Abstract
Three-dimensional, protein-based hydrogel scaffolds that successfully mimic in vivo extracellular matrix microenvironments are desirable for tissue engineering and regenerative medicine applications, and can provide highly capable in vitro tissue analogues. However, the fabrication of protein-based scaffolds with uniform porosity, thin walls and durable mechanical properties remains a challenging prospect that might be overcome by integrating advances in microfabrication and protein functionalization. Towards this goal, herein, we report the successful fabrication of a highly ordered, gelatin-based inverted colloidal crystal (ICC) hydrogel platform that is robust and supports high levels of cell function. In particular, the utilization of colloidal templating microfabrication strategies together with highly substituted, photocrosslinkable gelatin methacryloyl (GelMA) allowed us to fabricate protein-based three-dimensional scaffolds with uniform pore interconnectivity, structural stability and tailorable degradation properties. The resulting GelMA ICC scaffolds provided cell attachment sites and promoted intercellular interaction of hepatocytes, which resulted in improved cell function compared to a flat 2D system. The results demonstrate the potential of GelMA ICC scaffolds to become an effective tissue engineering platform for drug screening and regenerative medicine.
Similar content being viewed by others
Introduction
In the human body, cells reside in three-dimensional (3D) extracellular matrix (ECM) environments, and a large number of tissue engineering studies have demonstrated that mimicking this environment in engineered 3D systems allows resident cells to exhibit in vivo-like behaviour in terms of cell morphology and function.1, 2, 3 Such capabilities represent key advances in the field because, when using conventional strategies like two-dimensional (2D) substrates, cultured cells (for example, primary hepatocytes) rapidly lose their functional capacity as well as their morphology.4 On the other hand, 3D tissue engineering strategies such as sandwich and spheroid cultures along with hydrogel and microfluidic platforms have demonstrated excellent potential for overcoming these challenges and highlight the merits of applying materials science approaches to design biomaterials for biological applications.5, 6, 7
Among the design strategies, ICC hydrogel scaffolds are one of the most promising 3D systems and possess uniform pores that are hexagonally arranged and interconnected.8 The ICC configuration is typically prepared from a base polymeric material and a sacrificial colloidal crystal, and the size of the pores is tunable which allows excellent control over nutrient and oxygen diffusion.9 Such capabilities have proven advantageous across a wide range of tissue engineering fields, including liver tissue engineering for which ICC scaffolds have enabled spheroid morphologies to assemble along with long-term viability, and function,10 as well as eliciting an in vivo-like response against nanoparticles.11 In the earliest examples, the ICC scaffolds were made from synthetic polyacrylamide materials that are biologically inert and lack support for important cell–ECM interactions.12, 13 More recently, ECM-mimicking collagen-coated polyethylene glycol diacrylate ICC scaffolds have been developed that allow hepatocytes to attach to the cavity walls, significantly improving both cell–cell and cell–ECM interactions.14, 15 In contrast to bare polyethylene glycol diacrylate ICC scaffolds that promoted the formation of cellular aggregates on account of weak cell-scaffold interactions, the collagen coating on the ECM-mimicking ICC scaffold facilitates the formation of sheet-like cell morphologies led to higher albumin production in a manner that could be controlled by the degree of collagen coating.14 While these findings demonstrate the significance of integrating ICC scaffold designs with protein functionalization, engineering challenges remain, including the fact that synthetic polymers such as polyethylene glycol diacrylate are not biodegradable and the multi-step process to prepare protein-coated ICC scaffolds is cumbersome. As a promising alternative, a fully protein-based ICC hydrogel scaffold would be advantageous, especially if it were possible to construct the scaffold with a biologically active protein acting as the monomer. However, to date, the preparation of protein-based ICC hydrogels had faced significant obstacles arising from low mechanical strength of the scaffold, limited protein solubility and high viscosity during the preparation process, and high cost of the materials.14, 15, 16, 17, 18, 19, 20, 21
ICC scaffolds composed of natural biopolymers, such as alginate,16, 17, 18 chitin19, 20 and chitosan,21, 22, 23 have been reported. As these protein materials have high viscosity in aqueous solution, the protein concentrations must be kept below 5 w/v% during fabrication in order to ensure effective penetration into the ICC lattice interstices. To enhance mechanical strength, these ICC systems are commonly prepared in the presence of ionic or chemical crosslinkers (for example, calcium chloride,16, 17, 18 1-[3-(dimethylamino)-propyl]-3-ethylcarbondiimide hydrochloride17, 18 and genipin19, 23). Ionically crosslinked hydrogels can be formed quickly in an aqueous solution containing divalent cations, and covalently crosslinked hydrogels can be prepared by including chemical agents that link together protein monomers to form a permanent network structure with improved stability. However, the mechanical stability of ionically crosslinked hydrogels in an aqueous solution is compromised by the reversibility of ionic bonds,24 and covalently crosslinked hydrogels require long curing times (1–24 h)17, 18 that render them unsuitable for simultaneous curing and cell encapsulation.25 One of the more promising covalent crosslinking strategies has been realized based on photocrosslinking in which light is used as a cue to promote rapid polymerization of photosensitive reactive monomer units, in turn enabling a high degree of spatiotemporal control over the crosslinking process and facilitating its usage for a wide range of biological applications.25, 26 Indeed, photopolymerization supports the rapid formation of a hydrogel network with greater homogeneity than small molecule-based chemical methods17, 26, 27, 28 and is attractive for exploring within the context of improved protein-based hydrogel designs.
To successfully mimic ECM microenvironments, the selection of protein material is critical and a leading choice is collagen, which is the main structural protein of extracellular matrices.29 Gelatin is a partially hydrolysed form of collagen,30 and offers a practical functional alternative that can be functionalized into photopolymerizable GelMA.31 Importantly, GelMA retains the biological activities of gelatin such as cell attachment and enzymatic degradation,32 and also offers tailorable mechanical properties which have led to its extensive use in 3D bioprinting and many other bioengineering applications.25, 27, 28, 32, 33, 34 For these reasons, GelMA has attracted significant interest as a material for preparing optimized 3D tissue analogues,25 and a porous GelMA scaffold for cell culture applications has been reported.35 The latter work also highlighted the challenges of fabricating porous, protein-based hydrogel scaffolds with hierarchical architectures as well as the importance of establishing robust materials science strategies to achieve the target objective. In particular, the fabrication process utilized soft alginate sacrificial beads that did not form colloidal crystals, and as a result the platform lacked a high degree of structural ordering.
