Abstract
We fabricated and tested encoders and decoders based on a multiplex, DNA-based electrochemical biosensor that uses electronic (electrochemical) signals as its readout. These devices use two or more sequence-specific DNA probes, with each being modified with a distinct redox reporter. These probes, when interrogated together, serve as encoders and decoders, converting patterns that are encoded and decoded by the presence or absence of specific DNA sequences into specific electronic outputs. We demonstrated these multifunctional, bio-electrochemical devices, for example, 4-to-2 and 8-to-3 encoders and 1-to-2 and 2-to-3 decoders. Accordingly, these devices bridge the division between DNA-based devices and silicon-based electronics.
Similar content being viewed by others
Introduction
In recent decades, serious attempts have been made to implement computational approaches, models and paradigms that utilize the information transfer and processing abilities of naturally occurring and modified biomolecules.1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 A decade ago, for example, Adleman performed a now-classic experiment in which computations were conducted using DNA molecules, demonstrating that biology can provide new computing substrates.12, 13, 14, 15 In the decade since Adleman's invention, reports have been made of a wide and diverse range of DNA logic devices.16, 17, 18, 19, 20, 21, 22 Winfree, Stojanovic, Willner, Katz and their co-workers have, for example, developed optically reported ‘AND’, ‘OR’ and ‘SET–RESET’ logic gate operations.7, 23, 24, 25, 26, 27, 28, 29, 30, 31 Although the above examples serve as promising proofs of principle (see also refs 27,28,29,30,31,32,33,34,35), it remains necessary to create complex, multi-component devices on a single biomolecular platform to achieve increased computational complexity and develop realistic DNA-based information processing systems. In this study, we fabricated and tested encoders and decoders based on a multiplex, DNA-based electrochemical biosensor that uses electronic (electrochemical) signals as its readout. These devices uses two or more sequence-specific DNA probes that, when interrogated together, serve as encoders, converting patterns that are encoded and decoded by the presence or absence of specific DNA sequences into specific electronic outputs.
Materials and methods
Reagents
The following reagents were used as received: 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES; Sigma Aldrich, St Louis, MO, USA), fetal calf serum (Sigma Aldrich), ferrocenecarboxylic acid(FC-COOH), N-hydroxysuccinimide (Sigma Aldrich), N-hydroxysulfosuccinimide sodium salt (NHS), N-(3-(dimethylamino)propyl)-N′-ethylcarbodiimide hydrochloride (Fluka, St Louis, MO, USA) and 3,7-bis(N-(3-carboxypropyl)-N-methylamino)phenothiazin-5-ium perchlorate (MB-NHS, EMP-Biotech GmbH, Berlin, Germany), FC-NHS, anthraquinone (AQ)-NHS (methylene blue (MB)-NHS, EMP-Biotech).
The probe and target DNA sequences (Biosearch Technologies, Novato, CA, USA) used were as follows: probe,
5′-HS-C6-TGGATCGGCGTTTTATT-C7-NH2-3′ (for AQ-NHS);
5′-HS-C6-AGACAAGGAAAATCCTTCAATGAAGTGGGTCG-C7-NH2-3′ (for FC-NHS)
5′-HS-C6-AGCCCATTTATCCGTTCCTCCTAGTGGTGTGC-C7-NH2-3′ (for MB-NHS)
5′-HS-C6-GGCGAGGTACGACGGCCAGCCTCGCCGAGCCGTCG-C7-NH2-3′ (for Signal-on MB-NHS)
5′-AATAAAACGCCGATCCA-3′ (AQ-DNA)
5′-CGACCCACTTCATTGAAGGATTTTCCTTGTCT-3′ (FC-DNA)
5′-AGGCAAGGAGGA-3′ (MB-DNA)
5′-AATAAAACGCCGATCCATTTTTTTTCGACCCACTTCATTGAAGGATTTTCCTTGTCT-3′ (AQ+FC-DNA)
5′-AATAAAACGCCGATCCATTTTTTTTAGGCAAGGAGGA -3′ (AQ+MB-DNA)
5′-CGACCCACTTCATTGAAGGATTTTCCTTGTCTTTTTTTTTAGGCAAGGAGGA-3′ (FC+MB-DNA)
5′-AATAAAACGCCGATCCATTTTTTTTCGACCCACTTCATTGAAGGATTTTCCTTGTCTTTTTTTTTAGGCAAGGAGGA -3′ (AQ+FC+MB-DNA)
5′-GCTGGCCGTCGTTTTAC-3′ (Signal-on MB-DNA)
5′-AATAAAACGCCGATCCATTTTTTTTGCTGGCCGTCGTTTTAC-3′ (AQ+Signal-on MB-DNA)
5′-CGACCCACTTCATTGAAGGATTTTCCTTGTCTTTTTTTTTGCTGGCCGTCGTTTTAC-3′ (FC+Signal-on MB-DNA).
