ABSTRACT
Based on the sequence information of Arabidopsis PIN1, two cDNAs encoding PIN homologues from Brassica juncea, Bjpin2 and Bjpin3, were isolated through cDNA library screening. Bjpin2 and Bjpin3 encoded proteins containing 640 and 635 amino acid residues, respectively, which shared 97.5% identities with each other and were highly homologous to Arabidopsis PIN1, PIN2 and other putative PIN proteins. BjPIN2 and BjPIN3 had similar structures as AtPIN proteins. Northern blot analysis indicated that Bjpin2 was expressed in stem, leaf and floral tissues, while Bjpin3 was expressed predominantly in stem and hypocotyls. Two promoter fragments of pin genes, Bjpin-X and Bjpin-Z, were isolated by 'genome walking' technique using primers at 5′-end of pin cDNA. Promoter-gus fusion studies revealed the GUS activities driven by Bjpin-X were at internal side of xylem and petal; while those driven by Bjpin-Z were detected at leaf vein, epidermal cell and cortex of stem, vascular tissues and anther. Results of the pin genes with different expression patterns in B. juncea suggested the presence of a gene family.
Similar content being viewed by others
INTRODUCTION
Auxin is critical for plant embryo development, morphogenesis and especially polar differentiation and growth. Polar auxin transport (PAT), detected in stem and roots of all higher plants, was involved in plant apical dominance, tropic growth, vascular and floral tissue differentiation, plant pattern formation and lateral root development, etc (for reviews, seeNi, et al, 20001; Berleth and Sachs, 20012). Current model of PAT, chemiosmotic hypothesis (Rubery and Sheldrade3 and Raven4), posited the occurrence of PAT through the action of cellular auxin influx and efflux carriers locating on the plasma membrane of transporting cells, and the direction of PAT depends on the basal location of the efflux carriers.
The first molecular support of the model came from the characterization of a candidate auxin influx carrier, AUX1 (auxin resistant1), which was a presumptive membrane protein similar to amino acid permeases5 and specifically expressed in the root. Further isolation and characterization of two genes, Atpin1 and Atpin2, indicated that the primary function of the AtPIN1 protein was for PAT in the inflorescence axis and AtPIN2 in root tissues6, 7, 8, 9, 10. AtPIN1 involved in the auxin transport from stem to root tip and was critical for symmetry differentiation of embryo, the initiation and radial position of plant lateral organs, pattern formation of leaf vein11, 12, 13. AtPIN2 played a role in root gravitropic response together with AUX19. AtPIN1 was a membrane protein locating at the basal end of auxin transport-competent cells in vascular tissues7. AtPIN2-expressing yeast cells were resistant to toxic fluoroindoles, an IAA analogy8, and released preloaded IAA more rapidly than control cells demonstrated that AtPIN2 transported IAA out of the yeast cells and indeed a component of auxin efflux carriers9. The Arabidopsis mutant pin1 showing normal tropic responses of shoot and root10 also indicated that there maybe other pin genes controlling the lateral auxin transport pathways. Presence of other cDNAs encoding PIN homologues indicated a multi-gene family in Arabidopsis genome that, however, suggested the specific roles of pin genes in auxin related plant growth and development. Analyses of Atpin1 (widely expressed in all the tissues tested) and Atpin2 (specifically expressed in the root) indicated their expression patterns were closely linked to their physiological functions. As a conclusion, isolation of pin genes, especially studies on their expression patterns will provide informations on functional mechanisms of PAT during plant growth and development, especially polar differentiation.
We are interested in the functions of PAT in plant cells. From Brassica juncea we have isolated an auxin efflux carrier (BjPIN1) and performed functional characterization through transgenic approaches of Arabidopis14. Here we report the isolation of two putative pin-gene promoters and two cDNAs encoding PIN homologous of B. juncea, as well as the studies on their expression pattern analysis.
MATERIALS AND METHODS
Materials
Enzymes used for DNA restriction and modification were obtained from Boehringer (Mannheim, Germany). DNA primers for polymerase chain reaction (PCR) and “LAPCR in vitro Cloning” kit used for 'genome walking' were obtained from TaKaRa Biotechnology (Dalian, China). 5-bromo-4-chloro-3-indolyl b-D-glucuronic acid (X-Gluc) was obtained from Sigma (USA). Radiochemicals [α-32P]dCTP was obtained from Yahui Company (Beijing, China). The leaf lZAPII cDNA library of Brassica juncea was kindly provided by Pua E-C. (National University of Singapore).
