ABSTRACT
DNA degradation is a biochemical hallmark in apoptosis. It has been demonstrated in many cell types that there are two stages of DNA fragmentation during the apoptotic execution. In the early stage, chromatin DNA is cut into large molecular weight DNA fragments, although the responsible nuclease(s) has not been recognized. In the late stage, the chromatin DNA is cleaved further into short oligonucleosomal fragments by a well-characterized nuclease in apoptosis, the caspase-activated DNase (CAD/DFF40). In this study, we demonstrate that large molecular weight DNA fragmentation also occurs in Xenopus egg extracts in apoptosis. We show that the large molecular weight DNA fragmentation factor (LDFF) is not the Xenopus CAD homolog XCAD. LDFF is activated by caspase-3. The large molecular weight DNA fragmentation activity of LDFF is Mg2+-dependent and Ca2+-independent, can occur in both acidic and neutral pH conditions and can tolerate 45°C treatment. These results indicate that LDFF in Xenopus egg extracts might be a new DNase (or DNases) responsible for the large DNA fragmentation.
Similar content being viewed by others
INTRODUCTION
Apoptosis, a morphologically distinct form of programmed cell death, has attracted considerable attention for its implicated roles in embryonic development, tissue homeostasis and protection against diseases. A cell in apoptosis undergoes cell shrinkage, cytoskeleton collapse, nuclear envelope disassembly, chromatin condensation into apoptotic bodies, DNA fragmentation, protein degradation, and so on. DNA fragmentation is considered to be the biochemical hallmark for this suicidal process. In most apoptotic cell types, there are two stages of DNA degra-dation. In the early stage, chromatin DNA is cleaved into large molecular weight chromatin DNA fragments. In the late stage, the fragmented chromatin DNA is digested further into short oligonucleosomal DNA fragments by the nucleases1, 2; however, in some cell types only large molecular weight fragments were observed3, 4. So far, more than twenty nucleases have been identified and impli-cated as being responsible for the late stage DNA degra-dation, such as DFF40/CAD/CPAN5, 6, 7, DNase I8, inducible-lymphocyte Ca2+/Mg2+-dependent endonuclease9, cyclophilins10, NUC-111, endonuclease G12, DNA topoisomerase IIα and so on13. Among these implicated nucleases, DFF40 is one of the well-characterized. DFF40 was first identified as one subunit of DNA fragmentation factor in the cytoplasm of Hela cells14, and subsequently proved to be the homolog of caspase-activated DNase (CAD) in mouse6. In non-apoptotic cells, DFF 40 was in complex with the inhibitory DNA fragmentation factor DFF45/ICAD. Besides its inhibitory function on CAD nuclease activity, DFF45/ICAD also acts as a CAD/DFF40-specific folding chaperone, which is essential for apoptosis15. During apoptosis, ICAD/DFF45 is cleaved by activated caspase-3, releasing CAD/DFF40 to digest chromosome DNA into oligonucleosomal fragments5, 6, 16. CAD and ICAD exist in the cells of a variety of species. In Xenopus egg extracts, we have identified the CAD and ICAD homologs and re-named them as XCAD and XICAD (previously termed as XAD and IXAD respectively) 17, 18. The XCAD/XICAD complex performs the same acti-vation mechanism as the CAD/ICAD complex does in DNA fragmentation in apoptosis in Xenopus egg extract. Furthermore, human ICAD can inhibit XCAD activity and Xenopus XICAD can inhibit human CAD17, 18, 19.
Another well-defined DNase for the late stage oligo-nucleosomal DNA digestion is endonuclease G10, 20. Endonuclease G is normally situated in mitochondria in non-apoptotic cells and relocates into the nucleus during apoptosis, where it digests the chromatin DNA into oligonucleosomal fragments21, 22.