As such, the fabrication of 3D protein-based ICC hydrogel platforms with highly ordered architectures and structural stability remains an outstanding goal and would be useful for preparing 3D replicas of human organs such as the liver.36, 37, 38 In this regard, the liver is one of the most challenging organs to mimic in engineered tissues, and addressing this challenge is important because cellular function is strongly correlated with the architectural arrangement.39 From a design perspective, the microscopic units in which liver cells reside in the body are lobules that are roughly hexagonal in shape,40 and thus ICC platforms arranged hexagonally are an ideal choice. For these reasons, we hypothesized that a 3D GelMA ICC hydrogel platform could be successfully established to mimic the hexagonal lobule structure while retaining ECM-like biological activity, and hence provide a superior 3D environment for culturing liver cells. Based on this goal, herein, we report the production of highly ordered, GelMA ICC hydrogel scaffolds that were engineered using a colloidal templating technique with optimized preparations of highly substituted, photocurable GelMA. Systematic examinations were conducted in terms of structural stability, uniform pore interconnectivity and tailorable degradation properties within the context of important biological applications, including in vitro tissue analogues for liver tissue engineering. To our knowledge, this is the first report of a highly ordered, protein-based ICC hydrogel platform for tissue engineering.
Experimental procedures
Fabrication and degradation of gelatin-based ICC hydrogel scaffolds
GelMA samples with different degrees of substitution (DS) were synthesized as seen in Figure 1a according to the literature.27 Briefly, type A gelatin (175 bloom) was dissolved at 10 w/v% at 60 °C in two buffer systems (phosphate buffered saline (PBS) and 0.1 M carbonate-bicarbonate (CB) buffer). GelMA samples were prepared by the reaction of gelatin with methacrylic anhydride (94%) at a feed ratio of methacrylic anhydride (1 ml) to gelatin (10 g) at 50 °C for 3 h in three different conditions (PBS, CB and CB with pH maintenance at 9.0) in a time-lapse loading manner. After 3 h of reaction, the solutions were readjusted to pH of 7.4, filtered, dialysed using PALL Minimate TFF Capsule (Ann Arbor, MI, USA) with 10 kDa MWCO at 40 °C for 1 day, lyophilized and stored at −20 °C until further use. The DS of GelMA was quantitatively measured by 2,4,6-trinitrobenzene sulfonic acid and verified by 1H-NMR (Avance I 400 MHz, Bruker, Rheinstetten, Germany) in deuterium oxide.
GelMA ICC scaffold preparation. (a) Gelatin methacryloyl (GelMA) synthesis scheme. GelMA synthesis was conducted at a feed ratio of methacrylic anhydride (MAA)/gelatin (0.1 ml per 1 g) in different buffer systems (PBS, 0.1 M CB, and 0.1 M CB with pH 9 adjustment). (b) Storage moduli of GelMA bulk hydrogels (30 w/v% in distilled water with 1 w/v% I2959) as a function of degrees of substitution (DS) of 52, 76 and 98%. (c) Schematic illustration of the fabrication process of GelMA ICC scaffolds by colloidal templating. ICC, inverted colloidal crystal.
Polystyrene beads of 138±2.0 μm diameter (Duke Scientific Corporation, Palo Alto, CA, USA) were self-assembled in 70% ethanol solution in polypropylene moulds of 6 mm diameter by shaking for 2 days and annealed at 134 °C for 6 h to obtain lattices. GelMA samples with different DS were dissolved at various concentrations (5, 10, 20, 30 and 40 w/v%) in distilled water at 50 °C. The solutions of GelMA containing 1% 2-hydroxy-4′-(2-hydroxyethoxy)-2-methylpropiophenone (I2959) were soaked into the lattices by centrifugation at 15 000 r.p.m. at 40 °C for 10 min. The GelMA infiltrated lattices were cured by ultraviolet light (UV; 365 nm at 100 mW cm−2) for 10 min, and then polystyrene beads were removed using tetrahydrofuran. The GelMA ICC scaffolds were sterilized with 70% ethanol solution, washed three times with PBS, and finally stored in distilled water or PBS at 4 °C until further use. The fabricated scaffolds with different concentrations and different DS were evaluated in terms of structural integrity and dimensional stability. All chemicals were purchased from Sigma-Aldrich (St Louis, MO, USA) unless specified.
Rheological measurements
Mechanical properties of aqueous GelMA solutions (30 w/v%) containing I2959 (1 w/v%) were characterized with sinusoidal shear rheometry. Frequency-sweep measurements were conducted using a rheometer (Anton Paar Physica MCR 501, Graz, Austria), equipped with a temperature-controllable glass plate, UV curing system (365 nm, 100 mW cm−2), and a 25 mm cone-plate geometry with an angle of 1°. The measurement conditions were at 2% strain amplitude at an oscillation frequency of 0.1–10 Hz. Similarly, the storage modulus of each GelMA ICC scaffold was measured with a 10 mm diameter parallel-plate geometry at 0.1 % strain and 0.1 Hz. The temperature was maintained at 37 °C throughout the measurements.
Accelerated enzyme degradation study
ICC scaffolds made of GelMA (30 w/v%) with different DS were tested for enzymatic degradation in 1 mg ml−1 of collagenase (125 CDU mg−1 solid) in Hank’s Balanced Salt Solution, containing 3 mM CaCl2. Surface morphology of GelMA ICC scaffolds was observed through an optical microscope during the degradation, and mass loss of GelMA ICC scaffolds was also measured.
Cell culture
Human hepatocellular carcinoma cells (Huh 7.5, Apath, New York, NY, USA) were maintained in Dulbecco’s modified Eagle’s medium (Hyclone, South Logan, UT, USA) with 10% fetal bovine serum (Hyclone) and 1% penicillin/streptomycin (Life Technologies, Carlsbad, CA, USA) in a humidified atmosphere at 37 °C with 5% CO2. The medium was changed every 3 days. Before cell seeding, GelMA ICC scaffolds were placed in 24-well plates and washed with PBS and kept in 2 ml media for 30 min. After aspiration of the media, an amount of 25 μl (1 × 106 cells) was carefully pipetted on top of each GelMA ICC scaffold and incubated for 3 h. The scaffolds were then transferred into a new 24-well plate with each well containing 1 ml medium. As a control, the same number of cells was seeded on 2D GelMA substrates of approximately 2 mm thickness in six-well plates. For both conditions, media were changed every day.
Viability and proliferation assays of Huh-7.5 within the GelMA ICC scaffold
The cell viability in 3D GelMA ICC scaffolds and on 2D GelMA substrates was characterized using Live/Dead Cell Viability/Cytotoxicity kit (Life Technologies). Briefly, 4 μM calcein-acetoxymethyl (calcein-AM) and 8 μM ethidium homodimer-1 (EthD-1) in media were added to the samples and incubated for 1 h at 37 °C. The cytoplasm of live cells and the nuclei of dead cells were stained by calcein-AM (green) and EthD-1 (red), respectively and were observed through a confocal microscope (TCS SP8, Leica, Mannheim, Germany). Numbers of live and dead cells were counted using the ImageJ software package.
