Abstract
Previous studies have shown that Arabidopsis equilibrative nucleoside transporters (AtENTs) possess transport activities when produced in yeast cells and are differentially expressed in Arabidopsis organs. Herein, we report further analysis on the nucleoside transport activities and transcriptional patterns of AtENT members. The recombinant proteins of AtENTs 3, 6, and 7, but not those of AtENTs 1, 2, 4, and 8, were found to transport thymidine with high affinity. Contrary to previous suggestion that AtENT1 may not transport uridine, this work showed that recombinant AtENT1 was a pH-dependent and high-affinity transporter of uridine. When grown on MS plates, the AtENT3 knockout plants were more tolerant to the cytotoxic uridine analog 5-fluorouridine than wild-type plants and the knockout plants of AtENT1 or AtENT6. Consistent with this observation, the AtENT3 knockout line exhibited a significantly decreased ability to take up [3H]uridine via the roots when compared with wild-type plants and the plants with mutated AtENT1 or AtENT6. This indicates that AtENT3, but not AtENTs 1 and 6, is the main transporter for uridine uptake in Arabidopsis roots. The transcription of AtENTs 1, 3, 4, 6, 7, and 8 was regulated in a complex manner during leaf development and senescence. In contrast, the six AtENT members were coordinately induced during seed germination. This work provides new information on the transport properties of recombinant AtENT proteins and new clues for future studies of the in vivo transport activities and physiological functions of the different ENT proteins in Arabidopsis plants.
Similar content being viewed by others
Introduction
Nucleoside transporters are conserved transmembrane proteins found in animals, yeast, fungus, higher plant, and several species of bacterium 1, 2, 3. They are involved in transporting nucleosides and analogs across cell membranes and are thus important for the salvage pathway of nucleotide synthesis 2, 3, 4. On the basis of the differences in primary structure and mechanism of action, nucleoside transporters have been divided into two families. The equilibrative nucleoside transporters (ENTs) have 11 transmembrane domains in their primary structure 1, 2, 5. Typical ENTs (such as hENTs 1 and 2 from human) transport substrates down their concentration gradients and do not rely on transmembrane pH gradient for their transport activities 1, 6, 7. But, recently, several ENTs from kinetoplastid protozoans (Leishmania donovani, Trypanosoma brucei), humans, and the higher plant Arabidopsis thaliana have been found to transport nucleosides in a pH-dependent manner 8, 9, 10, 11, 12, 13. The concentrative nucleoside transporters (CNTs), which generally have 13 transmembrane domains in their primary structure and are Na+ or H+-dependent symporters, transport substrates against existing concentration gradients 2, 14, 15. Although CNT transport activities have been found in a variety of animal tissues and cells 15, 16, bioinformatic analysis has not found CNT encoding genes in the completely sequenced Arabidopsis genome 1.
Seven ENT members are actively expressed in Arabidopsis 1, 17. Recombinant AtENT1 expressed in yeast cells has been found to transport adenosine, guanosine, and cytidine in a pH-dependent manner 10, 18. However, it has been suggested that AtENT1 may not transport uridine based on competition assay 10, although uridine is efficiently transported by most ENT proteins characterized to date 19, 20. The recombinant proteins of AtENTs 3, 4, 6, and 7 have all been found to transport adenosine, guanosine, cytidine, and uridine in a facilitative manner 17, 18. In contrast, the yeast-expressed AtENT8 is unable to mediate the uptake of any of the four nucleosides (adenosine, guanosine, cytidine, and uridine) tested so far 18. Compared to AtENTs 1, 3, 4, 6, 7, and 8, there is still no information on the transport function of recombinant AtENT2. It is also unknown whether the recombinant proteins of AtENT members are able to transport thymidine and by what mechanism(s) they do this. In addition to mediating the uptake of nucleosides, there is now genetic evidence that AtENTs 3 and 8 may be involved in transporting isopentenyladenine riboside (iPR) and trans-zeatin riboside (tZR), which are nucleoside-type cytokinins in higher plants 21. Interestingly, analysis of knockout mutant has indicated that AtENT1 does not display significant transport activity towards nucleoside-type cytokinin 21, although its transcripts are found abundantly in both vegetative and reproductive organs of Arabidopsis 17, 22. Apart from the analysis of Arabidopsis ENTs, a recent study has shown that rice may encode four ENT proteins, and the recombinant protein of one member (OsENT2) mediates the transport of nucleosides and nucleoside cytokinins when produced in yeast cells 23.
Comparative analysis of relative transcript levels has shown that the expression patterns of AtENT members in Arabidopsis vegetative and reproductive organs are highly complex, with all seven members expressed in leaves and flowers, four members (AtENTs 1, 3, 6, and 8) in roots, four members (AtENTs 1, 4, 6, and 8) in stems, and four members (AtENTs 1, 3, 6, and 8) in siliques 17. For the members found in the same organ, their transcript levels may also vary dramatically 17. This raises an important question on how the transport activities of different AtENT members, some of which have been demonstrated in vitro using recombinant proteins, are regulated in different organs or during different developmental processes in Arabidopsis plants. The answer to this question, which is essential for future studies of the physiological functions of different AtENT members, depends on a more complete understanding of the transport properties of recombinant AtENT proteins and the expression profiles of the AtENT members during important developmental processes, and the availability of efficient methods for determining the in vivo transport functions of different AtENT members in given Arabidopsis organs or developmental processes.
On the basis of the above information, the main objectives of this work were to generate new information on the transport activities of recombinant AtENT proteins and the transcriptional patterns of six AtENT members during leaf development and senescence and seed germination. Furthermore, effort was also made to investigate the ENT member(s) involved in uridine uptake in Arabidopsis roots, which led to the identification of AtENT3 as the main transporter for uridine uptake in this organ.