Unlike that of the late stage apoptotic DNA fragmentation, the mechanism for the early stage DNA fragmentation largely remains to be elucidated, although some reports indicated that CAD also plays a role in large molecular weight DNA fragmentation23. Recent experiments with CAD-depleted sub-lines of chicken DT40 cells showed that the large molecular weight DNA fragmentation activity was not seriously affected by the CAD depletion, which suggested that CAD is not the responsible DNase for the early stage chromatin DNA fragmentation24.
In this study, using exogenous cytochrome c to induce apoptosis in the Xenopus egg extract containing mouse liver nuclei, we found that a two-stage DNA fragmentation also existed in this cell-free system. During the apoptotic process, exogenous chromatin DNA was firstly digested into large molecular weight DNA fragments by an unknown large molecular weight DNA fragmentation factor (LDFF). The large molecular weight DNA fragments were then digested into short oligonucleosomal DNA fragments by XCAD. We partially characterized the LDFF and showed that LDFF is Mg2+-dependent and Ca2+-independent. LDFF remains active in an acidic optimal pH condition and could tolerate up to 45°C heat treatments.
MATERIALS AND METHODS
Preparations of mouse liver nuclei and Xenopus egg extract S-150
Mouse liver nuclei and crude egg extract were prepared as described18, 25. The high speed supernatant (termed extract S-150) was prepared by centrifuging the crude egg extract at 150,000 g for 2 h (HITACHI 55P-72, RPS 50-II rotor). Prior to centrifugation, separate additions of aprotinin and leupeptin (Sigma) were added to give final concentrations of 6 μg/ml and 8 μg/ml, respectively. After being stored in liquid nitrogen, frozen extract S-150 can be induced into apoptosis by addition of 1 μM cytochrome c after being thawed rapidly17, 18.
Apoptotic chromatin DNA fragmentation assay
The final concentration of 1 μM cytochrome c (from horse heart, Sigma) was added into 50 μl extract S-150 with 106 mouse liver nuclei. The extract was incubated at 23°C for 2 h, then mixed with 450 μl buffer D (0.1 M Tris CHCl, pH 8.5, 5 mM EDTA, 0.2 M NaCl, 0.2% w/v SDS, and 0.2 mg/ml protease K) and incubated at 37°C for another 3 h or overnight. NaCl was added to a final concentration of 1.5 M, centrifuged at 10,000 g for 20 min at room temperature and the resultant pellet was discarded. The chromatin DNA in the suspension was precipitated with equal volume of 100% ethanol, and then centrifuged at 14,000 g for 5 min. After washing once with 70% ethanol, the DNA pellet was dissolved at 37°C for 2 h in 20 μl TE containing 200 μg/ml RNase A. For nucleosomal fragments detection, 10 μl DNA was loaded onto a 1.5% agarose gel and subjected to electrophoresis. For large molecular weight DNA fragments detection, 10 μl DNA was loaded onto 0.4% agarose gel and subjected to electrophoresis in 0.5×TBE buffer for 8 h at 4°C.
Prokaryotic expression and purification of human DFF40 and DFF45, and in vitro nuclease activity assay of recombinant DFF40
The coding region of CAD/DFF40 and ICAD/DFF45 were amplified from the total RNA of Hela cells by RT-PCR, then were inserted into pET28a (+) (Invitrogen) respectively. For protein expression, the E. coli BL21 (pLys) were transfected with pET28a(+)-CAD/DFF40 or pET28a(+)-ICAD/DFF45. Protein expression was induced by adding IPTG into the medium to a final concentration of 1 mM and allow the bacteria to grow for additional 4 h at 30°C. The cells were then collected for protein purification.
ICAD/DFF45 was expressed as a soluble fraction and was purified by His6×Talon-spin column (Clontech) according to the manual provided. Proteins were desalted using a G-25 column and then concentrated to 2 mg/ml by 10K cut-off centrifuges (Nanosap). Glycerol was added to a final concentration of 50% and the proteins were stored in aliquots at -20°C.