Immunostaining and confocal microscopy
Cell-seeded samples were collected at different time points (1, 3, 6 and 9 days) for immunocytochemistry. The samples were washed twice with PBS, fixed with 4% paraformaldehyde for 5 min, permeabilized with 0.1% Triton X-100 for 30 min, washed twice with PBS, and incubated in a 3% bovine serum albumin blocking buffer for 1 h. Albumin and cytochrome P450 (CYP) 3A4 were separately bound with specific primary antibodies (sc-271605 and sc-53850, respectively, Santa Cruz Biotechnology, Santa Cruz, CA, USA) overnight at 4 °C. After being washed three times with PBS, they were incubated with a secondary antibody conjugated with Alexa Flour 555 (Life Technologies). Simultaneously, filamentous actins (f-actin) were stained with Alexa Flour 488 labelled phalloidin (Life Technologies) for 2 h at room temperature. Next, the samples were washed twice with PBS, and then nuclei were stained with 10 μg ml−1 4,6-diamidino-2-phenylindole, dihydrochloride (Life Technologies) for 10 min. Images of stained cells were captured and reconstructed with confocal microscopes (LSM710 with a ZEN program, Carl Zeiss, Oberkochen, Germany, or TCS SP8, Leica).
Western blot assay
Intracellular protein amount at different time points (1, 3, 6 and 9 days) were qualitatively measured by western blot assay. First, total proteins were extracted with the Protein Extraction kit (Life Technologies), and the amount was determined using BCA protein assay kit (Thermo Fisher Scientific, Waltham, MA, USA). The protein samples were boiled with 4 × Laemmli sample buffer (Bio-Rad, Hercules, CA, USA) for 5 min, and 20 μl of each boiled sample was loaded into wells of a 10% polyacrylamide gel. Subsequently, electrophoresis was performed using the Mini PROTEAN III Cell (Bio-Rad). Separated proteins were transferred onto nitrocellulose membrane (Bio-Rad) and stained with Ponceau S staining solution to ascertain the loading process. The stained membranes were washed with a blocking buffer (10 mM Tris-HCl, 150 mM NaCl, 0.1% Tween 20 and 3% nonfat dry milk), and separately incubated for 2 h at room temperature in the blocking buffer with a 1:1000 diluted primary antibodies (albumin: sc-271605, E-cadherin: sc-21791 and beta-actin: sc-47778, Santa Cruz Biotechnology). After being washed with the blocking buffer three times for 15 min, the membrane was probed with a 1:2000 diluted secondary antibody (#1706516, Bio-Rad) for 2 h. The membrane was then washed three times for 15 min and developed with Clarity and Clarity Max Western ECL Blotting Substrates (Bio-Rad). Chemiluminescent signal was detected with a luminescent image analyser (LAS-4000, GE Healthcare Life Sciences, Buckinghamshire, UK).
Gene expression analysis
The expression levels of regulative and functional genes of cells were quantified at different time points (1, 3, 6 and 9 days). First, total RNA was isolated with TRIzol reagent (Life Technologies) and reverse transcribed with primers (listed in Supporting Supplementary Table S1) and iScript Reverse Transcription Supermix (Bio-Rad). The synthesized complementary DNAs were amplified, based on real-time quantitative polymerase chain reaction (RT-qPCR) with the SYBR select Master Mix for CFX (Life Technologies) in the CFX connect Real-Time PCR system (95 °C for 20 s, 40 cycles of 10 s at 95 °C and 40 s at 60 °C). All reactions were run in three times, and 2−ΔΔCT method was used for analysing the data. The value of each gene was normalized against the housekeeping gene, glyceraldehyde 3-phosphate dehydrogenase.
Morphological characterization
To investigate the microscopic morphology, a field emission scanning electron microscope (FESEM; JSM-7600F, JEOL, Tokyo, Japan) was utilized. ICC scaffolds and cell-seeded samples fixed with 4% paraformaldehyde were treated by sequential ethanol dehydration at 25, 50, 75, 95 and 100% for 15 min each. The scaffolds were stored at −80 °C and were lyophilized for 48 h. The scanning electron microscopy samples were sputter-coated (JFC-1600, JEOL) with Pt to a thickness of 10 nm, and the images were taken with an acceleration voltage of 5 kV.
Statistical analysis
Statistical analysis was performed using the Microsoft Excel statistical analysis software package. Comparisons between two treatments were made using a two-tailed pair student’s t-test. One-way ANOVA was used to test for differences among at least three groups. The standard deviation was calculated and presented for each treatment group (mean±s.d.). The level of significance was set at P<0.05 (#, P<0.05; ##, P<0.01; ###, P<0.001).
Results and discussion
As the DS is an important parameter for controlling GelMA crosslink density, mechanical properties and degradation speed, GelMA samples with various DS were synthesized for fabrication of ICC scaffolds as seen in Figures 1a and b.27 GelMA samples with different DS of 52, 76 and 98% were prepared at the feed ratio of methacrylic anhydride (1 ml) to gelatin (10 g) in three different systems (PBS, CB and CB at pH 9), respectively.
To fabricate GelMA ICC scaffolds, colloidal crystals were prepared by the self-assembly of polystyrene beads of 138.1±2.2 μm diameter (Figure 1c). The dried polystyrene beads were subsequently annealed to form template lattices. Next, aqueous GelMA solutions (5, 10, 20, 30 and 40 w/v%) were infiltered to the lattices by centrifuging at 40 ºC. Owing to reduced ionic intermolecular interactions, GelMA is highly water soluble at this temperature and solutions of 30 w/v% had viscosity values of 394±97 mPa s for 52% DS, 191±19 mPa s for 76% DS and 215±27 mPa s for 98% DS at 37 °C at 50 s−1 as shown in Supplementary Figure S1. Furthermore, the bulk stiffness of GelMA hydrogels (30 w/v% in distilled water with 1 w/v% I2959) at 52, 76 and 98% DS were 22, 61 and 99 kPa, respectively. All solutions were observed to penetrate into lattice interstices. After UV crosslinking of GelMA-infiltered lattices, polystyrene templates were removed using tetrahydrofuran and clean GelMA ICC scaffolds were obtained.