Materials and methods
Plant materials, general molecular methods, and radiochemicals
The Col-0 ecotype of Arabidopsis thaliana was used throughout this work. General conditions for Arabidopsis growth were described previously 17, 24. The standard medium for germinating Arabidopsis seeds and growing Arabidopsis plants on agar plates was 1/2 MS (pH 5.8) supplemented with 1% (w/v) sucrose and 0.7% (w/v) phytagar (Sigma). The seeds of the T-DNA insertion lines of AtENT1 (SALK_104866), AtENT3 (SALK_131585), and AtENT6 (SALK_088083) were obtained from Arabidopsis Biological Resource Center (http://www.biosci.ohio-state.edu/). Plants homozygous for T-DNA insertion were identified from the T3 population and confirmed in the T4 generation using genomic PCR (with appropriate oligonucleotide primers listed in Table 1) according to the methods detailed in the website http://signal.salk.edu/. For DNA and RNA manipulations, the molecular methods described by Sambrook and Russell 25 were generally followed. The tritiated nucleosides, uridine and thymidine, were purchased from Amersham Biosciences and were used immediately for uptake assays.
Plasmid construction and yeast transformation
Yeast expression vectors (p112AINE and p181AINE) and their uses in expressing recombinant AtENTs were described previously 17, 26, 27. The fui1 deletion strain (BY4741; MAT a; his3Δ1; leu2Δ0; met15Δ0; ura3Δ0; YBL042c::kanMX4) was employed for investigating uridine transport activities of recombinant AtENTs as reported before 17. The yeast strain BJ2168 (MATa; prc1-407; prb1-1222; pep4-3; leu2; trp1; ura3-52) (obtained from ATCC/Yeast Genetic Stock Center) was used for expressing AtENTs in the studies on thymidine transport. The coding sequences of AtENTs 1, 2, 3, 4, 6, 7, and 8 were amplified using high-fidelity DNA polymerase by PCR with the oligonucleotide primers listed in Table 1. The resulting fragments were each cloned into p181AINE after digestions by appropriate restriction enzymes, producing the yeast expression constructs pAtENT1, pAtENT2, pAtENT3, pAtENT4, pAtENT6, pAtENT7, and pAtENT8, respectively. For studying thymidine transport by recombinant AtENTs (see below), an additional construct expressing thymidine kinase (TK) was prepared. This was done by amplifying TK coding sequence from the mammalian cell expression vector pPNT 28 using appropriate oligonucleotide primers (Table 1) and cloning the resultant fragment into p112AINE, giving rise to the construct pTK. The correctness of all cloning experiments was verified using DNA sequencing. Yeast transformation was conducted using a lithium acetate method 29. In addition to developing the yeast strains expressing AtENTs, a recombinant yeast strain containing the empty p181AINE vector was also prepared and employed as the negative control in examining the transport activities of recombinant AtENTs. We were unable to test the potential transport activity of AtENT5 in this work, because its cDNA was not isolated in either our study 17 or that by other investigators 18. Our recent investigation using promoter-GUS transgenic plants showed that AtENT5 was not expressed in Arabidopsis plants under normal growth condition (unpublished data).
Thymidine and uridine transport by recombinant AtENTs
In yeast cells, de novo synthesis of dTMP can be inhibited by treatment with methotrexate (MTX) and sulfanilamide (SA) 30. It is also known that yeast cells lack an endogenous high-affinity thymidine uptake system and thymidine kinase 31. On the basis of these findings, a previous study has developed a reliable system for evaluating the thymidine transport activity of hENT1 by simultaneously expressing hENT1 and thymidine kinase, and examining yeast growth on the medium containing methotrexate and sulfanilamide and thymidine 32. In this work, we have adapted this system for investigating potential thymidine transport activities of recombinant AtENT proteins. For the growth assays, two sets of analyses were conducted. The first set employed the yeast cells containing p181AINE or the AtENT expression construct but not the construct expressing thymidine kinase. The second set used the yeast cells that were doubly transformed with the AtENT and thymidine kinase constructs. The recombinant cells were cultured in liquid medium at 30 °C to an OD600 of 2, serially diluted, and then spotted onto the selective plates (without leucine for the first set, and lacking both leucine and tryptophan for the second set) supplemented with 6 μg/ml SA, 50 mg/ml MTX, and 150 μM thymidine. The plates were incubated at 30 °C for three days before yeast growth was examined. Subsequent to the growth assay, direct uptake experiments were undertaken to characterize the pH dependency and kinetics of [3H]thymidine transport by AtENT members as previously described 17.
To examine potential uridine transport by AtENT1 expressed in yeast cells, the recombinant fui1 strains containing p181AINE, pATENT1, pAtENT3, or pAtENT8 were firstly grown on the plates without leucine (for selecting the plasmid construct) but supplemented with 500 μM 5-fluorouridine (Sigma). Because 5-fluorouridine is a cytotoxic uridine analog, growth inhibition in the presence of this drug would indicate potential uridine transport by the recombinant yeast strain. Recombinant AtENT3, but not AtENT8, has previously been found to transport uridine 17, 18. They were employed here as the positive and negative controls of uridine transport, respectively. After the growth assay, the pH dependency and kinetics of [3H]uridine transport by recombinant AtENT1 were further analyzed using direct uptake assays as detailed previously 17.