DFF40 was expressed almost exclusively in insoluble bodies. After sonication, the suspension was centrifuged at 12,000 g for 15 min. The pellets were washed twice with wash buffer (50 mM Sodium Phosphate, pH 7.0, 300 mM NaCl, 1% Triton X-100), and dissolved in denaturation buffer (10 mM HEPES-KOH, pH 7.2, 50 mM KCl, 5 mM MgCl2, 5 mM EGTA, 10 mM DTT, 20% glycerol, 6 M guanine-HCl). After incubation at 30°C for 1 h to completely denature the proteins, the solutions were diluted at 1:100 with renaturation buffer (10 mM HEPES-KOH, pH 7.2, 500 mM NaCl, 50 mM KCl, 5 mM MgCl2, 5 mM EGTA, 100 mM DTT, 20% glycerol, 50 μg/ml DFF45) and incubated for additional 2 h. A 2 μl-aliquot of the reaction was incubated with 0.1 μg caspase-3 and 106 mouse liver nuclei in a final volume of 50 μl adjusted with nuclease buffer (10 mM HEPES-KOH, pH 7.2, 50 mM KCl, 5 mM MgCl2, 5 mM EGTA, 20% glycerol, 0.1 mM PMSF, 1 mg/ml BSA) at 37°C for 4 h. The DNA was then extracted for nucleosomal and large molecular weight fragments detection.
RESULTS
Two-stage chromatin DNA degradation in the cell-free apoptotic system
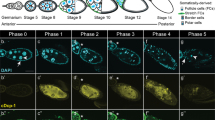
It has been previously reported that cytochrome c could induce apoptosis in Xenopus egg extract17, 18, 26. To study the chromatin DNA degradation, we established the cell-free apoptotic system using egg extract and mouse liver nuclei with the induction of cytochrome c. After adding cytochrome c and mouse liver nuclei to the extract for a while, the nuclei soon underwent apoptosis with chromatin condensation, margination, and nuclear fragmentation, which was observed by fluorescence microscopy (Fig 1B). To determine the chromatin DNA fragmentation, we extracted the DNA from the fragmented nuclei at various time points and separated the DNA on 1.5% agarose gel. It showed clearly that apoptotic nucleosomal DNA ladders appeared after 0.5 h of incubation. To see if chromatin DNA was degraded into large molecular weight fragments, we separated the same batch of DNA on 0.4% agarose gel. We found that chromatin DNA was also degraded into 50 kb fragments progressively (Fig 1C and D) and that the large molecular weight DNA fragmentation took place prior to nucleosomal fragmentation (Fig 1C and D, 0.5 h lane). These results suggest that in the egg extract, apoptosis also occurs as a two-stage DNA fragmentation process, similar to what was found in somatic cells.
Two-stage chromosome DNA degradation exists in apoptosis of egg extracts. Egg extract S-150 containing mouse liver nuclei was induced to apoptosis by adding cytochrome c to a final concentration of 1 μM. After incubation at 23°C for 2 h, the nuclei were stained with DAPI, and then observed by fluorescence microscopy. (A) Control; (B) Induction with cytochrome c. At the time indicated, the chromatin DNA was extracted and separated on 1.2% agarose for detecting the oligonucleosomal DNA (C) or 0.4% agarose for detecting the large molecular weight DNA fragments (D).
Biochemical features of the large molecular weight DNA fragmentation factor (LDFF)
Since we observed large molecular weight DNA frag-mentation, we presumed that there must be at least one factor responsible for this DNA fragmentation activity. We called this factor the large molecular weight DNA fragmentation factor (LDFF). Previously we had identified a 27 KD DNase in egg extract, termed XCAD, which was activated during apoptosis. XCAD could digest chromosome DNA into nucleosomal fragments17, 18. To identify if LDFF was XCAD, we compared the biochemical features of LDFF to those of XCAD.
First, we tested the sensitivities of LDFF and XCAD to diverse ions. We observed that EDTA, Zn2+ or Mn2+ could inhibit not only oligonucleosomal DNA fragmentation as reported18 but also the large molecular weight DNA fragmentation activity (Fig 2A and B). However, EGTA could only extensively inhibit XCAD activity for oligonucleosomal fragmentation, but not LDFF for the large molecular weight DNA fragmentation (Fig 2A and B), indicating XCAD was a Ca2+, Mg2+-dependent DNase, and LDFF was Mg2+-dependent and Ca2+-independent.