Bacteria and plants
Escherichia coli strain XL-1 Blue (Stratagene, La Jolla, USA) was used for library screening and DNA cloning. Agrobacterium strain LBA4404 was used for plant transformation. B. juncea (Indian mustard) and Nicotiana tabacum plants were grown in the green house with a 12 h light (26°C) and 12 h dark (20°C) period.
Isolation of the Bjpin cDNAs
Bjpin cDNAs were isolated as described14. In short, two reverse degenerated primers were designed based on the nucleotide sequences of Atpin18 and two other putative pin sequences and used to amplify the Pin-homologous cDNA fragments from cDNA library of B. juncea. Specific primers were then used to isolate the full-length cDNAs through PCR-based screening14, 15.
DNA sequencing was performed by TaKaRa Biotechnology (Dalian, China). Computational analysis was performed with the help of the programs of the Wisconsin Genetics Computer Group (GCG Package, Version 10.1). The BLAST search programs were used for sequence comparisons on DNA and amino acid sequences in GeneBank, EMBL, dbEST and SwissProt databases.
Northern blot analysis
RNAs were prepared from well-watered B. juncea plants according to the protocol of Logemann et al16 and quantitiated spectrophotometrically at 260 nm. RNA (20 μg per lane) was electrophoretically separated in denaturing agarose gels and blotted onto Hybond-N+ membranes (Amersham, USA). RNA was fixed onto the membrane by heating at 80°C for 2 h. PCR products of around 400-bps of the Bjpin2 and Bjpin3 (encoded the amino acid residues at positions 305-441 and 305-436, respectively) were served as [α-32P]dCTP-labeled hybridization probe. Membranes were hybridized and washed as described17. Blots were exposed to Fuji X- films between intensifying screens for 3–4 d at −70°C.
Isolation of the promoter regions of pin genes
Promoter regions of pin genes were isolated using “LAPCA in vitro cloning” kit. 5 mg genomic DNA was digested with Hind III or EcoR I and then ligated with corresponding adaptors. Ligation products was then denatured at 94°C for 5 min and used as templates for 1st round PCR with primers C1 (5′-GTACATATTGTCGTTAGAACGCGTAATACGACTCA-3′) and BjPS1 (5′-TACAGAGCCGTAGGCGAGGATCATAG-3′). PCR was performed as follows: 94°C for 30 sec; 58°C for 90 sec; 72°C for 3 min; 30 cycles. PCR products were then diluted for 10 times and 1 ml was used as templates for 2nd PCR using primers C2 (5′-CGTAATACGACTCACTATAGGGAGA-3′) and BjPS2 (5′-CGAGGATCATAGCGACGTAGAG-3′) under same conditions as described above. PCR products were separated on the agarose gel and subcloned into TA vectors. Colonies harbouring PCR products were sequenced (TaKaRa Biotechnology , Dalian) and analyzed with programs of the Wisconsin Genetics Computer Group (GCG Package, Version 10.1) and PLACE.
Plant transformation with binary constructs harbouring promoter regions of pin genes
Promoter regions before ATG of Bjpin-X was amplified via PCR using standard primer SP6 and GW1.2A (5′-CTTATTTACGGTGGAATATAGAGGG-3′) with Pfu taq polymerase and then digested with Hind III, resulted DNA fragment was then ligated to pBSK precut with Sma I /Hind III. Resulted plasmid DNA was digested with Hind III / BamH I and DNA fragment was ligated to pBI101.3 cut with same enzymes.
Same cloning strategy was used for Bjpin-Z, primers SP6 and Bjpin1S (5′-CTTCTTTCTGGTTGATTAGAGAG-3′) were used for amplification, restriction enzyme Sal I was used for cloning procedures.
The resulted plasmids, containing the promoter-uidA coding region, were transformed into A. tumefaciens strain LBA4404 for tobacco transformation. Transformation, using leaf disc as materials for transformation, were performed as described.