Investigation of AtENT member(s) involved in uridine transport in Arabidopsis roots
Two sets of experiments were conducted in this investigation. In the first set of experiments, the seeds of wild-type Arabidopsis plants and the homozygous T-DNA knockout mutant plants of AtENT1, AtENT3 or AtENT6 (designated as atent1-1, atent3-1 and atent6-1, respectively) were germinated on either the standard medium or the standard medium supplemented with various concentrations (10 to 400 μM) of 5-fluorouridine. The germination and seedling growth were inspected at weekly intervals. Seed germination and seedling survival in the presence of higher concentrations of 5-fluorouridine would indicate decreased uptake of this cytotoxic uridine analog via the root system. The germination and growth experiment was also conducted with the pH of the MS medium adjusted to pH 4.8 in order to facilitate uridine uptake into Arabidopsis roots through the pH dependent mechanism. In the second set of experiments, wild-type plants and the atent1-1, atent3-1 and atent6-1 plants were compared for their abilities to take up tritiated uridine via the root system. Wild-type plants and the knockout mutants were grown vertically on the standard medium for three weeks. Four samples, each composed of 20 intact roots, were prepared from the vertically grown wild-type plants and the three knock-mutants, respectively. The four samples were each incubated in 20 ml of 5 μM Mes/KOH buffer (pH 5.5) containing 5 mM [3H]uridine on a horizontal shaker as described previously 18, 33. At 2, 4, 6 and 8 h after the start of the incubation, triplicate samples (100 μl each) were withdrawn from each incubation solution and their levels of radioactivity were determined using liquid scintillation counting (1450 Microbeta, Wallac Oy). The amounts of [3H]uridine uptake by the root samples were deduced based on the decreases of radioactivity in the incubation media 18.
Transcriptional patterns of six AtENT members during leaf development and senescence and seed germination
To assess the transcriptional patterns of AtENTs 1, 3, 4, 6, 7 and 8 during leaf development and senescence, leaves were collected from 5-week-old wild-type Arabidopsis plants before bolting, and were divided into six developmental stages: Y (young), FX (mature fully expanded), OS (onset of senescence, 5% leaf area showing yellow), ES (early stage of senescence, 25% leaf area showing yellow), MS (middle stage of senescence, 50% leaf area showing yellow), and LS (late stage of senescence, more than 75% leaf area showing yellow) 34. To compare the transcriptional patterns of the six AtENT members during seed germination, fresh wild-type Arabidopsis seeds were vernalized for 2 days at 4 °C before imbibitions. They were then germinated on the plates with standard medium. Samples were collected at days 1, 2, 3, 5, 7, 9 and 11 after the germination started 35. Total RNA samples were prepared using a commercial kit (RNeasy Plant Mini kit, Qiagen), further purified using RNase-free DNase (RNase-Free DNase set, Qiagen), and converted to cDNAs through reverse transcription. The cDNA contents of different reverse transcription reactions were normalized by amplifying tubulin transcripts 17. The relative transcript levels of the six AtENT members were then evaluated using normalized cDNA mixtures with different numbers of amplification cycles (24, 26, 28, and 30). The consistency and reliability of the obtained patterns were checked in at least three independent experiments. AtENT2 was not included in above transcriptional analysis because its transcription was too low to be monitored reliably in leaves 17, and was undetectable in germinating seeds (unpublished data).
Results
Demonstration of thymidine transport by recombinant AtENTs 3, 6, and 7 and uridine transport by recombinant AtENT1
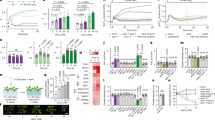
In the growth assay designed to study potential thymidine transport by recombinant AtENTs (with the endogenous dTMP synthesis in the yeast cells inhibited by drug treatment, and the presence of 150 μM thymidine in the medium), the yeast cells harboring single p181AINE or pAtENT constructs did not grow (Figure 1A). Coexpression with the pTK construct restored growth to the cells harboring pAtENT3, pAtENT6, or pAtENT7, but growth improvement of the pAtENT6 cells was less than that of the pAtENT3 or pAtENT7 cells (Figure 1A). In contrast, no growth was observed for the cells simultaneously expressing pTK and p181AINE, pAtENT1, pAtENT2, pAtENT4, or pAtENT8 (Figure 1A). This indicated that the recombinant proteins of AtENTs 3, 6, and 7, but not those of AtENTs 1, 2, 4 and 8, were capable of transporting thymidine when expressed in yeast cells. Furthermore, AtENT6 may be less efficient than AtENT3 and AtENT7 in mediating the transport of thymidine across yeast cell membrane. In direct uptake assays, recombinant AtENTs 3, 6 and 7 showed significant uptake of tritiated thymidine (Figure 1B), and thymidine uptake by the three AtENT members was all found to be resistant to inhibition by the protonophore CCCP (data not shown). Kinetic analysis showed that the Km and Vmax values of thymidine transport by recombinant AtENT3 were 2.3 μM and 328.49 pmol/mg protein/min, respectively (Figure 1C). The corresponding values determined for recombinant AtENT7 were 3.1 μM and 240.01 pmol/mg protein/min, respectively, and those for recombinant AtENT6 were 3.3 μM and 209.19 pmol/mg protein/min, respectively. Taken together, these results suggested that recombinant AtENTs 3, 6 and 7 could mediate high-affinity thymidine transport in a pH independent manner in yeast cells, but the affinity of thymidine transport by recombinant AtENT6 was relatively lower than that by recombinant AtENTs 3 or 7.
Analysis of potential thymidine transport by seven AtENT members. (A) Growth assay using serially diluted yeast cultures. The growth medium contained 6 mg/ml methotrexate, 50 mg/ml sulfanilamide, and 150 μM thymidine. In the absence of pTK (expressing thymidine kinase, left panel), no growth was observed in the yeast cells expressing the control vector p181AINE or the pAtENT constructs. In the presence of pTK (right panel), substantial growth was observed for the cells expressing pAtENTs 3, 6 or 7, but not those expressing p181AINE or pAtENTs 1, 2, 4 or 8. (B) Time course experiment. Compared to the cells expressing the control vector p181AINE, those expressing pAtENT3 exhibited significant transport of tritiated thymidine. The uptake rates and standard errors were calculated using the data from three independent assays. Similar results were obtained for the cells expressing pAtENT6 or 7 (data not shown). (C) Kinetics of thymidine transport into intact yeast cells expressing pAtENT3. Transport was allowed to proceed for 2 min in each of the indicated concentrations of tritiated thymidine. Uptake rates and standard errors were calculated using the data (after subtracting the uptake rates of the control cells expressing p181AINE) from three independent assays. The Km and Vmax values of thymidine transport by recombinant AtENT3 were 2.3 μM and 328.49 pmol/mg protein/min, respectively. Using the same protocol, the Km and Vmax values of thymidine transport by recombinant AtENT6 were determined to be 3.3 μM and 209.19 pmol/mg protein/min, respectively, whereas the corresponding values calculated for recombinant AtENT7 were 3.1 μM and 240.01 pmol/mg protein/min, respectively.