Following preparation, we first investigated structural integrity by evaluating mechanical stiffness. GelMA ICC scaffolds made of 5 and 10 w/v% GelMA solutions were mechanically weak regardless of DS owing to use of low concentrations of GelMA that resulted in thin ICC scaffold walls. These samples collapsed easily, and were difficult to manipulate with tweezers as presented in the optical images in Figure 2a. At above 20 w/v% concentrations, GelMA ICC with the highest DS (98%) provided excellent structural integrity, as they were self-supporting and able to withstand manipulation with tweezers (Supplementary Figure S2), whereas those with 52 and 76% DS exhibited such structural integrity only at concentrations above 30 w/v%. Furthermore, the GelMA ICC scaffolds at 30 w/v% were measured for mechanical stiffness, and their storage moduli exhibited a range of 1–4 kPa (Supplementary Figure S3), depending on the DS of GelMA. These results highlighted the ability of combining colloidal crystal templating of high concentrated GelMA and efficient UV crosslinking to overcome the common drawbacks of poor structural integrity and mechanical properties in protein-based hydrogels.
Demonstration of structural integrity of GelMA ICC scaffolds. (a) Optical images of GelMA ICC scaffolds with different GelMA concentrations (5, 10, 20, 30 and 40 w/v%) and different DS (52, 76 and 98%). GelMA ICC scaffolds made from GelMA above 20 w/v% exhibited good structural integrity in different DS samples. (b) The diameter of GelMA ICC scaffolds versus different concentrations. GelMA ICC scaffolds made from GelMA above 20–30 w/v% maintained dimensional stability (n=2; each sample was measured three times in different angles) and showed an easy-to-handle feature. ICC, inverted colloidal crystal; GelMA, gelatin methacryloyl.
Regarding the dimensional size of GelMA ICC scaffolds, GelMA ICC scaffolds with a lower DS were slightly more swollen as shown in Figure 2b. GelMA ICC samples prepared below 20 w/v% easily collapsed when removed from aqueous media due to insufficient mechanical support. The overall dimensions of GelMA ICC scaffolds prepared above 30 w/v% were well maintained. Therefore, GelMA ICC scaffolds of 30 w/v% with three different DS (52, 76 and 98%), exhibiting good structural stability, were used in further investigations. The scanning electron microscopy micrographs and microscope images of GelMA ICC scaffolds (30 w/v%, DS=98%) (Figure 3a and Supplementary Figure S4) show that GelMA ICC scaffolds featured a hexagonal honeycomb-like microstructure with uniform cavities of about 139 μm diameter and connection window channels of about 30 μm at all DS (52, 76 and 98%). GelMA ICC scaffolds displayed six uniform connection window channels in each cavity as presented in the 3D confocal images. In the ICC system, the size of the cavities and the connection channels between cavities are controlled by the size of the self-assembled polystyrene beads and the annealing temperature, respectively, influencing the size of the cell constructs and cell–cell interactions between pores. Uniformity and interconnectivity of pores in ICC scaffolds is also important, not only for diffusion of molecules (oxygen and nutrients) but also for homogeneous cell distribution, eliciting greater control over cellular activities.41 For these reasons, we used 140 μm diameter polystyrene PS beads in order to meet permeability constraints of oxygen and nutrients within the cell constructs (150–200 μm),42 and gently annealed the sacrificial beads so as to achieve highly regular pore formation across all DS (Figure 3b). Using our systematic approach, we found that tightly annealed colloidal crystal templating and a high concentration of photocrosslinkable GelMA solution above 20% are the main important factors to produce well-defined protein-based ICC scaffolds with homogeneous microporosity, interconnectivity and robust mechanical properties.
Micro-scale features of ICC scaffolds made from GelMA (30 w/v%, DS=98%). (a) Optical microscopic images of assembled colloidal crystals made of polystyrene spheres, optical microscopic images of GelMA ICC scaffolds, scanning electron microscopic (SEM) images of GelMA ICC scaffolds, and confocal images of fibronectin-coated GelMA ICC scaffolds via immunohistochemical staining (fibronectin in red). (b) Dimensions of GelMA ICC scaffolds with different DS (52, 76 and 98%). (a) The diameter of colloidal beads (provided by the manufacturer). (b) Diameters of the cavity (n=10; mean±s.d.). (c) Diameters of connection channels (n>140; mean±s.d.). GelMA ICC scaffolds exhibited a highly ordered microporous and interconnected architecture. ICC, inverted colloidal crystal; DS, degree of substitution; GelMA, gelatin methacryloyl.
Tailorable biodegradability is an essential requirement of scaffolds for tissue engineering especially for in vivo applications. Here, we conducted accelerated enzymatic degradation tests of GelMA ICC scaffolds using collagenase at 1 mg ml−1 (125 U ml−1). As presented in Figure 4, the degradation of GelMA ICC scaffolds by the enzyme was apparent, and their degradation speed was highly dependent on the DS of the GelMA base material, which influences ICC crosslink density. The lower the DS of GelMA ICC scaffolds, the faster the degradation. GelMA ICC scaffolds with 52% DS lost a half of their masses within 2 h. Degradation half-lives of GelMA ICC scaffolds of 76 and 98% DS were around 3 and 4 h, respectively. Also, the morphological features of GelMA ICC scaffolds during the degradation were monitored as seen in Figure 4a. The time profile of their morphological changes was in good accordance with that of their mass loss (Figure 4b). The cavity rings of the first layers of GelMA ICC scaffolds with 52 and 76% DS disintegrated after 1 and 2 h, respectively, whereas those of GelMA ICC with 98% DS started to lose their cavity integrity between 4 and 6 h. It has been established that GelMA ICC scaffolds maintain the collagenase cleavage site (-R-Pro-X-Gly-Pro-R-, X: a neutral amino acid) even after the chemical modification process, as methacrylic anhydride reacts mainly with free amino groups of gelatin.25, 32 These results demonstrate that GelMA ICC scaffolds undergo enzymatic degradation even with relatively high concentration (30 w/v%), and that this degradation can be controlled via the designed DS of each GelMA system.
Accelerated enzymatic degradation study of GelMA ICC scaffolds. (a) Micro-scale surface morphology observation of GelMA ICC scaffolds with different DS (scale bar=100 μm) during enzymatic degradation. (b) Mass loss of GelMA ICC scaffolds with different DS at 1 mg ml−1 collagenase Type 1A solution (125 U ml−1) (n=3; mean±s.d.). GelMA ICC scaffolds experienced enzymatic degradation, and GelMA ICC with a lower DS degraded more quickly. ICC, inverted colloidal crystal; GelMA, gelatin methacryloyl.
