ABSTRACT
Wheat 1BL/1RS translocations are widely planted in China as well as in most of the wheat producing area in the world for their good qualities of disease resistance and high yield. 1BL/1RS translocations are however poor in bread making, partially caused by a family of small monomeric proteins, ω-secalins, which are encoded by genes on 1RS. Based on published sequence of a rye ω-secalin gene we designed a pair of primers to cover the whole mature protein coding sequence. A major band could be amplified from 1BL/1RS translocations but not from euploid wheat. Using this primer set we conducted PCR amplification by using high fidelity Pfu polymerase on the genomic DNAs and cDNAs purified from a 1BL/1RS translocation Lankao 906. Sequencing analysis indicated that this gene family contains several members of 1150 bp, 1076 bp, 1075 bp, 1052 bp and 1004 bp genes, including two pseudogenes and three active genes. The gene transcripts were differentially expressed in developing seeds.
Similar content being viewed by others
INTRODUCTION
The short arm of rye 1R chromosome contains several disease resistance genes such as leaf rust, stem rust, stripe rust and powdery mildew. The 1BL/1RS translocation, involving the short arm of rye chromosome 1R and the long arm of wheat chromosome 1B was originally selected for its resistance to diseases 1, 2. Since 1BL/1RS translocations were introduced into China during early 70s last century, they have been widely used in wheat breeding programs. Statistic data showed that among 179 wheat cultivars and recently bred wheat lines 38% are 1BL/1RS translocations. In some major wheat growing area 1BL/1RS translocations account for up to 59% of the total wheat cultivars 3. Except of the advantage in disease resistance, the translocation is also useful for its positive effects on agronomic traits including yield performance, yield stability and wide adaptation 4, 5, 6, 7, 8, 9, 10. Furthermore, 1BL/1RS translocations carrying new disease resistance characteristics have been developed via wide crosses between wheat and rye or triticale. The 1BL/1RS translocation became one of the most frequently used alien introgressions in wheat breeding programs throughout the world 11. However, serious defects in bread processing such as poor mixing tolerance, superficial dough stickness and low bread volume have been brought about by the translocation 12, 13, 14, 15, 16. Recently a correlation between the translocation and the poor qualities in noodle processing has also been reported 17.
Locus Sec-1 in 1RS chromosome arm encodes two kinds of seed storage proteins ω-secalins and 40K γ-secalins 18, 19. ω-secalins and 40K γ-secalins are separated groups of genes although they are tightly linked to each other 20. It is believed that the poor quality of 1BL/1RS translocations in bread processing is partially caused by the expression of ω-secalins, which are a family of small monomeric proteins related to wheat ω-gliadin 14. Up to date only three ω-secalin genes, two from a rye cv and one from a wheat/rye translocation have been reported 21, 22. Lankao 906 is a high yield wheat 1BL/1RS translocation with high resistance to rust and powdery mildew. Its bread making quality is however poor. The objective of the present study is to use homoeologous cloning method to clone the ω-secalin gene family from Lankao 906 and to analyze the gene family from molecular level. This will provide useful information for the future production of quality wheat 1BL/1RS translocations.
MATERIALS AND METHODS
Plant materials
Three wheat 1BL/1RS translocations Lankao 906, Bobwhite, R59 and six non 1BL/1RS translocations Chinese Spring, Zhongyou 9507, Gaocheng 8901, Yangmai 158, Yangmai 10 and Wenmai 6 were used.
Extraction of DNA and RNA
CTAB method 23 was used to isolate total DNA from young leaves. Total RNA was extracted from developing seeds about 15 d post anthesis with TRIzol method (Tianweishidai Company, China).
Synthesis of cDNA
cDNA was synthesized from above total RNA by using the Super ScriptTM II first strand synthesis system (Invitrogen).