The probe DNA was modified at the 5′-terminus with a C6-disulfide [HO(CH2)6-S-S-(CH2)6-] linker and at the 3′-end with an amine group for redox probe conjugation.
Labeling of DNA probes
Conjugation of the redox labels to the probe DNA was achieved via the coupling of the X-NHS ester (X=MB, FC, AQ) conjugates with a 5′-alkyl-amino-modified, single-stranded DNA. To accomplish this conjugation, 10 μl of 200 μM 5′-alkyl-amino DNA was added to 50 μl of a 0.5 M sodium bicarbonate solution (pH 8.5), and the X-NHS (1 M) was dissolved in 10 μl dimethyl sulfoxide. These two solutions were added to make a final volume of 70 μl and stirred for 4 h at room temperature in the dark. The mixture was subsequently desalted using a spin column (EMP-Biotech) and purified by reverse phase-high-performance liquid chromatography (C18 column). Stock solutions of these modified DNAs were stored at −20 °C for future use. The yield of the modified DNA, as estimated using high-performance liquid chromatography and mass spectrometry, is ∼20%.
Electrode preparation and sensor fabrication
Before sensor fabrication, gold-disk electrodes (2 mm diameter, CH Instruments, Austin, TX, USA) were cleaned both mechanically (by polishing with diamond and with alumina oxide slurries, successively) and electrochemically (through successive scans in sulfuric acid solutions), as previously described. The DNA probe was reduced for 1 h at the room temperature in the dark in 10 mM tris(2-carboxyethyl)phosphine hydrochloride (Molecular Probes, Carlsbad, CA, USA) and was diluted to a final concentration of 1.0 μM in HEPES/NaClO4 buffer (10 mM HEPES and 0.5 M NaClO4, pH 7.0, as was used in all the experiments to follow unless otherwise noted). The gold electrodes were incubated in this solution (50 nM AQ-DNA probe and 100 nM MB-DNA probe for 1-to-2 decoders, 100 nM MB-DNA probe, 100 nM FC-DNA probe and 50 nM AQ-DNA probe for 2-to-3 decoders) for 1 h at (room temperature for 1-to-2 decoders, 50 °C for 2-to-3 decoders) in the dark, rinsed with distilled, deionized water and incubated in 3 mM 6-mercapto-1-hexanol in deionized water for 30 min. Subsequently, the electrodes were rinsed in the deionized water and stored in HEPES/NaClO4 buffer for future use.
Sensor measurement
Fabricated sensors were interrogated using square wave voltammetry with a 50-mV amplitude signal at a frequency of 60 Hz in the absence and presence of fully-complementary target (2 μM). The electrodes were incubated for 30 min with the appropriate concentration of target DNA in HEPES/NaClO4 buffer or 20% fetal calf serum mixed with HEPES/NaClO4 buffer. Relative signal changes were computed by the relative change in square wave voltammetry peak currents with respect to background current (square wave voltammetry peak current in the absence of target).
Results and Discussion
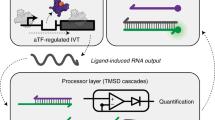
Our encoders are based on the E-DNA sensor, a previously described electrochemical sensor36, 37 architecture that consists of an electrode-attached, redox-reporter-modified DNA probe (Figure 1). These E-DNA sensors turn ‘off’ (that is, their redox reporter produces less faradic current) when the hybridization of the target to the DNA probe forces the redox reporter away from the electrode. We adapted these sensors into encoders by fabricating two or more of them on a single interrogating electrode. To distinguish between the signals arising from every sensor, each of the sensors was modified with a different redox tag and thus produced a signal at its own unique redox potential. Specifically, in this work, we used E-DNA sensors using the redox reporters: AQ (potential at −0.54 V vs Ag/AgCl), MB (−0.27 V) and FC (+0.18 V). Because the probe DNA is fully covalent, the platform architecture is stable in complex media. Of note, we can perform the sensors in 20% serum (diluted with buffered saline to control the pH and ionic strength; Figure 1).