To examine potential uridine transport by recombinant AtENT1 more vigorously in this work, we conducted both growth and direct uptake assays. In the growth assay, AtENTs 3 and 8, whose uridine transport activity has been determined previously 17, 18, were employed as positive and negative controls of uridine transport, respectively. From the results shown in Figure 2A, it is clear that the fui1 deletion cells containing p181AINE or AtENT8 were resistant to growth inhibition by 5-fluorouridine, suggesting lack of uridine transport in these cells. In contrast, the growth of the fui1 deletion cells expressing pAtENT1 or pAtENT3 was inhibited by 5-fluorouridine (Figure 2A). Based on the degree of growth inhibition, the pAtENT3 cells appeared more susceptible to 5-fluorouridine than the pAtENT1 cells (Figure 2A). This indicated that uridine transport occurred in the fui1 cells expressing pAtENT1 or pAtENT3, and that uridine uptake by the pAtENT3 cells might be stronger than that in the pAtENT1 cells. In direct uptake experiments, the uptake of tritiated uridine by the pAtENT1 cells was significantly inhibited by CCCP (Figure 2B), indicating the requirement of transmembrane pH gradient for efficient uridine uptake by AtENT1 in yeast cells. Kinetics analysis revealed that the Km and Vmax values of uridine transport by AtENT1 in yeast cells were 3.9 μM and 102.37 pmol /mg protein/min, respectively (Figure 2C). This demonstrated that recombinant AtENT1 could mediate high-affinity uridine transport in a pH-dependent manner in yeast cells. However, uridine transport by AtENT1 may not be as strong as that by AtENT3, because the Km and Vmax values of uridine transport by the latter have previously been shown to be 3.2 μM and 232.50 pmol/mg protein/min, respectively 17. This may explain why the pAtENT3 cells were more susceptible to growth inhibition by 5-fluorouridine than the pAtENT1 cells observed in this work (Figure 2A).
Analysis of uridine transport by recombinant AtENT1. (A) Growth assay using serially diluted yeast cultures. The FUI1 deletion yeast cells expressing the control vector p181AINE or pAtENT8 grew extensively in the presence of the cytotoxic uridine analog 5-fluorouridine, indicating lack of uridine transport in these cells. In contrast, the growth of the FUI1 deletion cells expressing pAtENT1 or pAtENT3 was much limited, suggesting the occurrence of uridine transport by these cells. (B) Time course experiment. Compared to the FUI1 deletion cells expressing the control vector p181AINE, those expressing pAtENT1 exhibited significant transport of tritiated uridine. The uptake rates and standard errors were calculated using the data from three independent assays. Specific uridine transport mediated by recombinant AtENT1 was abolished in the presence of the protonophore CCCP. (C) Kinetics of uridine transport into intact yeast cells expressing pAtENT1. The conditions for the transport assay were the same as those described in Figure1. The Km and Vmax values of uridine transport by recombinant AtENT1 were determined to be 3.9 μM and 102.37 pmol/mg protein/min, respectively.
Analysis of AtENT member(s) involved in uridine transport in Arabidopsis roots
Although our previous study has shown that four AtENT members (1, 3, 6 and 8) are transcribed in wild-type Arabidopsis roots 17, knockout mutants were available for only AtENTs 1, 3 and 6, but not for AtENT8 21. Consequently, it was not possible to include AtENT8 in this analysis. RT-PCR experiment confirmed the lack of the transcripts for AtENTs 1, 3 or 6 in their respective knockout mutants (atent1-1, atent3-1 and atent6-1, Figure 3A). Furthermore, the knockout of one AtENT member did not affect the transcription of the other AtENT members (Figure 3A). When grown on the standard medium, the knockout mutants of AtENTs 1, 3, or 6 germinated and grew as normally as wild-type plants (Figure 3B). The presence of 10 μM 5-fluorouridine in the MS medium had a similar inhibitory effect on wild-type plants as well as the knockout mutants of AtENTs 1, 3 or 6 (Figure 3B). The presence of higher concentrations of 5-fluorouridine (= 20 μM) in the medium, although not affecting seed germination, strongly retarded the growth of wild-type plants and the knockout mutants of AtENT1 or AtENT6 (Figure 3B). In contrast, the growth of the AtENT3 knockout plants was much less inhibited (Figure 3B). The AtENT3 knockout plants still survived even with the concentration of 5-fluorouridine in the medium raised to 400 μM (data not shown). Considering that recombinant AtENT1 was shown to transport uridine in a pH-dependent manner in this work, the above growth experiment was repeated with the pH of the MS medium adjusted to 4.8. But identical results were obtained (data not shown).
Effect of the cytotoxic uridine analog 5-fluorouridine on the growth of wild-type (WT) plants and the knockout mutants of AtENTs 1, 3 or 6. (A) Lack of AtENT1, AtENT3 or AtENT6 transcripts in their respective knockout mutants (atent1-1, atent3-1, atent6-1). The knockout of one AtENT member did not affect the transcription of the other AtENT members. (B) In the absence of 5-fluorouridine, wild-type plants and the knockout mutants (atent1-1, atent3-1, atent6-1) were indistinguishable in their germination and growth. The presence of 10 μM 5-fluorouridine had a similar inhibitory effect on wild-type plants and the three knockout mutants. When 5-fluorouridine was supplied at higher concentrations of (≥20 μM), the growth of wild-type plants and the atent1-1 and atent6-1 mutants was more strongly inhibited than that of the atent3-1 mutant. Note that seed germination of wild-type plants and the three knockout mutants was not affected by 5-fluorouridine. The photographs were taken at 2 weeks after the start of seed germination.