PCR
PCR primers was designed using DNAStar software based on the published sequence from a ω-secalin gene present on the short arm of 1RS 22. The primers covered the whole mature protein coding sequence and most of the signal peptide coding region. The primer sequences were as follows: ω-sec-P1 (sense) 5′-accttcctcatctttgtcct-3′, ω-sec-P2 (antisense) 5′-ccgatgcctataccactact-3′. PCR reaction was carried out by using Pfu PCR Kit KP-101 (Tianweishidai company) and PTC-100™ Programmable Thermal Controller (MJ Research) with the following program: 3 min at 94°C (initial denaturation), and 30 cycles of 45 sec at 94°C, 45 sec at 65°C, and 1.5 min at 72°C. Additional 7 min extension was followed after the 30 cycles. The 50 μl reaction volume contained 20 mM Tris-HCl (pH8.8), 10 mM KCl, 2 mM MgSO4, 0.2 mM of each dNTP, 0.4 μM of each sense and antisense primers, 3.75 U Pfu DNA polymerase and about 100 ng of template DNA or 1 μl cDNA from the cDNA synthesis system. Partial PCR products were subjected to electrophoresis in 1×TAE buffer (40 mM Tris-Acetate, 1 mM EDTA) on 1% agarose gel, and visualized by ethidium bromide staining.
Molecular cloning and DNA sequencing of PCR products
PCR products were purified with PCR product purification kit DP-204 (Tianweishidai Company). The purified PCR products were then cloned into pGEM-T Easy vector with blunt end DNA fragments cloning kit VT405 (Tianweishidai Company). Transformation was performed with TOP10 competent E. coli cells by heat shock method. Positive clones were selected by Colony PCR and their plasmids were extracted with plasmid extraction kit DP-103 (Tianweishidai Company). Sequencing of PCR products was carried out from both ends of the inserted fragments with T7 and SP6 primers respectively. Nucleotide sequences of the cloned fragments were finally determined with a 3730 DNA Sequencer (ABI) and the sequence data were analyzed with a computer program DNAStar.
Southern and Northern blotting
For southern blot analysis 15 μg genomic DNA was digested with BamH I and EcoR I respectively. For northern blot analysis 10 μg total RNA was used. DNA was transferred to nylon membrane from agarose gel as described by Read and Mann 24. The transfer of RNA from agarose gel to the membrane was performed with formaldehyde method according to Sambrook and Russell 25. One of ω-secalin sequences 1076 bp-1 was used to prepare probes with oligo labeling method as described by Feinberg and Vogelstein 26.
RESULTS
Primer specificity
Based on the sequence of published ω-secalin gene 22, a 1076 bp fragment should be amplified with primers ω-sec-P1 and ω-sec-P2 as described in Materials and Methods. As expected a major band above 1 kb was amplified from genomic DNAs of all of the three 1BL/1RS translocations and no amplification occurred in all of the six non 1BL/1RS wheat cultivars (Fig. 1), indicating that the primers could be used to amplify the ω-secalin genes specifically.
Sequences of ω-secalin genes from Lankao 906
The genomic DNA and the cDNA from Lankao 906 were used as templates to conduct PCR amplification by using high fidelity Pfu polymerase. The PCR products were checked by electrophoresis (Fig. 2), purified and cloned into pGEM-T Easy plasmid. Forty two genomic and sixty nine cDNA clones were selected for sequencing. Among these clones except of those 1076bp fragments that have the same size of the target sequence, there were many clones containing shorter insertions and only one clone with longer insertion. The shortest fragment amplified from cDNA was only 158 bp, while the longest fragment was 1150 bp amplified from genomic DNA. Except of those 1076 bp fragments, the clones containing fragment of 1004 bp represent the main population (Tab. 1).
Sequence comparisons between different length genomic clones
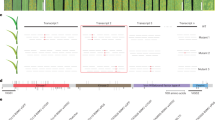
In comparison with the 1076bp target sequence, all of the shorter clones contain a deletion in different regions. Fig. 3 shows comparison of some representative sequences. All of the 1075bp clones lacked one base at the position 34. All of the 1052 bp clones were deletions of a 24 bp fragment from 410 to 423 base pairs. Most of the 1004 bp clones lacked a 72 bp fragment from nucleotides 916 to 987, except that one clone contains deletion of nucleotides 906–977. All the clones less than 1000 bp contain deletions at different positions. The 1150bp clone contains one base pair deletion at nucleotide 704 and three insertions of 45, 12 and 18 base pairs at positions of 1016, 1035 and 1050 respectively (Fig. 4).