The sensitivities of XCAD and LDFF to diverse ions. Egg extract S-150 was induced to apoptosis by adding cytochrome c to a final concentration of 1 μM. After induction for 2 h, 105 mouse liver nuclei were added to apoptotic extract with a final concentration of 5 mM Zn2+, 5 mM Mn2+, 5 mM EGTA or 5 mM EDTA, respectively. After incubation at 23°C for additional 2 h, total chromatin DNA was extracted and separated on 1.5% agarose gel for detection of the oligonucleosomal fragmentation (A) and 0.4% agarose gel for the large molecular weight DNA fragmentation (B).
Secondly, the optimal pH for XCAD and LDFF activities in apoptotic DNA fragmentation were examined. We found that the optimal pH for XCAD for oligonucleosomal fragmentation were 7.4 and 8.0, while the optimal pH for LDFF for large molecular weight DNA fragmentation were 5.0, 7.4 and 8.0 (Fig 3A, B). These results indicated that LDFF, but not XCAD, could keep active under acidic condition.
The sensitivities of XCAD and LDFF to different pH. The extract S-150 was induced to apoptosis by 1 μM final concentration of cytochrome c. After induction for 2 h, the apoptotic extract were mixed with 100 mM citrate-sodium citrate buffer in different pH as indicated, and 105/ml mouse liver nuclei were added, then the mixture was incubated for 2 h. The total chromatin DNA was extracted and separated on 1.5% agarose gel for the detection of oligonucleosomal fragmentation (A) and 0.4% gel for the large molecular weight DNA fragmentation (B).
Finally, the sensibility of LDFF and XCAD to the tempe-ratures was also tested. The results showed that the optimal temperature for both XCAD and LDFF was 23°C (Fig 4A, B), a good temperature for Xenopus laevis. However, it was surprising that LDFF could tolerate up to 45°C heat treatment for 10 min (Fig 4A, B).
The sensitivities of XCAD and LDFF to diverse temperatures. The extract S-150 was induced to apoptosis by 1 μM final concentration of cytochrome c. After induction for 2 h, the apoptotic extract was treated at varies temperatures for 10 min and centrifuged at 14,000 rpm for 10 min.Then the mouse liver nuclei were added to the supernatant, and after incubated at 23°C for 2 h, the total chromatin DNA was extracted and separated on 1.5% agarose gel for the detection of oligonucleosomal fragmentation (A) and 0.4% gel for the large molecular weight DNA fragmentation (B).
From the above biochemical analysis, we concluded that LDFF was not XCAD, but a different DNase (s).
LDFF was activated by caspase-3.
Caspase-3 plays a key role in apoptosis. During apop-tosis, active caspase-3 cleaves ICAD/DFF45, releasing CAD/DFF40 to digest chromatin DNA into olignucleosomal fragments. To test if LDFF is also activated by caspase-3, we added caspase-3 inhibitor, Ac-DEVD-CHO, to the egg extract S-150 together with cytochrome c and mouse liver nuclei. We found that both the large molecular weight and the oligonucleosomal DNA fragmentation activities were inhibited (Fig 5A and B), suggesting LDFF was also activated by caspase-3.
LDFF was activated by caspase-3 during apoptosis. 1 μM final concentration of caspase-3 inhibitor, Ac-DEVD-CHO, together with 1 μM cytochrome c and 105 mouse liver nuclei were adde to the egg extract S-150. After incubation at 23 °C for 2 h, DNA was extracted and separated on 1.5% agrose gel to examine the nucleosomal fragmentation (A) and on 0.5% agarose gel to examine large DNA fragmentation (B). Extract without cytochrome c was used as the negative control.