To understand the effect of the protein-based microarchitecture of 3D GelMA ICC scaffolds on cells, we used Huh7.5 liver cells as surrogate model cells because they are highly available and retain important hepatic functions such as liver-specific gene expression, hepatic enzyme activity and permissibility to HCV infection. Liver cells reside in microscopic lobules that are roughly hexagonal in shape, the ECM structural proteins of which are composed mainly of collagen.29 As GelMA ICC scaffolds mimic the liver lobule microenvironment, we hypothesized that GelMA ICC scaffolds of highly ordered micro-porosity and interconnectivity, as well as ECM-derived protein activities, would be an advantageous biomimetic platform for liver tissue engineering and drug screening applications. We therefore prepared GelMA ICC cell-laden constructs and assessed their organization, cell viability, growth, and functions in 2D and 3D GelMA (30 w/v%, 98% DS) systems.
After seeding Huh7.5 cells into the cavities of GelMA ICC scaffolds, Huh7.5 cells were rapidly infiltered through the cavities and attached to the cavity walls. Cell areas in the cavity grew increasingly large over 9 days of culture as presented in the live/dead micrographs in Figure 5a. The images show that Huh7.5 cells were highly viable (above 80%) both in 3D ICC scaffolds and on 2D substrates (Figure 5b). In GelMA ICC scaffolds, Huh7.5 cells stretched along the cavity walls and formed increasingly thick cell sheets over time. The thickness of 3D cell sheets more than doubled, from 22 μm on day 1 to 50 μm by day 9 (P<0.0001, one-way ANOVA). Three-dimensional confocal images (blue: nucleus; green: actin; red: CYP3A4 in Figures 6a and b) and scanning electron microscope images (Figure 6c) exhibit the clear morphological differences between the cells grown in 3D GelMA scaffolds and those grown on 2D GelMA substrates. Huh7.5 cells in GelMA ICC scaffolds initially anchored to the cavity walls and gradually grew into to the cell sheet constructs. On day 9, most of the cavities were filled with cells which then expanded to form 3D multi-cell clusters. This trend was prominent in the first layer and second layer (Supplementary Figure S5). On the other hand, cells on 2D GelMA substrates initially formed thin, scattered islands composed of several cells on day 1 which later merged to form a broader, albeit still very thin, cell sheet.
Evaluation of Huh7.5 cell viability and growing pattern in 3D GelMA ICC scaffolds and on 2D GelMA substrates. (a) Live/Dead staining images. Green and red indicate live and dead cells, respectively. (b) Cell viability values were quantified from Live/Dead staining images by ImageJ. Cells both in 2D and 3D showed high viability above 80% (for each group, three pictures were analysed containing >250 cells per image, mean±s.d.). (c) The cell multilayer thickness in cavities of GelMA ICC scaffolds was quantified by analysing the confocal images with ImageJ. The cell layers grew increasingly thick over a 9-day culture period (n=22; mean±s.d., P<0.0001, one-way ANOVA). ICC, inverted colloidal crystal; GelMA, gelatin methacryloyl.
Characterization of Huh7.5 morphologies in 3D GelMA ICC scaffolds and on 2D GelMA substrates during the culture. Huh7.5 cells in 3D GelMA scaffolds formed hepatic multilayer constructs in a 3D manner, whereas Huh7.5 cells on 2D GelMA substrates grew into 2D cell sheets. (a) Orthogonal projection images: The immunofluorescent images were taken via confocal microscopy (red: CYP3A4; green: f-actin; blue: nuclei). (b) 3D reconstruction images were taken via confocal microscopy (red: CYP3A4; green: f-actin; blue: nuclei). (c) Microscopic images via s.e.m. (scale bar=50 μm). ICC, inverted colloidal crystal; GelMA, gelatin methacryloyl.
Cell proliferation in 3D GelMA ICC scaffolds and 2D GelMA substrates was quantified by colorimetric assay using CCK-8 reagent. The proliferation of cells in 2D substrates gradually increased up to day 6 and then dropped abruptly on day 9 (Supplementary Figure S6). In contrast, the CCK values of cells in 3D GelMA ICC scaffolds changed moderately over the culture period. Although there could be some limitations for the reagent to diffuse within the whole layers of GelMA ICC 3D scaffolds compared to 2D substrates, it showed that cells on 2D substrates seemed to grow faster. This result is similar to previous reports with primary hepatocytes in two different systems,43 in which hepatocytes in a spheroid system were less proliferative than those in a monolayer system. On the other hand, albumin production in hepatospheroids was well maintained compared with the 2D system, where albumin production abruptly dropped. These results indicate that hepatocytes within 3D cultures experience a higher degree of cellular interactions than those in 2D cultures, helping the cells better maintain functionality.
In order to understand the influence of the GelMA ICC scaffolds on cell functions, Huh7.5 cells on 2D GelMA substrates and in 3D GelMA ICC scaffolds were assessed by immunocytochemical staining and western blot assay as presented in Figure 7. The target proteins of immunocytochemical staining were albumin and CYP3A4, which are an essential secretory protein and a drug metabolic enzyme, respectively. The images display the morphological changes of Huh7.5 cells in 2D and 3D and reveal intracellular albumin and CYP3A4 production. Overall, Huh7.5 cells in GelMA ICC scaffolds appeared to produce higher concentrations of albumin and showed markedly higher CYP3A4 expression over the life of the culture. By contrast, cellular activity on GelMA 2D substrates was less optimal in that protein expression grew rapidly in the early stages only to peak and decline dramatically later on (Figures 7a and b). These results show a direct correlation between cell morphologies and cell functions. Hepatocytes lose their original morphology on 2D substrates, and consequently, essential functions are hampered.4 Thus, 3D culture systems such as the spheroid culture and the sandwich culture have been used to preserve the native hepatocyte phenotype and prolong healthy function.7 As for 3D GelMA ICC scaffolds, the presence of interconnected, hexagonally arranged concave geometries can offer cells a biomimetic environment that is conducive to cell–materials and cell–cell interactions, facilitating the attachment of innate cell peptides and consequent cell growth. Moreover, a systematic comparison of GelMA ICC scaffolds with current 3D systems would underscore the advantages and disadvantages of promising 3D cell systems for in vitro tissue-specific analogues.
Evaluation of liver-specific functions of Huh7.5 cell constructs in 3D GelMA ICC scaffolds and on 2D GelMA substrates. (a, b) Confocal microscopy via immunohistochemistry. (red: (a) albumin and (b) CYP3A4; green: f-actin; blue: nuclei). (c) Western blot results of E-cadherin, albumin and GAPDH. Cell constructs in 3D GelMA ICC scaffolds appeared to produce more albumin and to express more CYP3A4 protein, compared to those on 2D GelMA substrates. E-cadherin and albumin in 3D GelMA ICC scaffolds were expressed gradually more and more, whereas those on 2D GelMA substrates were expressed at a high level on day 3 and then dwindled. ICC, inverted colloidal crystal; GAPDH, glyceraldehyde 3-phosphate dehydrogenase; GelMA, gelatin methacryloyl.