There is no switch of reading frame for clones 1052 and 1004 bp in comparison with the 1076 bp target sequence. However the deletion of the one base pair in both 1075 bp and 1150 bp clones caused premature protein termination (Fig. 4 and 5).
The genomic clones containing similar lengths are nearly identical in sequences. For example, eleven sequences from thirteen 1076 bp clones showed 97 to 99.9% sequence identity in nucleotides. The minor difference may be contributed by errors occurred during PCR amplification.
Comparison of cDNA sequences to the genomic sequences
Four groups of cDNA clones have been obtained, ranging from 1076 bp, 1052 bp, 1051 bp to 1004 bp. As expected, the target 1076 bp cDNA clone is identical to its corresponding genomic clone. Furthermore, all of the 1052 bp and 1004 bp cDNA clones have identical respective genomic clones, suggesting that these clones are representing the real genes. Compared to the 1052 bp clone, the 1051 bp sequence contained one more deletion at position 395. Since only one 1051 bp cDNA clone but no corresponding genomic clone has been found, it is likely that the 1051 bp cDNA clone is a PCR product occurred during amplification rather than a real gene transcript.
The 15 genomic clones less than 1000 bp do not have any corresponding cDNA clones, suggesting that these short sequences may be produced during PCR amplification. Indeed, when the 1076 bp target gene was used as a template for PCR reaction, about 6% of the PCR products are incomplete sequences. Similarly, cDNA clones shorter than 1 kb do not show any corresponding genomic clones. Thus the clones shorter than 1 kb are not the real gene products of ω-secalin family.
Southern and Northern blotting analyses
To further characterize ω-secalin gene family in 1BL/1RS translocation lines, Southern blot was performed on Lankao 906. As shown in Fig. 6, in 1BL/1RS translocation line Lankao 906 three to four major bands ranging from 2.2 kb to 10 kb could be detected from BamH I and EcoR I digestions respectively, indicating that there are at least three ω-secalin related genes in 1RS. The 10 kb and the 2.2 kb fragment from respective BamH I and EcoR I digestions show higher intensity than other bands (Fig. 6). Non 1BL/1RS translocation line Chinese Spring was used as a control (Fig. 6A).
Total RNAs from seeds of 1BL/1RS translocation line Lankao 906 (Fig. 7, line B) and non 1BL/1RS translocation line Chinese Spring (Fig. 7, line A) were analyzed on Northern blot, probed with the 1076 bp ω-secalin gene sequence. At least two populations of transcripts can be found in Lankao 906, while the transcript with higher molecular weight exhibits higher expression level (Fig. 7, line B). In Chinese Spring, a non 1BL/1RS translocation line, two bands above the molecular weight of transcripts detected in Lankao 906 can also be found, which may be contributed by wheat ω-gliadins that have higher molecular weight than rye ω-secalins (Fig. 7). Compared to Chinese Spring, Lankao 906 gave stronger bands hybridized to ω-secalin sequence as shown by Southern blot (Fig. 6) but weaker gene expression as shown by Northern blot (Fig. 7). Whether ω-secalin gene is preferentially expressed during different developmental stages needs to be further analyzed.
DISCUSSION
In this report, we use homoeologous cloning method to clone ω-secalin gene family from a 1BL/1RS wheat translocation line. At present, three 1076 bp ω-secalin genes have been reported 21, 22. Except of the 1076 bp sequences, we have found additional clones containing 1150 bp, 1075 bp, 1052 bp and 1004 bp sequences. Among these clones, the ones of 1076-, 1052- and 1004-bp sequences may represent the real genes of ω-secalin family since there are corresponding genomic and cDNA clones. The 1075 bp and the 1150 bp genomic clones may be pseudogenes since no corresponding cDNA clones can be found. Identical 1150 bp sequence has been also obtained by Clarke BC (personal communication), it is thus believed to be a real ω-secalin gene. In comparison with the 1076 bp ω-secalin sequence, the ORF in the 1150 bp clone has been changed, due to one base pair deletion that leads to a premature termination.
On the other hand, a single 1051 bp cDNA sequence obtained has no corresponding genomic clone, suggesting that it may be produced during PCR amplification instead of a real gene.