E-DNA biosensors comprise an oligonucleotide probe attached to an electrode via an alkane thiol and modified with a redox reporter. For this experiment, we used three redox reporters (left) methylene blue (MB), (middle) ferrocene (FC) and (right) anthraquinone (AQ). Shown here are linear probe E-DNA sensors, which utilize an unstructured DNA probe that allows the redox reporter to approach in close proximity to the electrode surface, thereby ensuring rapid electron transfer and efficient redox of the MB (at −0.27 V), FC (at +0.18 V) and AQ (at −0.54 V) label. Target binding forms a rigid double strand, reducing the approach efficiency and lowering the observed current.
As a first test of our approach, we designed a 4-to-2 encoder, which compresses four input states into two output states (Figure 2), and defined the concentration of various DNA targets as inputs and the signal change in the faradic current from attached MB and FC redox reporters as outputs (see Supplementary Information for details of the square wave voltammetric protocols used). For input, the presence of the relevant DNA targets at concentrations >500 nM defined the ‘1’ states, and lower (to 0 M) concentrations defined the ‘0’ states. (The relatively high concentrations of the inputs induce distinctive change in the signal outputs and make the responsive time short for our devices.) As outputs, we defined signal changes (changes in the reduction peak of the MB or FC) of greater than 25% and less than 25% as the ‘1’ and ‘0’ states, respectively.
To fabricate this 4-to-2 encoder device, we used two linear E-DNA probes using previously reported MB and FC moieties, which were immobilized together on the surface of a single gold electrode via self-assembled monolayer chemistry (top left). When interrogated via square wave voltammetry (SWV), these probes responded to their respective targets (two different DNA molecules) via decrease in the faradic current (bottom left, right). Together, these probes comprise the encoder for which four input combinations induced different electrochemical output currents when probed using SWV. The truth table for 4-to-2 encoder is shown.
The initial state of the encoder was seen in the absence of any DNA target; the system was reset to this state before each input operation via a 30-s wash in distilled water, which disrupted the hybridization and regenerates the sensors. Four DNA target inputs, D0 to D3, were compressed into two electronic outputs, X and Y, which are the currents observed in the MB and FC reduction peaks, respectively. D0, a DNA sequence, did not hybridize with either of the probes nor did it alter the MB (X=0) or FC (Y=0) peaks. D1, which is complementary to the FC-modified probe, decreased the signal of its redox peak (Y=1) but did not change the signal at the MB redox potential (X=0). D2, which is complementary to the MB-modified probe, did not affect the signal at the FC redox potential (Y=0) but decreased the signal at the MB redox potential (X=1). D3, which contains DNA complementary to both the MB- and FC-modified probes, decreased in both redox peaks (X=1, Y=1). Overall, these operations resulted in a truth table (Figure 2) characteristic of a 4-to-2 encoder.
Building on our initial studies of the 4-to-2 encoder, we fabricated an 8-to-3 encoder. Therefore, we added AQ as a third redox reporter, which provided a third, independent output signal. Similar to the previous case, the presence of each DNA at concentrations >500 nM defines the ‘1’ state, and lower (to 0 M) concentrations of these molecules define the ‘0’ states. Eight DNA sequence inputs, specifically, D0 to D7, were thus compressed by the encoder into three electronic outputs, X, Y and Z at the redox potentials of AQ, MB and FC, respectively. As outputs, we again defined signal changes (changes in the relevant reduction peak) of greater than 25% and less than 25% as the ‘1’ and ‘0’ states, respectively. On the basis of the above definitions, an 8-to-3 operation was thus realized by controlling the concentrations of DNA for the AQ-, MB- and FC-modified probes. The truth table and schematic representation of this encoder are presented in Figure 3; they demonstrate an easy realization of a relatively complex encoder by our multiplexed sensor platform.
To fabricate this 8-to-3 encoder device we used three linear E-DNA probes using MB, ferrocene (FC) and anthraquinone (AQ) redox reporters, which were immobilized together on the surface of a single gold electrode via self-assembled monolayer chemistry (left). When interrogated via square wave voltammetry (SWV), these probes responded to their respective targets (three different cDNA) via decreases in the faradic current (right). Together, these probes comprise the encoder for which eight input combinations induced different electrochemical output currents when probed via SWV. The truth table for 8-to-3 encoder is shown.
A decoder is a device that reverses the action of an encoder, undoing the encoding such that the original information can be retrieved. To build the decoders, we used ‘signal-on’ E-DNA sensors consisting of a short, single-stranded DNA pseudoknot that forms two stem-loop structures in which a portion of each loop forms one strand of the stem of the other stem-loop. Such sensors turn ‘on’ (that is, the faradic current increases) when the hybridization of the target to the probe liberates a flexible, single-stranded element that allows the redox reporter to approach the electrode.