The wild-type plants and the knockout mutants of AtENTs 1, 3 or 6 were also compared for their abilities to take up tritiated uridine in a direct uptake assay employing the roots. From Figure 4, it is clear that the level of [3H]uridine in the roots of atent3-1 plants was consistently and significantly lower than those in the roots of wild-type, atent1-1 or atent6-1 plants (Figure 4). Interestingly, the roots of wild-type, atent1-1 and atent6-1 plants did not differ extensively in their levels of [3H]uridine (Figure 4).
Uptake of tritiated uridine by the roots of wild-type plants and the knockout mutants of AtENT1, AtENT3 or AtENT6. Four samples (each consisted of 20 intact roots) were prepared from vertically grown wild-type plants and the three knock-mutants, respectively. They were then each incubated in 20 ml of 5 μM Mes/KOH buffer (pH 5.5) containing 5 μM [3H]uridine. Three aliquots (100 μl each) were withdrawn from each incubation solution at given time points, and their levels of radioactivity were determined using liquid scintillation counting. The amounts of [3H]uridine uptake by the root samples were deduced based on the loss of radioactivity in the incubation solutions.
Comparison of the transcriptional patterns of six AtENT members during leaf development and senescence and seed germination
Using semiquantitative (relative) RT-PCR, the transcriptional patterns of AtENTs 1, 3, 4, 6, 7 and 8 during leaf development and senescence and seed germination were evaluated. Reproducible transcriptional patterns were obtained in independent trials. A typical result on the transcriptional patterns of the six AtENT members during leaf development and senescence is shown in Figure 5. The transcription of AtENTs 1, 3, 6 and 8 was detected throughout leaf development and senescence, whereas that of AtENTs 4 and 7 was mainly found in the leaves that were mature or, at the onset, early and middle stages of senescence (Figure 5). Three AtENT members (1, 4 and 6) showed increased transcript levels during the development from immature to fully expanded leaves (Figure 5). On the other hand, the transcript levels of the six AtENT members were all generally up-regulated at the onset of leaf senescence (Figure 5). The transcript levels of AtENTs 1 and 3 remained high even at the final stage of leaf senescence (Figure 5).
Evaluation of the transcriptional patterns of six AtENT members during leaf development and senescence. Total RNA samples were prepared from Arabidopsis leaves at six different stages [Y (young), FX (mature fully expanded), OS (early stage or onset of senescence, 5% leaf area showing yellow), ES (middle stage of senescence, 25% leaf area showing yellow), MS (late stage of senescence, 50% leaf area showing yellow), and LS (final stage of senescence, more than 75% leaf area showing yellow)]. Following reverse transcription, the cDNA contents of individual reverse transcriptions were normalized by amplifying tubulin transcripts. The transcript levels of six AtENT members were then evaluated using normalized cDNA samples. The kinetics of PCR was monitored by amplifying tubulin transcripts using different numbers of amplification cycles (24, 26, 28, and 30). Identical transcriptional patterns were obtained in independent experiments.
A representative result on the transcript levels of the six AtENT members during seed germination is shown in Figure 6. The transcription of the six members was all induced during the early stage of seed germination (from day 1 to day 3, Figure 6), and their transcript levels were either increased steadily (for AtENTs 3, 4, 6, 7, and 8), or remained relatively constant (for AtENT1), as germination continued (Figure 6). The transcript levels of AtENTs 1, 3, 4 and 6 were generally higher than those of AtENTs 7 and 8 (Figure 6).
Evaluation of the transcriptional patterns of six AtENT members during seed germination. Total RNA samples were prepared from germinating Arabidopsis seeds (at days 1, 2, 3, 5, 7, 9 and 11 after the start of germination). The transcript levels of the six AtENT members were evaluated using the cDNA samples that had been normalized by amplifying tubulin transcripts. The kinetics of PCR was checked by amplifying tubulin transcripts using different numbers of amplification cycles (24, 26, 28, and 30). The transcriptional patterns shown were reproducible in independent assays.
Discussion
From the available literatures, there is ample evidence for thymidine uptake, transport between tissues or cells, and utilization in plant growth. For example, artificial feeding experiments have revealed that thymidine uptake is correlated with DNA synthesis 36, 37, 38, 39. Thymidine kinase activity, which is essential for thymidine salvage, is detected during period of intense growth (such as seed germination) 40, 41, 42. In sugarcane suspension cells and germinating zygotic embryos of white spruce, significant catabolism of thymidine is also observed 37, 39. However, against the background of the above studies, potential thymidine transporters in plant cells have not been characterized previously. Using growth and direct uptake assays, the present study has, for the first time, found that the recombinant proteins of AtENTs 3, 6 and 7, but not those of AtENTs 1, 2, 4 and 8, are capable of transporting thymidine with high affinity and in a pH-independent manner. The failure of recombinant AtENT8 in transporting thymidine is in line with its inability to transport adenosine, guanosine, cytidine, and uridine observed before 18, and further suggests that AtENT8 may be a transporter with highly specialized substrate specificity. On the other hand, lack of thymidine transport by recombinant AtENTs 1 and 4 is surprising, considering that the two proteins have previously been found to transport adenosine, guanosine, cytidine, and uridine in yeast cells [10, 18, this work]. Recombinant hENT1 has been found to transport a variety of nucleosides including thymidine 1, 32. By contrast, the FUI1 from Saccharomyces cerevisiae shows high uridine selectivity in its transport function 32. From these findings, it is clear that ENTs from different species (or even from the same species) may differ in their permeant specificities, although the structural basis responsible for such difference remains to be elucidated in future studies.