Analysis of transgenic plants
Transgenic tobacco plants were confirmed via PCR using primers located in promoter region, for Bjpin-X, primers GW1.2S (5′-CTTCTGCTCTTGCATGATCTTCG-3′) and W1.2A (see above), for Bjpin-Z, primers Bjpin1S (see above) and GW07A (5′-TATTATCGCCGGATGAGTGGCA-3′) were used. PCR was performed as follows: 94°C for 5 min, 1 cycle; 94°C for 1 min, 55°C for 1 min, 72°C for 1 min, 35 cycles; with a final extension at 72°C for 10min.
GUS activities were determined using 5-bromo-4-chloro-3-indolyl b-D-glucuronic acid (X-Gluc) as substrate18.
RESULTS
Isolation of Bjpin2 and Bjpin3 cDNAs from B. juncea
Using degenerated primers, a DNA fragment with a size of 550bp was amplified and sequenced, the deduced peptide sequence showed 75% identities with isolated AtPIN1 at position 1 ∼ 145, indicating a putative PIN-like fragment of B. juncea. PCR-based screening of B. juncea cDNA library was then performed and two cDNAs, Bjpin2 and Bjpin3, were isolated. There were in-frame stop codens before the translation start code 'ATG' in both cDNAs, which indicated that full-length open reading frames were included. The predicted proteins of BjPIN2 and BjPIN3 contained 640 and 635 amino acid residues respectively, and had a molecular mass of 70 kDa, similar to that of previously reported AtPIN16 and AtPIN2/EIR1 8, 9. The calculated isoelectric points of BjPIN2 and 3 were around 8.8.
BjPIN2 and BjPIN3 shared 97.5% homologies with each other and were highly homologous to Arabidopsis PIN proteins (Tab 1), with the highest identities (93%) with AtPIN3 (accession No. AF087818).
Structural organization of the BjPIN proteins
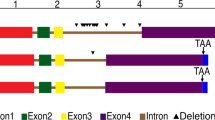
Fig 1 shows the alignment of BjPIN2 and BjPIN3 with BjPIN1, AtPIN1, AtPIN2 and other PIN-like proteins in Arabidopsis. Comparison of all the proteins revealed the presence of conserved regions locating at both N- and C-terminus, which are highly hydrophobic. Presence of around 11 predicted transmembrane regions in BjPIN2 and BjPIN3 was similar with that in AtPIN1. A non-conserved central region is predominantly hydrophilic.
Expression pattern analyses of Bjpin2 and Bjpin3
Northern blot analysis was performed to detect the transcription levels of both Bjpin2 and Bjpin3. The DNA fragments encoding the non-conserved (varied) regions of Bjpin2 and Bjpin3 were served as hybridization probes. As shown in Fig 3A and Fig 3B, single band of transcripts of approximately 2.5 kb, consistent with the length of both cDNAs, was detected. Bjpin2 mRNA was detected in stem, leaf, and floral tissues, and the Bjpin3 mRNA predominantly in stem and hypocotyls. No detectable mRNA was found in root tissues.
Expression pattern analyses of Bjpin genes Northern blot analysis of Bjpin2 and Bjpin3 transcription levels. A: Bjpin2; B: Bjpin3. Plant tissues were as follows: Ro-root; St-stem; Le-leaf; Fl-flower; Fr-fruit; Hy-hypocotyl; seed. Equal amounts of RNA (shown in bottom lines) were used for hybridization. C: Confirmation of transgenic tobacco plants harbouring Bjpin-Z::GUS construct via PCR. Transformed vector (Lane 1), genomic DNA of untransformed (Lane 2) and transformed plants (Lane 3-5) were used as templates for PCR. M: DNA ladder. GUS expression detection in transgenic tobacco plants. GUS activities driven by Bjpin-X were detected in stem (D), vascular tissues (E) and petal (F); those driven by Bjpin-Z were detected in anther (G), leaf vein (H, I), epidermis and cortex of stem (J), stem vascular tissues (K); and driven by Bjpin1 were in stem vascular tissues (L). Arrows showed the positions of GUS expressions.