To demonstrate our technique, we designed a 1-to-2 decoder that defines the concentration of a DNA target as input and the signal change in the faradic current from the redox reporters AQ and MB as outputs. We designed the encoder using one pseudoknot E-DNA probe (signal-on) and one linear E-DNA probe (signal-off; Supplementary Figures S1–S3). For input, the presence of DNA at concentrations >500 nM defines the ‘1’ state, and lower (to 0 M) concentrations of these molecules define the ‘0’ state. For output, we defined signals below and above 150 nA as ‘1’ and ‘0’, respectively. When no input was applied, the system produced 183 nA at the AQ potential (X=0) and 110 nA at the MB potential (Y=1). D0, which contained both the targets for the MB signal-on probe and the AQ signal-off probe, hybridizes with the MB and AQ probe, respectively, increasing the signal at the MB potential to 189 nA (Y=0) and decreasing the signal at the AQ potential to 85 nA (X=1). Overall, these operations resulted in a truth table (Figure 4) characteristic of a 1-to-2 decoder.
To fabricate this 1-to-2 decoder device, we used a linear probe modified with anthraquinone (AQ) moiety and a signal-on DNA sensor modified with methylene blue (MB) moiety, both of which were immobilized on the surface of a single gold electrode via self-assembled monolayer chemistry. When interrogated via square wave voltammetry (SWV), these probes responded to the target (involving the target for the MB signal-on probe and AQ signal-off probe and hybridizing with the AQ and MB probe) via changes in the faradic current. The truth table for 1-to-2 decoder is shown.
Building on the studies of the 1-to-2 decoder, we designed a 2-to-3 decoder that defines the concentration of a DNA target as input and the signal change in the faradic current from the redox reporters MB, FC and AQ as outputs. For this device, we used one signal-on and two signal-off E-DNA probes (Figure 5). As inputs, the presence of the relevant DNA targets at concentrations > 500 nM define the ‘1’ state, and lower (to 0 M) concentrations of these molecules define the ‘0’ state. For output, we defined the signal lower than 75 nA and larger than 75 nA as ‘1’ and ‘0’, respectively. When no input was applied, the system produced 105 nA at the AQ potential (X=0), 63 nA at the MB potential (Y=1) and 99 nA at the FC potential (Z=0). D0, which contained both the targets for the MB signal-on probe and the FC signal-off probe, hybridizes with the MB and FC probe, increasing the signal at the MB potential to 115 nA (Y=0) and decreasing the signal at the FC potential to 16 nA (Z=1). Moreover, the AQ potential was measured to be 101 nA (X=0). D1, which contained both the target for the MB signal-on probe and the AQ signal-off probe, hybridized with the MB and AQ probe, thereby increasing the signal at the MB potential to 97 nA (Y=0) and decreasing the signal at the AQ potential to 24 nA (X=1). Furthermore, the FC potential was measured to be 93 nA (Z=0). Overall, these operations resulted in a truth table (Figure 5) characteristic of a 2-to-3 decoder.
To fabricate this 2-to-3 decoder device, we used two previously described linear probes modified with ferrocene (FC) and anthraquinone (AQ) moieties and a signal-on DNA sensor modified with MB moiety, all of which were immobilized on the surface of a single gold electrode via self-assembled monolayer chemistry. When interrogated via square wave voltammetry (SWV), these probes responded to their respective targets (two different cDNA: one for the MB signal-on probe and FC signal-off probe, and the other for MB signal-on probe and AQ signal-off probe) via changes in the faradic current. For output, we defined the signal lower than 75 nA and larger than 75 nA as ‘1’ and ‘0’, respectively. The truth table for 2-to-3 decoder is shown.
Conclusion
In this study, we demonstrated a set of multifunctional, bio-electrochemical devices that realize the functions of encoding and decoding, converting bio-molecular information into electronic output signals. We used the electronic (electrochemical) signal as output, which is practical (Supplementary Figures S4–S6) and can be easily used as computing devices (most of the logic operations designed before were realized through fluorescence output). Accordingly, these devices bridge the barrier between DNA-based devices and silicon-based electronics. Although the construction of a DNA-based computer is currently extremely ambitious, even in long term, simple and dedicated computing tasks, such as those illustrated here, may prove quite amenable to DNA-based devices. Some specific problems that traditional electronic circuits cannot address include, for example, labeling and tracking nano- and micrometric objects and monitoring and controlling biological processes. Potential applications of DNA-based encoders and decoders for labeling and sensing nano- and micro-scale particles, as well as data processing, can also be envisioned, especially given that the molecular-scale dimensions of DNA suggest that parallel, integrated devices could be constructed realizing more complicated functions.