Previous investigation has indicated that AtENT1 expressed in yeast cells may not transport uridine 10. However, this suggestion was solely based on the competition of [3H]adenosine transport by unlabeled uridine using AtENT1 expressed in the ade2 yeast strain in which the FUI1 gene encoding the high-affinity uridine transporter (Km: 22 μM, 43) was not deleted. Under this condition, the competition of [3H]adenosine transport by unlabeled uridine may not be efficient because uridine might have been transported into the ade2 cells via FUI1. To exclude this possibility, we have chosen to use the FUI1 deletion strain to express AtENT1 in this work. Based on the growth assay (for sensitivity to 5-fluorouridine conferred by ectopic AtENT1 expression) and the time course and kinetics experiments (using direct uptake of [3H]uridine), we conclude that recombinant AtENT1 can mediate high-affinity uptake of uridine in a pH dependent manner. The finding of uridine transport by recombinant AtENT1 is in agreement with the notion that this nucleoside is an efficient permeant for most ENT proteins from eukaryotic cells, and can be transported by either typical ENTs (such as hENT1 from human and AtENT3 from Arabidopsis) in a facilitative manner or those ENTs (e.g., hENT3 from human and AtENT1 from Arabidopsis) that employ pH dependent mechanism during their function [1, 13, 17, 19, 32, this work].
The information on thymidine transport by recombinant AtENTs 3, 6 and 7 and uridine transport by recombinant AtENT1 in the present study complemented existing knowledge on nucleoside transport capabilities of different recombinant AtENT members. Based on the results gathered so far [10,17,18, this work], it is deduced that, in Arabidopsis, the transport of adenosine, guanosine, uridine, and cytidine may be more complex than that of thymidine, because the former may involve five AtENT members and two transport mechanisms (pH dependent transport by AtENT1 and facilitative transport by AtENTs 3, 4, 6 and 7), whereas the latter may employ three transporters (AtENTs 3, 6 and 7) and only the facilitative transport mechanism. In addition to the contribution discussed above, this work also represents the first attempt to investigate potential nucleoside transport by AtENT2 and showed that its recombinant protein did not transport thymidine. This may be caused by one or more of the following possibilities. Yeast expression system may not be suitable for revealing the transport function of AtENT2. Certain ENT protein (such as FUN26 from S. cerevisiae, 43) is not functional when produced in yeast cells, but displays clear nucleoside transport function when expressed in Xenopus oocytes. Consequently, it may be beneficial to use the Xenopus oocyte expression system in investigating the potential transport function of recombinant AtENT2. Alternatively, the permeant specificity of recombinant AtENT2 may be narrower compared to that of recombinant AtENTs 1, 3, 4, 6, and 7. Consequently, more nucleosides and analogs should be tested for AtENT2 transport in future experiments.
Beyond the studies using recombinant AtENT proteins, a major challenge ahead is to understand how the transport activities of the different AtENT members are regulated in vivo in different Arabidopsis organs or during important developmental processes. As the first steps to meet the above challenge, the present study set out to identify AtENT member(s) involved in uridine transport in Arabidopsis roots, and to compare the transcriptional patterns of six AtENT members during leaf development and senescence and seed germination. The fact that only the knockout mutant of AtENT3 could survive in the presence of higher concentrations of 5-fluorouridine (≥20 μM) indicates that the absence of AtENT3 is associated with decreased uptake of the uridine analog and hence tolerance to this toxic compound. On the other hand, the presence of AtENT3 in wild-type plants and the knockout mutants of AtENT1 or AtENT6 appears to confer uptake of 5-fluorouridine and much greater growth inhibition. Importantly, according to the differences in the levels of tritiated uridine accumulated in the roots during the in vivo uptake assay, the AtENT3 knockout mutant displays a significantly decreased ability to take up tritiated uridine via its roots when compared with wild-type plants or the knockout mutants of AtENT1 or AtENT6. Taken together, these results suggest that AtENT3 is a functional high-affinity uridine transporter in Arabidopsis roots. AtENTs 1 and 6 do not transport uridine as efficiently as AtENT3 in this organ. The high tolerance of the AtENT3 knockout mutant to 5-fluorouridine also indicates that AtENT8 (expressed in the atent3-1 background) may not play a significant role in the uptake of exogenous uridine via the roots. This may not be surprising since recombinant AtENT8 produced in yeast cells has been shown not to transport tritiated uridine 18. The finding that AtENTs 1 and 6 do not participate in uridine uptake significantly in Arabidopsis roots is, however, unexpected, considering that the recombinant proteins of the two AtENTs can mediate high-affinity uridine transport in yeast cells [18, this work]. One possibility is that the uridine uptake functions of the two members in Arabidopsis roots may be highly regulated. This suggestion may also be relevant to the observation that AtENT1 does not transport nucleoside-type cytokinin significantly in vivo 21, despite the fact that AtENT1 transcripts accumulate abundantly in both the vegetative and reproductive organs of Arabidopsis plants 17, 22. Nevertheless, the identification of AtENT3 as the main transporter for uridine uptake in Arabidopsis roots in this work demonstrates that the deployment of AtENT knockout mutants, coupled with an understanding of their transcriptional patterns, can lead to efficient methods for investigating the in vivo transport functions of different AtENT members in future studies.