Effects of DFF40/CAD and DFF45/ICAD in the large molecular weight DNA fragmentation
What kind of roles do DFF40/CAD and DFF45/ICAD play in large molecular weight DNA fragmentation is still under controversial. Some reports showed that DFF40/CAD is also responsible for large molecular weight DNA fragmentation23, while some other reports indicated it does not24. To confirm whether DFF40/CAD was responsible for the large molecular weight DNA fragmentation, we incubated the purified active human DFF40 with mouse liver nuclei. We found that with DFF40 the chromatin DNA could only be degraded into oligonucleosomal ladders, but not the large molecular weight DNA fragments (Fig 6A, B). This observation confirmed that CAD/DFF40 does not have the large molecular weight DNA fragmentation activity.
Effect of CAD/DFF40 and ICAD/DFF45 on the large molecular weight DNA fragmentation. (A) and (B): 1 μM renatured human CAD/DFF40 was added to 105/μl mouse liver nuclei with 100 ng/ml caspase-3 and incubated at 37°C for 4 h. Total chromatin DNA was extracted and separated on 1.5% agarose gel for oligonucleosomal (A) and 0.4% agarose gel for large molecular weight DNA fragmentation detection (B). (C, D) and (E): Purified human CAD/DFF45 was added to the egg extract S-150 with the absence of 1 μM cytochrome c and incubated at 23°C for 2 h. Total DNA was extracted and separated on 1.5% agarose gel for oligonucleosomal (C) and large molecular weight DNA fragmentation detection (D). Also, the nuclei were stained with DAPI at the indicated time, and observed by fluorescence microscopy (E).
We also tested the effect of ICAD/DFF45 on the large molecular weight DNA fragmentation. We added the purified bacteria-expressed human ICAD/DFF45 to the extract S-150 with cytochrome c and mouse liver nuclei and found that human ICAD/DFF45 can only inhibit XCAD activity for the oligonucleosomal DNA fragmentation (Fig 6C) but not LDFF activity for the large molecular weight DNA fragmentation (Fig 6D). At the same time, it was found that DFF45 could only delay, but not inhibit, the occurrence of chromatin condensation and margination (Fig 6E).
These findings suggested that DFF40/CAD and DFF45/ICAD were not the critical factors for the large molecular weight DNA fragmentation.
DISCUSSION
In this paper we reported that there exists a two-stage DNA fragmentation processe during apoptosis in the cell free extract from Xenopus eggs, which is similar to what has been reported in somatic cells1, 2, 3, 4. In the first stage the chromatin DNA is cut into large molecular weight fragment (about 50 kb) and in the second stage into oligo-nucleosomal DNA ladders. However, there is an over-lap within the two different stages (Fig 1). We also confirmed that there are at least two different kinds of nucleases responsible for the DNA fragmentations: XCAD, a homolog of CAD, and LDFF. XCAD, which was identified previously17, 18, digests the chromatin DNA into oligonucleo-somal ladders, while LDFF will cut the chromatin DNA into the large molecular weight fragments. We have partially characterized LDFF with biochemical assays although we have not identified it.
It was further proved that LDFF was clearly not XCAD, but maybe a totally different kind of DNase(s). CAD/DFF40, the human homolog of XCAD, can substitute for XCAD in digesting chromatin DNA into oligonucleosomal DNA pieces; while it could not induce large molecular weight DNA fragmentation (Fig 6B). ICAD/DFF45, the specific inhibitor of CAD, which could also inhibit XCAD specifically17, 18, 19, could not inhibit LDFF activity for the large molecular weight DNA fragmentation (Fig 6D). LDFF was sensitive to Mg2+, but not to Ca2+ (Fig 2B), whereas XCAD was both Mg2+- and Ca2+-dependent (Fig 2A). LDFF could also resist up to 45°C treatment and low pH down to 5.0 (Fig 4 and 5). These results showed LDFF and XCAD functioned differently in large molecular weight and nucleosomal DNA fragmentation.