Western blot assay was conducted on albumin and E-cadherin (a transmembrane component of adherens junctions), with glyceraldehyde 3-phosphate dehydrogenase used as a normalizer. In the liver, E-cadherin is present in the periportal zone,44 where a higher amount of albumin was expressed.45 It has been reported that hepatospheroids, which are a well-studied configuration for enhanced cell–cell interactions and hepatic function maintenance,46 have high expression of E-cadherin.47 The results of western blot analysis show a close correlation between albumin and E-cadherin expression (Figure 7c). In GelMA 2D substrates, the expression levels of both proteins were maximized at day 3 when they formed flat clusters. Meanwhile, in GelMA ICC scaffolds, where cell constructs grew gradually and were interconnected between cavities in a 3D manner, the protein expression increased as culture time proceeded. These results suggest that GelMA ICC scaffolds with highly organized 3D interconnectivity and protein-derived biological activities could provide hepatic cell constructs with lasting intercellular interactions and promote continuous albumin production.
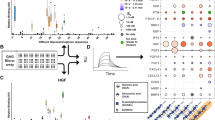
To examine a wide spectrum of cell functions at the gene expression level, hepatocyte-specific gene expression of Huh7.5 cells on 2D GelMA substrates and in 3D GelMA ICC scaffolds was quantified by RT-qPCR. Hepatocyte nuclear factors, regulating the expression of liver secretory proteins,48 and liver-specific genes including albumin, CYPs, alpha 1-antitrypsin, and glucose 6-phosphatase were chosen as hepatic markers as seen in Figures 8a–g. Among these genes, a drastic difference between 2D and 3D was observed in CYPs: The gene expression levels of CYP3A4 and CYP3A7 increased considerably in 3D GelMA scaffolds, whereas those in 2D GelMA substrates decreased by more than half after 9 days of culture. Further, hepatocyte nuclear factors and other secretory gene molecules were significantly upregulated in 3D GelMA ICC scaffolds compared with those in 2D GelMA substrates. The mRNA levels on cell junctions were also quantified. Figures 8h–i shows the expression level results of transmembrane components of adherent junctions (E-cadherin and N-cadherin). As for E-cadherin, the expression in 3D GelMA ICC scaffolds at day 9 was significantly higher than that in 2D GelMA substrates. This result is consistent with the western blot experiments, indicating an important advantage of 3D GelMA ICC cell constructs over 2D GelMA cell constructs for supporting functional hepatic cell growth. Of note, differences in N-cadherin expression between day 1 and day 9 were insignificant for both systems (P>0.05). As N-cadherin is related to cellular motility and increases during liver fibrosis,49 this result indicates that cells on GelMA systems exhibited non-mesenchymal characteristics. Other cell tight junction proteins (claudin-1 and zonula occludens (ZO)-1) were also assessed. In the liver, tight junctions exist between cells and play an important role in regulating paracellular diffusion and maintaining cellular polarity.50, 51 In 3D GelMA ICC scaffolds, the gene expression levels of both claudin-1 and ZO-1 increased after day 3 (Figures 8j–k). On the other hand, those in 2D GelMA substrates peaked on day 6 and did not show time-dependent behaviour. This result signifies enhanced intercellular communication and polarity maintenance in 3D GelMA ICC scaffolds. To summarize, these results on gene expression show upregulated hepatic functions in 3D GelMA ICC scaffolds compared with 2D GelMA substrates.
Effect of hepatic cell culture in 3D GelMA ICC scaffolds and on 2D GelMA substrates on liver-specific gene expression. Huh7.5 cells were cultured in both systems, and their RNA was extracted for the quantitative real-time PCR analysis of (a) AAT, (b) albumin, (c) CYP3A4, (d) CYP3A7, (e) G6Pase, (f) HNF4α, (g) HNF6, (h) E-cadherin, (i) N-cadherin, (j) claudin-1 and (k) ZO-1. The mRNA expression levels were normalized to GAPDH of the corresponding day and day 1 of the respective target gene. (n=3, mean±s.d.; #: P<0.05; ##: P<0.01; ###: P<0.001, for a comparison of 3D GelMA ICC scaffolds and 2D GelMA substrates.) AAT, alpha 1-antitrypsin; G6Pase, glucose 6-phosphatase; HNF, hepatocyte nuclear factor; ICC, inverted colloidal crystal; GelMA, gelatin methacryloyl.
Conclusion
In this study, we successfully constructed GelMA ICC hydrogel scaffolds possessing highly organized interconnected porous architectures and tunable biodegradation properties, and systematically characterized them with respect to structural and mechanical properties. The 3D GelMA ICC system negates the need for additional ECM coatings, which are cumbersome and may not assure reproducibility, and also possesses a collagenase biodegradation profile that can be tailored by adjusting the DS of the GelMA base material. Investigation of the GelMA ICC mechanical properties revealed high-dimensional stability as well as highly regular, interconnected pores that enabled excellent cell infiltration and subsequent cell–cell interactions between pores. Viability studies of hepatocytes loaded in 3D GelMA ICC scaffolds demonstrated enhanced cell–cell and cell–ECM interactions between pores over the 9-day culture period. Furthermore, the mRNA levels of multiple hepatocyte-specific genes in 3D GelMA ICC system were found to be significantly upregulated in comparison with a 2D GelMA system, which may be attributed to improved cell–cell and cell–material interactions from the 3D GelMA ICC geometry. Taken together, our results support that 3D GelMA ICC scaffolds lend excellent potential to the development of an improved, tunable, simple-to-produce artificial liver platform for use in applied settings such as drug screening, tissue engineering and beyond.
References
Cukierman, E., Pankov, R., Stevens, D. R. & Yamada, K. M. Taking cell-matrix adhesions to the third dimension. Science 294, 1708–1712 (2001).
Dutta, R. C. & Dutta, A. K. Cell-interactive 3D-scaffold; advances and applications. Biotechnol. Adv. 27, 334–339 (2009).
Schmeichel, K. L. & Bissell, M. J. Modeling tissue-specific signaling and organ function in three dimensions. J. Cell Sci. 116, 2377–2388 (2003).
Gómez-Lechón, M. J., Jover, R., Donato, T., Ponsoda, X., Rodriguez, C., Stenzel, K. G., Klocke, R., Paul, D., Guillén, I., Bort, R. & Castell, J. V. Long-term expression of differentiated functions in hepatocytes cultured in three-dimensional collagen matrix. J. Cell. Physiol. 177, 553–562 (1998).