Judging from the relative transcript levels, AtENTs 1, 3, 6, and 8 may be active throughout leaf development and senescence, and AtENTs 1 and 3 may be more dominant than AtENTs 6 and 8. In contrast to AtENTs 1, 3, 6 and 8, AtENTs 4 and 7 may mainly be active in the leaves that are mature or at the onset, early and middle stages of senescence. Despite the generalizations made above, the six AtENT proteins may differ from each other in the dynamic changes of their activities during the different stages of leaf development and senescence. Contrary to the complex activity patterns of AtENTs 1, 3, 4, 6, 7 and 8 during leaf development and senescence, the functions of the six members during seed germination may be coordinated. The six members are all induced during early germination, and their activities may remain high or increasing as germination continued. Assuming the six AtENT members exhibit one or more forms of transport activities towards nucleosides and/or analogs (i.e., nucleoside cytokinins) in vivo, leaf development and senescence may represent a developmental process with which the mechanism(s) that specify the differences in the transport functions of individual AtENT proteins can be studied. On the other hand, seed germination may be a suitable biological process for further investigations into the mechanism(s) that coordinate the transport functions of different ENT members. From the finding that higher concentration of 5-fluorouridine did not inhibit seed germination of wild-type plants and the knockout mutants of AtENTs 1, 3 or 6 in this work, it seems likely that the AtENT members induced during the early stage of seed germination may all be involved in transporting the nucleosides and/or analogs that are generated endogenously and utilized for early seedling growth. Further support for this proposition may come from the finding that nucleosides are the main tranported metabolites of nucleic acid catabolism in the cotyledons of germinating castor bean seeds 38.
In summary, this work has improved our understanding on the transport properties of recombinant AtENT proteins. But more importantly, we have embarked upon studying the in vivo transport functions of AtENT members. The identification of AtENT3 as the main transporter for uridine uptake in Arabidopsis roots opens the possibility to investigate more deeply into the molecular mechanism underlying nucleoside transport in plant cells. The revealing of the transcriptional patterns of the six AtENT members during leaf development and senescence and seed germination may provide model systems for future studies on the transport activities and physiological functions of different AtENT proteins in important developmental processes in Arabidopsis plants.
References
Hyde RJ, Cass CE, Young JD, Baldwin SA . The ENT family of eukaryote nucleoside and nucleobase transporters: recent advances in the investigation of structure/function relationships and the identification of novel isoforms. Mol Membr Biol 2001; 18:53–63.
Cabrita MA, Baldwin SA, Young JD, Cass CE . Molecular biology and regulation of nucleoside and nucleobase transporter proteins in eukaryotes and prokaryotes. Biochem Cell Biol 2002; 80:623–638.
Kong W, Engel K, Wang J . Mammalian nucleoside transporters. Curr Drug Metab 2004; 5:63–84.
Griffith DA, Jarvis SM . Nucleoside and nucleobase transport systems of mammalian cells. Biochim Biophys Acta 1996; 1286:153–181.
Baldwin SA, Beal PR, Yao SY, King AE, Cass CE, Young JD . The equilibrative nucleoside transporter family, SLC29. Pflugers Arch 2004; 447:735–743.
Griffiths M, Beaumont N, Yao SY, et al. Cloning of a human nucleoside transporter implicated in the cellular uptake of adenosine and chemotherapeutic drugs. Nat Med 1997; 3:89–93.
Acimovic Y, Coe I . Molecular evolution of the equilibrative nucleoside transporter family:identification of novel family members in prokaryotes and eukaryotes. Mol Biol Evol 2002; 19:2199–2210.
de Koning HP, Watson CJ, Jarvis SM . Characterization of a nucleoside/proton symporter in procyclic Trypanosoma brucei brucei. J Biol Chem 1998; 273:9486–9494.
Mäser P, Sütterlin C, Kralli A, Kaminsky RA . Nucleoside transporter from Trypanosoma brucei involved in drug resistance. Science 1999; 285:242–244.
Möhlmann T, Mezher Z, Schwerdtfeger G, Neuhaus HE . Characterization of a concentrative type of adenosine transporter from Arabidopsis thaliana (ENT1, At). FEBS Lett 2001; 509:370–374.
Sanchez MA, Tryon R, Green J, Boor I, Landfear SM . Six related nucleoside/nucleobase transporters from Trypanosoma brucei exhibit distinct biochemical functions. J Biol Chem 2002; 277:21499–21504.
Stein A, Vaseduvan G, Carter NS, Ullman B, Landfear SM, Kavanaugh MP . Equilibrative nucleoside transporter family members from Leishmania donovani are electrogenic proton symporters. J Biol Chem 2003; 278:35127–35134.
Baldwin SA, Yao SY, Hyde RJ, et al. Functional characterization of novel human and mouse equilibrative nucleoside transporters (hENT3 and mENT3) located in intracellular membranes. J Biol Chem 2005; 280:15880–15887.
Ritzel MW, Ng AM, Yao SY, et al. Molecular identification and characterization of novel human and mouse concentrative Na+-nucleoside cotransporter proteins (hCNT3 and mCNT3) broadly selective for purine and pyrimidine nucleosides (system cib). J Biol Chem 2001; 276:2914–2927.
Gray JH, Owen RP, Giacomini KM . The concentrative nucleoside transporter family, SLC28. Pflugers Arch 2004; 447:728–734.
Ritzel MW, Ng AM, Yao SY, et al. Recent molecular advances in studies of the concentrative Na+-dependent nucleoside transporter (CNT) family: identification and characterization of novel human and mouse proteins (hCNT3 and mCNT3) broadly selective for purine and pyrimidine nucleosides (system cib). Mol Membr Biol 2001; 18:65–72.
Li G, Liu K, Baldwin SA, Wang D . Equilibrative nucleoside transporters of Arabidopsis thaliana. cDNA cloning, expression pattern, and analysis of transport activities. J Biol Chem 2003; 278:35732–35742.
Wormit A, Traub M, Florchinger M, Neuhaus HE, Möhlmann T . Characterization of three novel members of the Arabidopsis thaliana equilibrative nucleoside transporter (ENT) family. Biochem J 2004; 383:19–26.
Cass CE, Young JD, Baldwin SA . Recent advances in the molecular biology of nucleoside transporters of mammalian cells. Biochem Cell Biol 1998; 76:761–770.