References
Yan, H. Nucleic acid nanotechnology. Science 306, 2048–2049 (2004).
Stojanovic, M. N. & Stefanovic, D. A deoxyribozyme-based molecular automaton. Nat. Biotechnol. 21, 1069–1074 (2003).
Seeman, N. C. From genes to machines: DNA nanomechanical devices. Trends Biochem. Sci. 30, 119–125 (2005).
de Silva, A. P. & Uchiyama, S. Molecular logic and computing. Nat. Nanotechnol. 2, 399–410 (2007).
Condon, A. Designed DNA molecules: principles and applications of molecular nanotechnology. Nat. Rev. Genet. 7, 565–575 (2006).
Solanki, P. R., Kaushik, A., Agrawal, V. V. & Malhotra, B. D. Nanostructured metal oxide-based biosensors. NPG Asia Mater. 3, 17–24 (2011).
Motornov, M., Zhou, J., Pita, M., Gopishetty, V., Tokarev, I., Katz, E. & Minko, S. ‘Chemical transformers’ from nanoparticle ensembles operated with logic. Nano Lett. 8, 2993–2997 (2008).
Li, J. Self-assembled supramolecular hydrogels based on polymer-cyclodextrin inclusion complexes for drug delivery. NPG Asia Mater. 2, 112–118 (2010).
Heath, J. R. & Ratner, M. A. Molecular electronics. Physics Today 56, 43–49 (2003).
Tour, J. M. Molecular electronics. Synthesis and testing of components. Accounts Chem. Res. 33, 791–804 (2000).
Ding, B. & Seeman, N. C. Operation of a DNA robot arm inserted into a 2D DNA crystalline substrate. Science 314, 1583–1585 (2006).
Mao, C. D., LaBean, T. H., Reif, J. H. & Seeman, N. C. Logical computation using algorithmic self-assembly of DNA triple-crossover molecules. Nature 407, 493–496 (2000).
Benenson, Y., Paz-Elizur, T, Adar, R, Keinan, E, Livneh, Z & Shapiro, E . Programmable and autonomous computing machine made of biomolecules. Nature 414, 430–434 (2001).
Liu, Q. H., Wang, L., Frutos, A. G., Condon, A. E., Corn, R. M. & Smith, L. M . DNA computing on surfaces. Nature 403, 175–179 (2000).
Adleman, L. M. Molecular computation of solutions to combinatorial problems. Science 266, 1021–1024 (1994).
Bozdemir, O. A., Guliyev, R., Buyukcakir, O., Selcuk, S., Kolemen, S., Gulseren, G., Nalbantoglu, T., Boyaci, H. & Akkaya, E. U . Selective manipulation of ICT and PET processes in styryl-bodipy derivatives: applications in molecular logic and fluorescence sensing of metal ions. J. Am. Chem. Soc. 132, 8029–8036 (2010).
Macdonald, J., Li, Y, Sutovic, M., Lederman, H., Pendri, K., Lu, W., Andrews, B. L., Stefanovic, D. & Stojanovic, M. N . Medium scale integration of molecular logic gates in an automaton. Nano Lett. 6, 2598–2603 (2006).
Pu, F., Liu, Z., Yang, X., Ren, J. & Qu, X. DNA-based logic gates operating as a biomolecular security device. Chem. Commun. 47, 6024–6026 (2011).
Li, T., Zhang, L., Ai, J., Dong, S. & Wang, E. Ion-tuned DNA/Ag fluorescent nanocluster as versatile logic device. ACS Nano 5, 6334–6338 (2011).
Xia, F., Zuo, X.l., Yang, R. Q., White, R. J., Xiao, Y., Kang, D., Gong, X., Lubin, A. A., Vallee-Belisle, A., Yuen, J. D., Hsu, B. Y. B. & Plaxco, K. W. Label-free, dual-analyte electrochemical biosensors: a new class of molecular-electronic logic gates. J. Am. Chem. Soc. 132, 8557–8859 (2010).
Ogihara, M. & Ray, A. Molecular computation: DNA computing on a chip. Nature 403, 143–144 (2000).
Benenson, Y., Gil, B., Ben-Dor, U., Adar, R. & Shapiro, E. An autonomous molecular computer for logical control of gene expression. Nature 429, 423–429 (2004).