Yoon, No, D., Lee, K. H., Lee, J. & Lee, S. H. 3D liver models on a microplatform: well-defined culture, engineering of liver tissue and liver-on-a-chip. Lab Chip 15, 3822–3837 (2015).
Cheng, Y., Zheng, F., Lu, J., Shang, L., Xie, Z., Zhao, Y., Chen, Y. & Gu, Z. Bioinspired multicompartmental microfibers from microfluidics. Adv. Mater. 26, 5184–5190 (2014).
Yamada, M., Utoh, R., Ohashi, K., Tatsumi, K., Yamato, M., Okano, T. & Seki, M. Controlled formation of heterotypic hepatic micro-organoids in anisotropic hydrogel microfibers for long-term preservation of liver-specific functions. Biomaterials 33, 8304–8315 (2012).
Kotov, N. A., Liu, Y., Wang, S., Cumming, C., Eghtedari, M., Vargas, G., Motamedi, M., Nichols, J. & Cortiella, J. Inverted colloidal crystals as three-dimensional cell scaffolds. Langmuir 20, 7887–7892 (2004).
Stein, A., Li, F. & Denny, N. R. Morphological control in colloidal crystal templating of inverse opals, hierarchical structures, and shaped particles. Chem. Mater. 20, 649–666 (2007).
Lee, J., Cuddihy, M. J., Cater, G. M. & Kotov, N. A. Engineering liver tissue spheroids with inverted colloidal crystal scaffolds. Biomaterials 30, 4687–4694 (2009).
Lee, J., Lilly, G. D., Doty, R. C., Podsiadlo, P. & Kotov, N. A. in vitro toxicity testing of nanoparticles in 3D cell culture. Small 5, 1213–1221 (2009).
Lee, J., Shanbhag, S. & Kotov, N. A. Inverted colloidal crystals as three-dimensional microenvironments for cellular co-cultures. J. Mater. Chem. 16, 3558–3564 (2006).
Liu, Y., Wang, S., Krouse, J., Kotov, N. A., Eghtedari, M., Vargas, G. & Motamedi, M. Rapid aqueous photo‐polymerization route to polymer and polymer‐composite hydrogel 3D inverted colloidal crystal scaffolds. J. Biomed. Mater. Res. Part A 83, 1–9 (2007).
Kim, M. H., Kumar, S. K., Shirahama, H., Seo, J., Lee, J. H. & Cho, N.-J. Phenotypic regulation of liver cells in a biofunctionalized three-dimensional hydrogel platform. Integr. Biol. 8, 156–166 (2015).
Kim, M. H., Kumar, S. K., Shirahama, H., Seo, J., Lee, J. H., Zhdanov, V. P. & Cho, N. J. Biofunctionalized hydrogel microscaffolds promote 3D hepatic sheet morphology. Macromol. Biosci. 16, 314–321 (2016).
Zhang, Y. & Xia, Y. Formation of embryoid bodies with controlled sizes and maintained pluripotency in three-dimensional inverse opal scaffolds. Adv. funct. mater. 22, 121–129 (2012).
Kuo, Y.-C. & Chung, C.-Y. TATVHL peptide-grafted alginate/poly(γ-glutamic acid) scaffolds with inverted colloidal crystal topology for neuronal differentiation of iPS cells. Biomaterials 33, 8955–8966 (2012).
Kuo, Y.-C. & Chen, C.-W. Inverted colloidal crystal scaffolds with induced pluripotent stem cells for nerve tissue engineering. Colloids Surf. B 102, 789–794 (2013).
Kuo, Y.-C. & Tsai, Y.-T. Inverted colloidal crystal scaffolds for uniform cartilage regeneration. Biomacromolecules 11, 731–739 (2010).
Kuo, Y.-C. & Tsai, Y.-T. Heparin-conjugated scaffolds with pore structure of inverted colloidal crystals for cartilage regeneration. Colloids Surf. B 82, 616–623 (2011).
Choi, S.-W., Xie, J. & Xia, Y. Chitosan-based inverse opals: three-dimensional scaffolds with uniform pore structures for cell culture. Adv. Mater. 21, 2997–3001 (2009).
Yang, J.-T., Kuo, Y.-C. & Chiu, K.-H. Peptide-modified inverted colloidal crystal scaffolds with bone marrow stromal cells in the treatment for spinal cord injury. Colloids Surf. B 84, 198–205 (2011).
Kuo, Y.-C. & Chiu, K.-H. Inverted colloidal crystal scaffolds with laminin-derived peptides for neuronal differentiation of bone marrow stromal cells. Biomaterials 32, 819–831 (2011).
Berger, J., Reist, M., Mayer, J. M., Felt, O., Peppas, N. A. & Gurny, R. Structure and interactions in covalently and ionically crosslinked chitosan hydrogels for biomedical applications. Eur. J. Pharmaceut. Biopharmaceut. 57, 19–34 (2004).
Klotz, B. J., Gawlitta, D., Rosenberg, A. J., Malda, J. & Melchels, F. P. Gelatin-methacryloyl hydrogels: towards biofabrication-based tissue repair. Trends Biotechnol. 34, 394–407 (2016).
Billiet, T., Gevaert, E., De Schryver, T., Cornelissen, M. & Dubruel, P. The 3D printing of gelatin methacrylamide cell-laden tissue-engineered constructs with high cell viability. Biomaterials 35, 49–62 (2014).
Lee, B. H., Shirahama, H., Cho, N.-J. & Tan, L. P. Efficient and controllable synthesis of highly substituted gelatin methacrylamide for mechanically stiff hydrogels. RSC Adv. 5, 106094 (2015).
Shirahama, H., Lee, B. H., Tan, L. P. & Cho, N. J. Precise tuning of facile one-pot gelatin methacryloyl (GelMA) synthesis. Sci. Rep. 6, 31036 (2016).
McCuskey, R. in Zakim and Boyer’s Hepatology 6th edn (eds. Boyer, T. D., Manns, M. P. & Sanyal, A. J.) 3–19 (W.B. Saunders, Saint Louis, 2012).
Eastoe, J. The amino acid composition of mammalian collagen and gelatin. Biochem. J. 61, 589 (1955).
Van Den Bulcke, A. I., Bogdanov, B., De Rooze, N., Schacht, E. H., Cornelissen, M. & Berghmans, H. Structural and rheological properties of methacrylamide modified gelatin hydrogels. Biomacromolecules 1, 31–38 (2000).
Yue, K., Trujillo-de Santiago, G., Alvarez, M. M., Tamayol, A., Annabi, N. & Khademhosseini, A. Synthesis, properties, and biomedical applications of gelatin methacryloyl (GelMA) hydrogels. Biomaterials 73, 254–271 (2015).