Vickers MF, Zhang J, Visser F, et al. Uridine recognition motifs of human equilibrative nucleoside transporters 1 and 2 produced in Saccharomyces cerevisiae. Nucleosides Nucleotides Nucleic Acids 2004; 23:361–373.
Sun J, Hirose N, Wang X, et al. Arabidopsis SOI33/AtENT8 gene encodes a putative equilibrative nucleoside transporter that is involved in cytokinin transport in planta. Journal of Integrative Botany 2005; 47:588–603.
Li J, Wang D . Cloning and in vitro expression of the cDNA encoding a putative nucleoside transporter from Arabidopsis thaliana. Plant Sci 2000; 157:23–32.
Hirose N, Makita N, Yamaya T, Sakakibara H . Functional characterization and expression analysis of a gene, OsENT2, encoding an equilibrative nucleoside transporter in rice suggest a function in cytokinin transport. Plant Physiol 2005; 138:196–206.
Hu Y, Bao F, Li J . Promotive effect of brassinosteroids on cell division involves a distinct CycD3-induction pathway in Arabidopsis. Plant J 2000; 24: 693–670.
Sambrook J, Russell RW . Molecular cloning: A laboratory manual, 3rd ed. New York : Cold Spring Harbor Laboratory Press, 2001.
Riesmeier JW, Willmitzer L, Frommer WB . Isolation and characterization of a sucrose carrier cDNA from spinach by functional expression in yeast. EMBO J 1992; 11:4705–4713.
Daram P, Brunner S, Persson BL, Amrhein N, Bucher M . Functional analysis and cell-specific expression of a phosphate transporter from tomato. Planta 1998; 206:225–233.
Tybulewicz VL, Crawford CE, Jackson PK, Bronson RT, Mulligan RC . Neonatal lethality and lymphopenia in mice with a homozygous disruption of the c-abl proto-oncogene. Cell 1991; 65:1153–1163.
Gietz D, St Jean A, Woods RA, Schiestl RH . Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res 1992; 20:1425.
McNeil JB, Friesen JD . Expression of the herpes simplex virus thymidine kinase gene in Saccharomyces cerevisiae. Mol Gen Genet 1981; 184:386–393.
Hogue DL, Ellison MJ, Young JD, Cass CE . Identification of a novel membrane transporter associated with intracellular membranes by phenotypic complementation in the yeast Saccharomyces cerevisiae. J Biol Chem 1996; 271:9801–9808.
Vickers MF, Mani RS, Sundaram M, et al, Cass CE . Functional production and reconstitution of the human equilibrative nucleoside transporter (hENT1) in Saccharomyces cerevisiae. Interaction of inhibitors of nucleoside transport with recombinant hENT1 and a glycosylation-defective derivative (hENT1/N48Q). Biochem J 1999; 339:21–32.
Narang RA, Bruene A, Altmann T . Analysis of phosphate acquisition efficiency in different Arabidopsis accessions. Plant Physiol 2000; 124:1786–1799.
Gepstein S, Sabehi G, Carp MJ, et al. Large-scale identification of leaf senescence-associated genes. Plant J 2003; 36:629–642.
Schmidt A, Su YH, Kunze R, et al. UPS1 and UPS2 from Arabidopsis mediate high affinity transport of uracil and 5-fluorouracil. J Biol Chem 2004; 279:44817–44824.
Ashihara H . Changes in activities of the de novo and salvage pathways of pyrimidine nucleotide biosynthesis during germination of black gram (Phaseolus mungo) seeds. Z Pflanzenphysiol 1977; 81:199–211.
Lesley S, Maretzki A, Nickell LG . Incorporation and degradation of 14C and 3H-labeled thymidine by sugarcane cells in suspension culture. Plant Physiol 1980; 65:1224–1228.
Kombrink E, Beevers H . 1983. Transport of purine and pyrimidine bases and nucleosides from endosperm to cotyledons in germinating castor bean seedlings. Plant Physiol 1980; 73:370–376.
Stasolla C, Loukanina N, Ashihara H, Yeung EC, Thorpe TA . Pyrimidine nucleotide biosynthesis and nucleic acid metabolism in embryos and megagametophytes of white spruce (Picea glauca) during germination. Physiol Plant 2002; 115:155–165.
Stasolla C, Loukanina N, Ashihara H, Yeung EC, Thorpe TA . Pyrimidine deoxyribonucleotide metabolism during maturation and germination of white spruce (Picea glauca) somatic embryos: metabolic fate of C-labeled cytidine, deoxycytidine and thymidine. Physiol Plantarum 2003b; 118:499–506.
Wanka F, Walboomers JM . Thymidine kinase and uridine kinase in corn seedlings. Z Pflanzenphysiol 1966; 55:458–465.
Wanka F, Bauer FW . On phosphorylation and deamination of pyrimidine and deoxy-pyrimidine nucleosides by enzymes from germinating corn seedlings. Z Pflanzenphysiol 1967; 58:175–186.
Vickers MF, Yao SY, Baldwin SA, Young JD, Cass CE . Nucleoside transporter proteins of Saccharomyces cerevisiae. Demonstration of a transporter (FUI1) with high uridine selectivity in plasma membranes and a transporter (FUN26) with broad nucleoside selectivity in intracellular membranes. J Biol Chem 2000; 275:25931–25938.
Acknowledgements
We thank Drs Yi Ping Tong and Hong Jie Li for comments on the manuscript. This work was supported by grants from the National Natural Science Foundation of China (39770491) and the Chinese Academy of Sciences (KSCX2-SW-304).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chen, K., Xu, M., Li, G. et al. Identification of AtENT3 as the main transporter for uridine uptake in Arabidopsis roots. Cell Res 16, 377–388 (2006). https://doi.org/10.1038/sj.cr.7310049
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7310049
Keywords
This article is cited by
-
Advances in Arabidopsis research in China from 2006 to 2007
Chinese Science Bulletin (2007)