According to the biochemical features, the best candidate for LDFF might be DNase II, an acid lysosomal nuclease. During apoptosis, DNase II relocates to nucleus27 and like LDFF, DNase II was also Ca2+-independent (Fig 3A)27. It was found that overexpression of DNase II could induce apoptosis in Chinese hamster ovary cells and this process could not be blocked by the caspase inhibitor benzyloxycarbonyl-Val-Ala-Asp-(O-methyl)-fluoro-methylketone28. These results suggest that DNase II functions downstream of caspase, which is in accordance with our result that LDFF was activated by caspase-3 (Fig 3B). In C. elegans, NUC-1, the DNase II homolog, plays an important role in complete DNA digestion during apoptosis. Mutation of NUC-1 could lead to DNA degradation slow-down29. All these suggested that DNase II might get involved in the process of apoptotic DNA fragmentation, and that DNase II is very similar to LDFF identified in our study.
In conclusion, there are at least two kinds of DNases in egg extract, XCAD and LDFF, responsible for the two-stage chromatin DNA fragmentation in apoptosis. XCAD is the homolog of CAD/DFF40 and is responsible for nucleo-somal DNA molecular weight chromatin fragmentation. But LDFF cuts chromatin DNA into large pieces. Although the activity of LDFF in digesting the chromatin DNA was partially characterized in this work, a lot remains to be done such as the cloning of LDFF gene.
References
Wyllie AH, Glucocorticoid-induced thymocyte apoptosis is associated with endogenous endonuclease activation. Nature 1980; 284(5756):555–6.
Cohen JJ, Duke RC . Glucocorticoid Activation of a Calcium-Dependent Endonuclease in Thymocyte Nuclei Leads to Cell-Death. J Immunol 1984; 132:38–42.
Filipski JJ . Leblanc T Youdale, et al. Periodicity of Dna Folding In Higher-Order Chromatin Structures. EMBO Journal 1990; 9:1319–27.
Ucker DS, Obermiller PS, Elkhart W, et al. Genome Digestion Is A Dispensable Consequence Of Physiological Cell-Death Mediated By Cytotoxic Lymphocytes-T. Mol Cell Biol 1992; 12 :3060–9.
Liu X, Li P, Widlak P, Zou H, et al. The 40-kDa subunit of DNA fragmentation factor induces DNA fragmentation and chromatin condensation during apoptosis. Proc Natl Acad Sci USA 1998; 95:8461–6.
Enari M, Sakahira H, Yokoyama H, et al. A caspase-activated DNase that degrades DNA during apoptosis, and its inhibitor ICAD. Nature 1998; 391(6662):43–50.
Halenbeck R, MacDonald H, Roulston A, et al. CPAN, a human nuclease regulated by the caspase-sensitive inhibitor DFF45. Curr Biol 1998; 8(9):537–40.
Ribeiro JM, Carson DA . Ca2+/Mg2+-dependent endonuclease from human spleen: purification, properties, and role in apoptosis. Biochemistry 1993; 32(35):9129–36.
Wu YC, Stanfield GM, Horvitz HR . NUC-1, a Caenorhabditis elegans DNase II homolog, functions in an intermediate step of DNA degradation during apoptosis. Genes Dev 2000; 14:536–48.
Parrish J, Li L, Klotz K, et al. Mitochondrial endonuclease G is important for apoptosis in C. elegans. Nature 2001; 412(6842):90–4.
Solovyan VT, Bezvenyuk ZA, Salminen A, et al. The role of topoisomerase II in the excision of DNA loop domains during apoptosis. J Biol Chem 2002; 277(24):21458–67.
Zhang J, Lee H, Agarwala A, Wen Lou D, Xu M . DNA fragmentation factor 45 mutant mice exhibit resistance to kainic acid-induced neuronal cell death. Biochem Biophys Res Commun 2001; 285:1143–9.
Zhang J, Liu X, Scherer DC, et al. Resistance to DNA fragmentation and chromatin condensation in mice lacking the DNA fragmentation factor 45. Proc Natl Acad Sci USA 1998; 95:12480–5.
