis of multi–protein interactions using FRET: application to the SUMO pathway. Protein Sci 2008; 17: 777–84.
Zhong W . Nanomaterials in fluorescence–based biosensing. Anal Bioanal Chem 2009; 394: 47–59.
Grigsby CL, Ho YP, Leong KW . Understanding nonviral nucleic acid delivery with quantum dot–FRET nanosensors. Nanomedicine (Lond) 2012; 7: 565–77.
Medintz IL, Mattoussi H . Quantum dot–based resonance energy transfer and its growing application in biology. Phys Chem Chem Phys 2009; 11: 17–45.
Hu LA, Zhou T, Hamman BD, Liu Q . A homogeneous G protein–coupled receptor ligand binding assay based on time–resolved fluorescence resonance energy transfer. Assay Drug Dev Technol 2008; 6: 543–50.
Lebakken CS, Riddle SM, Singh U, Frazee WJ, Eliason HC, Gao Y, et al. Development and applications of a broad–coverage, TR–FRET–based kinase binding assay platform. J Biomol Screen 2009; 14: 924–35.
Kim B, Tarchevskaya SS, Eggel A, Vogel M, Jardetzky TS . A time–resolved fluorescence resonance energy transfer assay suitable for high–throughput screening for inhibitors of immunoglobulin E–receptor interactions. Anal Biochem 2012; 431: 84–9.
Song Y, Madahar V, Liao J . Development of FRET assay into quantitative and high–throughput screening technology platforms for protein–protein interactions. Ann Biomed Eng 2011; 39: 1224–34.
Nguyen AW, Daugherty PS . Evolutionary optimization of fluorescent proteins for intracellular FRET. Nat Biotechnol 2005; 23: 355–60.
Tatham MH, Chen Y, Hay RT . Role of two residues proximal to the active site of Ubc9 in substrate recognition by the Ubc9.SUMO–1 thiolester complex. Biochemistry 2003; 42: 3168–79.
Liu Y, Song Y, Madahar V, Liao J . Quantitative Forster resonance energy transfer analysis for kinetic determinations of SUMO–specific protease. Anal Biochem 2012; 422: 14–21.
Shen L, Tatham MH, Dong C, Zagórska A, Naismith JH, Hay RT . SUMO protease SENP1 induces isomerization of the scissile peptide bond. Nat Struct Mol Biol 2006; 13: 1069–77.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liao, Jy., Song, Y. & Liu, Y. A new trend to determine biochemical parameters by quantitative FRET assays. Acta Pharmacol Sin 36, 1408–1415 (2015). https://doi.org/10.1038/aps.2015.82
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2015.82
Keywords
This article is cited by
-
Develop quantitative FRET (qFRET) technology as a high-throughput universal assay platform for basic quantitative biomedical and translational research and development
Med-X (2023)
-
Live imaging of apoptotic signaling flow using tunable combinatorial FRET-based bioprobes for cell population analysis of caspase cascades
Scientific Reports (2022)
-
An in vitro Förster resonance energy transfer-based high-throughput screening assay identifies inhibitors of SUMOylation E2 Ubc9
Acta Pharmacologica Sinica (2020)
-
Advanced FRET normalization allows quantitative analysis of protein interactions including stoichiometries and relative affinities in living cells
Scientific Reports (2019)
-
Protein–Protein Affinity Determination by Quantitative FRET Quenching
Scientific Reports (2019)