To support the homoeologous cloning results, our Southern blot analysis demonstrates that there are probably at least three ω-secalin genes located on 1RS. At least two ω-secalin genes are actively transcribed in the developing seeds of Lankao 906. The transcript with higher molecular weight expressed more.
The published three 1076 bp ω-secalin gene sequences are not 100% matched to our 1076 bp sequences, which may be due to the difference in materials used by different groups. Two sequences published by Hull 22 were from rye cv Gazelle and another sequence published by Clarke 21 was from a wheat translocation line containing a small interstitial segment from rye cv Imperial. The wheat 1BL/1RS translocation line we used is a progeny of a wide cross between a wheat cultivar with a hexaploid triticale cv Mzalenod Beer.
The 1004 bp and 1052 bp ω-secalin gene sequences have not been reported. It is unlikely that these sequences are derived from PCR amplification based on our observations that these sequences are discovered from both the genomic and cDNA clones with high frequencies up to 33% of the total clone sequenced. Besides, we could never detect these sequences when the 1076 bp clone was used as template. It is very much likely that the 1004 bp and the 1052 bp sequences are the real genes of ω-secalin family. Furthermore, proteins translated from both the 1004 bp and the 1052 bp genes share about 97% sequence identity to that from the 1076 bp gene, suggesting that they belong to the ω-secalin gene family and may play similar function.
In summary, by homoeologous cloning we have found five populations of ω-secalin genes, representing three actively transcribed genes, the 1076 bp, 1052 bp, 1004 bp sequences and two pseudogenes, the 1150 bp and the 1075 bp sequences. By Southern and Northern analyses, different populations of ω-secalin gene family have been found. The ω-secalin genes are differentially expressed in developing seeds. The biological function of the differential expression of ω-secalin genes needs to be further analyzed. An open question is whether suppression of ω-secalin gene expression would improve the wheat quality of 1BL/1RS translocations in bread making.
References
Zeller FJ . 1B/1R wheat-rye chromosome substitutions and translocations. In: Sears ER, Sears LMS, ed. Proc Intl Wheat Genet Symp, 4th, 1973; 209–21. Columbia, MO. 6–11 Aug. 1973. Missouri Agric Exp Stn, Columbia.
Zeller FJ, Hsam SLK . Broadening the genetic variability of cultivated wheat by utilizing rye chromatin. In: Sakamoto S, ed. Proc Intl Wheat Genet Symp, 6th, 1984; 161–73. Kyoto, Japan. 28 Nov.–3 Dec. 1983. Plant Germplasm Int, Kyoto Univeristy, Kyoto, Japan.
Zhou Y, He ZH, Zhang GS, et al. Utilization of 1BL/1RS translocation in wheat breeding in China. Acta Agronomica Sinica 2004; 30:531–5.
Carver BF, Rayburn AL . Comparison of related wheat stocks possessing 1B or 1BL. 1RS chromosomes: Agronomic performance. Crop Sci 1994; 34:1505–10.
McKendry AL, Tague DN, Miskin KE . Effect of 1BL. 1RS on agronomic performance of soft red winter wheat. Crop Sci 1996; 36:844–7.
Moreno-Sevilla B, Baenziger PS, Peterson CJ, Graybosch RA, McVey DV . The 1BL. 1RS translocation: Agronomic performance of F3-derived lines from a winter wheat cross. Crop Sci 1995; 35:1051–5.
Rajaram S, Mann CE, Ortiz-Ferrara G, Mujeeb-Kazi A . Adaptation, stability and high yield potential of certain 1RS. 1BL CIMMYT wheats. In: Sakamoto S, ed. Proc 6th Int Wheat Genetic Symp, 1984; 613–21. Kyoto, Japan. 28 Nov–3 Dec. 1983. Plant Germplasm Inst, Kyoto Univ, Kyoto, Japan.
Schlegel R, Meinel A . A quantitative trait locus (QTL) on chromosome arm 1RS of rye and its effect on yield performance of hexaploid wheats. Cereal Res Commun 1994; 22:7–13.
Villarreal RL, Rajaram S, Mujeeb-Kazi A, Toro E . The effect of chromosome 1RS. 1BL translocation on the yield potential of certain spring wheat (Triticum aestivum L.). Plant Breed 1991; 106:77–81.