Liu X, Zou H, Slaughter C, et al. DFF, a heterodimeric protein that functions downstream of caspase-3 to trigger DNA fragmentation during apoptosis. Cell 1997; 89(2):175–84.
Sakahira H, Iwamatsu SA, Nagata Specific chaperone-like activity of inhibitor of caspase-activated DNase for caspase-activated DNase. J Biol Chem 2000; 275:8091–6.
Halenbeck R, Roulston AMH, Chen TT, et al. CPAN, a human nuclease regulated by the caspase-sensitive inhibitor DFF45. Curr Biol 1998; 8:537–40.
Lu ZG, Yang W, Cao Q, et al. Identification of inhibitor of apoptosis specific DNase in egg extract. Chinese Sci Bull 2001; 46:1449–53.
Zhu S, Jiang ZF, Zhang B, et al. Identification of a DNase activated in egg extracts undergoing apoptosis. Chinese Sci Bull 1998; 43:522–6.
Yang W, Lu ZG, Zhang L, et al. Recombinant Human DFF45 Inhibits Apoptosis-specific Endonuclease in a Cell-free System of Egg Extracts. Acta Biochim Biophys Sin 2001; 33(1):82–6.
Loo van, Schotte GP, van Gurp M, et al. Endonuclease G: a mitochondrial protein released in apoptosis and involved in caspase-independent DNA degradation. Cell Death Differ 2001; 8(12):1136–42.
Li L, Luo YX, Wang X . Endonuclease G (EndoG) is an apoptotic DNase when released from mitochondria. Nature 2001; 412:95–9.
Zhan J, Xu M . Apoptotic DNA fragmentation and tissue homeostasis. Trends Cell Biol 2002; 12:84–9.
Morana SJ, Wolf CM, Li J, et al. The involvement of protein phosphatases in the activation of ICE/CED-3 protease, intracellular acidification, DNA digestion, and apoptosis. J Biol Chem 1996; 271(30):18263–71.
Samejima K, Tone S, Earnshaw WC . CAD/DFF40 nuclease is dispensable for high molecular weight DNA cleavage and stage I chromatin condensation in apoptosis. J Biol Chem 2001; 276:45427–32.
Liu X, Li P, Widlak P, et al. The 40-kDa subunit of DNA fragmentation factor induces DNA fragmentation and chromatin condensation during apoptosis. Proc Natl Acad Sci USA 1998; 95(15):8461–6.
Zhao Y, Lu Z, Wu M, et al. Effects of phosphocreatine on apo-ptosis in a cell-free system. J Biol Chem 2001; 276(37):34573–8.
Torriglia A, Perani P, Brossas JY, et al. L-DNase II, a molecule that links proteases and endonucleases in apoptosis, derives from the ubiquitous serpin leukocyte elastase inhibitor. Mol Cell Biol 1998; 18(6):3612–9.
Krieser RJ, Eastman A . The cloning and expression of human deoxyribonuclease II. A possible role in apoptosis. J Biol Chem 1998; 273(47):30909–14.
Wu YC, Stanfield GM, Horvitz HR . NUC-1, a caenorhabditis elegans DNase II homolog, functions in an intermediate step of DNA degradation during apoptosis. Genes Dev 2000; 14(5):536–48.
Acknowledgements
We gratefully thank Dr. Joe C WU from IDUN Pharmaceuticals Inc. (LaJolla, California) for purified recombinant caspase-3.
This work was supported by Major State Basic Research Program of China (No. G1999053905) and National Science Fund for Distinguished Young Scholars (No. 30225016).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
LU, Z., ZHANG, C. & ZHAI, Z. LDFF, the large molecular weight DNA fragmentation factor, is responsible for the large molecular weight DNA degradation during apoptosis in Xenopus egg extracts. Cell Res 14, 134–140 (2004). https://doi.org/10.1038/sj.cr.7290212
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290212
Keywords
This article is cited by
-
Detection of DNA ladder during cotyledon senescence in cotton
Biologia plantarum (2008)