Isolation of promoter regions of Bjpin-X and Bjpin-Z, which encoded other two isoforms of pin family from B. juncea
Two promoter fragments,Bjpin-X and Bjpin-Z, were isolated through genome walking technique based on the cDNA sequence of Bjpin1. After amplification using primers within adaptor and specific primers designed at 5′-coding region of Bjpin1, two DNA fragments, 1.2 kb (Bjpin-X, Fig 2A) and 0.7 kb (Bjpin-Z, Fig 2B), respectively, were amplified and sequenced. Comparison analyses showed that the coded peptides by Bjpin-X and Bjpin-Z, as well as their 5′- untranslated regions shared high homology with isolated Bjpin1, Bjpin2 and Bjpin3 but still with differences, which indicated that Bjpin-X and Bjpin-Z were promoter regions of other 2 pin genes from B. juncea than Bjpin1, Bjpin2 and Bjpin3. Sequence analyses further revealed the presence of basic elements “TAAT” and “CAAT”, and of particular interests, two pollen specific elements “AGAAA” in Bjpin-Z (Fig 2B).
Expression pattern analyses of Bjpin-X and Bjpin-Z
Both promoter regions of Bjpin-X and Bjpin-Z were subcloned into binary vector pBI101.3, and the resulted Bjpin-X::GUS and Bjpin-Z::GUS constructs were used for agrobacterium-mediated tobacco transformation. Comfirmation of resistant plants via PCR amplification indicated that both Bjpin-X::GUS andBjpin-Z::GUS had been integrated into tobacco genome (Fig 3C). At least 10 independent transgenic tobacco plants were used for GUS activity analyses. GUS activities, driven by Bjpin-X, were detected at xylem region (Fig 3D, E) and petal (Fig 3F); while those driven by Bjpin-Z were detected in mature anther and pollen (Fig 3G), leaf vein (Fig 3H, I), epidermis, cortex of stem (Fig 3J), and vascular tissues (Fig 3J, K).
DISCUSSION
Till now, at least 8 different pin-like genes have been detected in Arabidopsis genome and a few homologous genes in rice as well. Presence of multi-isoform auxin carriers, based on the extensive functions of auxin, suggest that some pin genes might share functional redundancy, yet each member of the pin gene family has its distinct functions. On the basis of distinct expression pattern and functions of AtPIN1 and AtPIN2/EIR1/AGR1 in the shoot and root, respectively6, 7, 8, 9, it is proposed that the tissue- or cell-specific auxin efflux carriers are responsible for multiple auxin transport pathways19. Presence of at least 5 pin-like genes in B. juncea, as demonstrated by our studies, indicated a pin-gene family in B. juncea. Different expression patterns of the pin-like genes in B. juncea suggest their different roles during developmental processes. Isolation and expression pattern analysis of pin genes will certainly provide hints on their roles during plant growth and development.
Bjpin2 and Bjpin3 were relatively high expressed in stem, which, however, suggests that both proteins might participate in the polar auxin transport in stem and play a specific role in stem or vascular tissue development. Recent studies have shown that AtPIN3 was expressed predominantly in endodermal cells with a lateral localization20. Since the isolated BjPIN proteins shows highest identity with AtPIN3, it is reasonable to assume that Bjpin2 and Bjpin3 might have a similar expression pattern with AtPIN3 in stem, and may play a role in growth that require lateral auxin distribution, such as shade avoidance21. Different expression patterns of Bjpin2 and Bjpin3 to isolated Bjpin114 and Arabidopsis Atpin1 and Atpin2, together with the difference between themselves, i.e. presence of Bjpin2 in leaf, floral tissues and stem while Bipin3 was relative specific in stem and hypocotyls, also indicated that they might have their distinct functions.
Different expression patterns of Bjpin-X, Bjpin-Z and isolated Bjpin114 at xylem (Fig 3E, K, L) indicated their specific roles during vascular development. Further, plant tropism growth was due to the accumulation of auxin at one side epidermis and cortex of stem (response to the stimulations) resulting in the affection of cell growth. Expression of Bjpin-X in both epidermis and cortex of stem, Bjpin-Z only at partial epidermis and cortex of stem, and isolated Bjpin1 at epidermis and cortex around stem indicated that Bjpin-Z might be involved in the unsymmetrical growth of stem by environmental stimulation such as light and gravity.
Expression of Bjpin-Z in anther and mature pollen indicated its role in reproductive growth. The relatively high level of auxin in plant anther is critical for anther development, while source of auxin, i.e. the biosynthesis site and transport in anther is still unclear. Plant floral tissues originate from leaf and auxin can be synthesized at edge of young leaf then transported to stem through vascular tissues22, 23. Recent studies showed that stigma may synthesize auxin and the auxin can be transported to floral base through polar transport resulting in the auxin gradient from stigma to receptacle and controlling of morphogenesis of pistil24. Expression of Bjpin-Z in anther may suggest the auxin synthesis capacity of pollen, even the role of Bjpin-Z during pollen germination and fertilization processes.