William MDHM, Mujeeb-Kazi A . Rapid detection of 1B, 1BL/1RS heterozygotes in the development of homozygous 1BL/1RS translocation stocks of Triticum turgidum (2n = 4x = 28). Genome 1993; 36:1088–91.
Braun HJ, Payne TS, Morgounov AI, van Ginkel M, Rajaram S . The challenge: One billion tons of wheat by 2020. In: Slinkard AE, ed. Proc. 9th Int. Wheat Genet Symp, 33–40. Saskatoon, Canada. 2-7 Aug. 1998. Univ Ext Press, Univ of Saskatchewen, Saskatoon, SK, Canada.
Burnett CJ, Lorenz KJ, Carver BF . Effect of the 1B/1R translocation in wheat on composition and properties of grain and flour. Euphytica 1995; 86:159–66.
Dhaliwal AS, Mares DJ, Marshall DR . Effect of 1B/1R chromosome translocation on milling and quality characteristics of bread wheats. Cereal Chem 1987; 64:72–6.
Dhaliwal AS, Mares DJ, Marshall DR . Measurement of dough surface stickiness associated with the 1B/1R chromosome translocation in bread wheats. Cereal Sci 1990; 12:165–75.
Lee JH, Graybosch RA, Peterson CJ . Quality and biochemical effects of a 1RS. 1BL wheat-rye translocation in wheat. Theor Appl Genet 1995; 90:105–12.
Seo YW, Graybosch RA, Peterson CJ, Shelton DR . Assessment of enzyme-linked immunoassay of rye secalins as a tool in the prediction of 1RS wheat quality. Cereal Chem 1995; 72:252–4.
Liu JJ, He ZH, Pena RJ, Zhao ZD . The effects of 1B/1R translocation on grain quality and noodle quality of bread wheat. Acta Agronomica Sinica 2004; 30:149–53.
Lawrence GJ, Shepherd KW . Chromosomal location of genes controlling seed proteins in species related to wheat. Theor Appl Genet 1981; 59:25–31.
Shewry PR, Bradberry D, Franklin J, White RP . The chromosomal locations and linkage relationships of the structural genes for the prolamin storage proteins (secalins) of rye. Theor Appl Genet 1984; 69:63–9.
Carrillo JM, Vazquez JF, Orellana J . Linkage relationships between the loci Sec-1 and Sec-3 in rye (Secale cereale L.). Heredity 1990; 64:125–30.
Clarke BC, Mukai Y, Apples R . The Sec-1 locus on the short arm of chromosome 1R of rye (Secale cereale). Chromosoma 1996; 105:269–75.
Hull GA, Halford NG, Kreis M, Shewry PR . Isolation and characterization of genes encoding rye prolamins containing a highly repetitive sequence motif. Plant Mol Biol 1991; 17:1111–5.
Saghai-Maroof MA, Soilman KM, Jorgensen RA, Allard RW . Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc Nat Acad Sci U S A 1984; 81:8014–8.
Reed KC, Mann DA . Rapid transfer of DNA from agarose gels to nylon membranes.Nucleic Acids Res 1985; 13:7207–21.
Sambrook J, Russell DW . Molecular cloning: A Laboratory Manual, 3rd ed. Cold Spring Harbor Laboratory Press. 2001:540–52.
Feinberg AP, Vogelstein B . A technique for radiolabelling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 1983; 132:6–13.
Acknowledgements
This research was supported by the National Basic Research Program of China (973) (No. 2004CB117200).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
CHAI, J., LIU, X. & JIA, J. Homoeologous cloning of ω-secalin gene family in a wheat 1BL/1RS translocation. Cell Res 15, 658–664 (2005). https://doi.org/10.1038/sj.cr.7290335
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290335
Keywords
This article is cited by
-
Genomic and functional genomics analyses of gluten proteins and prospect for simultaneous improvement of end-use and health-related traits in wheat
Theoretical and Applied Genetics (2020)
-
Genetic diversity in common wheat lines revealed by fluorescence in situ hybridization
Plant Systematics and Evolution (2019)
-
Characterization of ω-secalin genes from rye, triticale, and a wheat 1BL/1RS translocation line
Journal of Applied Genetics (2010)