Colosi, C., Shin, S. R., Manoharan, V., Massa, S., Costantini, M., Barbetta, A., Dokmeci, M. R., Dentini, M. & Khademhosseini, A. Microfluidic bioprinting of heterogeneous 3D tissue constructs using low-viscosity bioink. Adv. Mater. 28, 677–684 (2016).
Loessner, D., Meinert, C., Kaemmerer, E., Martine, L. C., Yue, K., Levett, P. A., Klein, T. J., Melchels, F. P., Khademhosseini, A. & Hutmacher, D. W. Functionalization, preparation and use of cell-laden gelatin methacryloyl-based hydrogels as modular tissue culture platforms. Nat. Protoc. 11, 727–746 (2016).
Kim, J., Bencherif, S. A., Li, W. A. & Mooney, D. J. Cell-friendly inverse opal-like hydrogels for a spatially separated co-culture system. Macromol. Rapid Commun. 35, 1578–1586 (2014).
Ananthanarayanan, A., Narmada, B. C., Mo, X., McMillian, M. & Yu, H. Purpose-driven biomaterials research in liver-tissue engineering. Trends Biotechnol. 29, 110–118 (2011).
Li, Y.-S., Harn, H.-J., Hsieh, D.-K., Wen, T.-C., Subeq, Y.-M., Sun, L.-Y., Lin, S.-Z. & Chiou, T.-W. Cells and materials for liver tissue engineering. Cell Transplant. 22, 685–700 (2013).
Bhatia, SN, Underhill, GH, Zaret, KS & Fox, IJ Cell and tissue engineering for liver disease. Science Translational Medicine 6: 245sr2–245sr2 (2014).
Elliott, N. T. & Yuan, F. A review of three-dimensional in vitro tissue models for drug discovery and transport studies. J. Pharmaceut. Sci. 100, 59–74 (2011).
Ho, C.-T., Lin, R.-Z., Chen, R.-J., Chin, C.-K., Gong, S.-E., Chang, H.-Y., Peng, H.-L., Hsu, L., Yew, T.-R. & Chang, S.-F. Liver-cell patterning lab chip: mimicking the morphology of liver lobule tissue. Lab Chip 13, 3578–3587 (2013).
Choi, S.-W., Zhang, Y. & Xia, Y. Three-dimensional scaffolds for tissue engineering: the importance of uniformity in pore size and structure. Langmuir 26, 19001–19006 (2010).
Curcio, E., Salerno, S., Barbieri, G., De Bartolo, L., Drioli, E. & Bader, A. Mass transfer and metabolic reactions in hepatocyte spheroids cultured in rotating wall gas-permeable membrane system. Biomaterials 28, 5487–5497 (2007).
Koide, N., Shinji, T., Tanabe, T., Asano, K., Kawaguchi, M., Sakaguchi, K., Koide, Y., Mori, M. & Tsuji, T. Continued high albumin production by multicellular spheroids of adult rat hepatocytes formed in the presence of liver-derived proteoglycans. Biochem. Biophys. Res. Commun. 161, 385–391 (1989).
Hempel, M., Schmitz, A., Winkler, S., Kucukoglu, O., Bruckner, S., Niessen, C. & Christ, B. Pathological implications of cadherin zonation in mouse liver. Cellular and Molecular Life Sciences 72, 2599–2612 (2015).
Racine, L., Scoazec, J. Y., Moreau, A., Chassagne, P., Bernuau, D. & Feldmann, G. Distribution of albumin, α1-inhibitor 3 and their respective mRNAs in periportal and perivenous rat hepatocytes isolated by the digitonin-collagenase technique. Biochem. J. 305 (Pt 1), 263–268 (1995).
Lin, R. Z. & Chang, H. Y. Recent advances in three-dimensional multicellular spheroid culture for biomedical research. Biotechnol. J. 3, 1172–1184 (2008).
Lin, R. Z., Chou, L. F., Chien, C. C. & Chang, H. Y. Dynamic analysis of hepatoma spheroid formation: roles of E-cadherin and β1-integrin. Cell Tissue Res. 324, 411–422 (2006).
Odom, D. T., Zizlsperger, N., Gordon, D. B., Bell, G. W., Rinaldi, N. J., Murray, H. L., Volkert, T. L., Schreiber, J., Rolfe, P. A., Gifford, D. K., Fraenkel, E., Bell, G. I. & Young, R. A. Control of pancreas and liver gene expression by HNF transcription factors. Science 303, 1378–1381 (2004).
Lee, J. M., Dedhar, S., Kalluri, R. & Thompson, E. W. The epithelial-mesenchymal transition: new insights in signaling, development, and disease. J. Cell Biol. 172, 973–981 (2006).
Vinken, M., Papeleu, P., Snykers, S., De Rop, E., Henkens, T., Chipman, J. K., Rogiers, V. & Vanhaecke, T. Involvement of cell junctions in hepatocyte culture functionality. Crit. Rev. Toxicol. 36, 299–318 (2006).
Kojima, T., Yamamoto, T., Murata, M., Chiba, H., Kokai, Y. & Sawada, N. Regulation of the blood-biliary barrier: interaction between gap and tight junctions in hepatocytes. Med. Electron Microscopy 36, 157–164 (2003).
Acknowledgements
This research is financially supported by the National Research Foundation (NRF) Competitive Research Programme (NRF-CRP10-2012-07) grant in Singapore and is also supported by the Creative Materials Discovery Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT, and Future Planning (2016M3D1A1024098).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
PUBLISHER’S NOTE
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations
Supplementary Information accompanies the paper on the NPG Asia Materials website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Lee, B., Shirahama, H., Kim, M. et al. Colloidal templating of highly ordered gelatin methacryloyl-based hydrogel platforms for three-dimensional tissue analogues. NPG Asia Mater 9, e412 (2017). https://doi.org/10.1038/am.2017.126
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/am.2017.126
This article is cited by
-
Organ-On-A-Chip Technology: An In-depth Review of Recent Advancements and Future of Whole Body-on-chip
BioChip Journal (2023)
-
Natural Scaffolds Used for Liver Regeneration: A Narrative Update
Stem Cell Reviews and Reports (2022)
-
Recent advances in the development of in vitro liver models for hepatotoxicity testing
Bio-Design and Manufacturing (2021)
-
Gelatin methacryloyl and its hydrogels with an exceptional degree of controllability and batch-to-batch consistency
Scientific Reports (2019)
-
Epithelial-mesenchymal transition of cancer cells using bioengineered hybrid scaffold composed of hydrogel/3D-fibrous framework
Scientific Reports (2019)