Elbaz, J., Wang, Z. G., Orbach, R. & Willner, I. pH-stimulated concurrent mechanical activation of two DNA ‘Tweezers’* A ‘SET-RESET’ logic gate system. Nano Lett. 9, 4510–4514 (2009).
Willner, I., Shlyahovsky, B., Zayats, M. & Willner, B. DNAzymes for sensing, nanobiotechnology and logic gate applications. Chem. Soc. Rev. 37, 1153–1165 (2008).
Weizmann, Y., Elnathan, R., Lioubashevski, O. & Willner, I. Endonuclease-based logic gates and sensors using magnetic force-amplified readout of DNA scission on cantilevers. J. Am. Chem. Soc. 127, 12666–12672 (2005).
Elbaz, J., Lioubashevski, O ., Wang, F ., Remacle, F ., Levine, R. D . & Willner, I . DNA computing circuits using libraries of DNAzyme subunits. Nat. Nanotechnol. 5, 417–422 (2010).
Seelig, G., Soloveichik, D., Zhang, D. Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Huang, Y., Duan, X., Cui, Y., Lauhon, L. J., Kim, K. H. & Lieber, C. M. Logic gates and computation from assembled nanowire building blocks. Science 294, 1313–1317 (2001).
Okamoto, A., Tanaka, K. & Saito, I. DNA logic gates. J. Am. Chem. Soc. 126, 9458–9463 (2004).
Saghatelian, A., Volcker, N. H., Guckian, K. M., Lin, V. S. Y. & Ghadiri, M. R. DNA-based photonic logic gates: AND, NAND, and INHIBIT. J. Am. Chem. Soc. 125, 346–347 (2003).
Stojanovic, M. N., Mitchell, T. E. & Stefanovic, D. Deoxyribozyme-based logic gates. J. Am. Chem. Soc. 124, 3555–3561 (2002).
Chen, X., Wang, Y., Liu, Q., Zhang, Z., Fan, C. & He, L. Construction of molecular logic gates with a DNA-cleaving deoxyribozyme. Angew. Chem. Int. Ed. 45, 1759–1762 (2006).
Miyoshi, D., Inoue, M. & Sugimoto, N. DNA logic gates based on structural polymorphism of telomere DNA molecules responding to chemical input signals. Angew. Chem. Int. Ed. 45, 7716–7719 (2006).
Gianneschi, N. C. & Ghadiri, M. R. Design of molecular logic devices based on a programmable DNA-regulated semisynthetic enzyme. Angew. Chem.-Int. Ed. 46, 3955–3958 (2007).
Liu, J. W., Cao, Z. H. & Lu, Y. Functional nucleic acid sensors. Chem. Rev. 109, 1948–1998 (2009).
Fan, C., Plaxco, K. W. & Heeger, A. J. Electrochemical interrogation of conformational changes as a reagentless method for the sequence-specific detection of DNA. Proc. Natl Acad. Sci. USA 100, 9134–9137 (2003).
Kang, D., Zuo, X., Yang, R., Xia, F., Plaxco, K.W. & White, R. J. Comparing the properties of electrochemical-based DNA sensors employing different redox tags. Anal. Chem 81, 9109–9113 (2009).
Acknowledgements
We are grateful to Professor Alan J Heeger for discussion. This research was supported by initiatory financial support from HUST. We also thank the HUST Analytical and Testing Center for allowing us to use its facilities.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on the NPG Asia Materials website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Kang, D., White, R., Xia, F. et al. DNA biomolecular-electronic encoder and decoder devices constructed by multiplex biosensors. NPG Asia Mater 4, e1 (2012). https://doi.org/10.1038/am.2012.1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/am.2012.1
Keywords
This article is cited by
-
Doping-induced changes in the structural, optical, magnetic and thermal properties of Ni:ZnS nanoparticles prepared by microwave-assisted chemical method
Applied Physics A (2021)
-
DNA Strand Displacement Reaction: A Powerful Tool for Discriminating Single Nucleotide Variants
Topics in Current Chemistry (2020)
-
Probing the effect of selenium substitution in kesterite-Cu2ZnSnS4 nanocrystals prepared by hot injection method
Journal of Materials Science: Materials in Electronics (2019)
-
A versatile DNA-supramolecule logic platform for multifunctional information processing
NPG Asia Materials (2018)
-
Dual-electrochromic bipolar electrode-based universal platform for the construction of various visual advanced logic devices
NPG Asia Materials (2017)