The differences of the expression patterns of Bjpin-Z and Bjpin1 in leaves, i.e. Bjpin-Z was restricted at vascular tissues while Bjpin1 was mostly at edge of leaves, suggested their different roles in leaf vascular development. Further expression of Bjpin1 at leaf trace indicated its role in auxin transport from leaf to stem.
**The nucleotide sequences reported in this paper have been submitted to the GenBankTM / EMBL Data Bank with accession numbers AJ249297 ( Bjpin2 cDNA), AJ249298 (Bjpin3 cDNA), AJ308229 (Bjpin-X) and AJ308230 (Bjpin-Z).
References
Ni WM, Chen XY, Xu ZH, Xue HW . Advance in study of polar auxin transport. Acta Botanica Sinica 2000; 42:221–8.
Berleth T, Sachs T . Plant morphogenesis: long-distance coordination and local patterning. Curr Opin in Plant Bio 2001; 4:457–62.
Rubery PH, Sheldrake AR . Carrier-mediated auxin transport. Planta 1974; 118:101–21.
Raven JA . Transport of indoleacetic acid in plant cell in relation to PH and electrical potential gradients, and its significance for polar IAA transport. New Phytol 1975; 74:163–72.
Bennett MJ, Marchant A, Green HG, May ST, Ward SP, Millner PA, et al. Arabidopsis AUX1 gene: a permease-like regulator of root gravitropism. Nature 1996; 273:948–50.
Gälweiler L, Guan C, Müller A, Wisman E, Mendgen K, Yephremov A, Palme K . Regulation of polar auxin transport by AtPIN1 in Arabidopsis vascular tissue. Science 1998; 282:2226–30.
Chen R, Hilson P, Sedbrook J, Rosen E, Casper T, Masson PH . The Arabidopsis thaliana AGRAVITROPIC1 gene encodes a component of the polar-auxin-transport efflux carrier. Proc Natl Acad Sci USA 1998; 95:15112–7.
Luschnig C, Gaxiola R, Grisafi P, Fink GR . EIR1, a root specific protein involved in auxin transport, is required for gravitropism in Arabidopsis thaliana. Genes Dev 1998; 12:2175–87.
Müller A, Guan C, Gälweiler L, Tanzler P, Huijser P, Marchant A, et al. AtPIN2 defines a locus of Arabidopsis for root gravitropism control. EMBO J 1998; 17:6903–11.
Okada K, Ueda H, Komaki MK, Bell CJ, Shimura Y . Requirement of the auxin polar transport in the early stage of Arabidopsis floral bud formation. Plant Cell 1991; 3:677–84.
Liu CM, Xu ZH, Chua NH . Auxin polar transport is essential for the establishment of bilateral symmetry during early plant embryogenesis. Plant Cell 1993; 5:621–30.
Reinhardt D, Mandel T, Kuhlemeier C . Auxin regulates the initiation and radial position of plant lateral organs. Plant Cell 2000; 12:507–18.
Vernoux T, Kronenberger J, Grandjean O . PIN-FORMED1 regulates cell fate at the periphery of the shoot apical meristem. Development 2000; 127:5157–65.
Ni WM, Chen XY, Xu ZH, Xue HW . Isolation and functional analysis of a Brassica juncea gene encoding a component of auxin efflus carrier. Cell Research 2002; 12(3):235–45.
Alfandari D, Darribere T . A simple PCR method for screening cDNA libraries. PCR Methods and Applications 1994; 4:46–9.
Logemann J, Schell J, Willmitzer L . Improved method for the isolation of RNA from plant tissues. Anal Biochem 1987; 163:21–6.
Xue HW, Pical C, Brearley C, Elge S, Müller -RÖber B . A plant 126-kDa phosphatidylinositol 4-kinase with a novel repeat structure: cloning and functional expression in baculovirus-infected insect cells. J Bio Chem 1999; 274:5738–45.
Jefferson RA, Kavanagh TA, Bevan MW . GUS fusions: b-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 1987; 6:3901–7.
Jones AM . Auxin transport: down and out and up again. Science 1998; 282:2201–2.
Bisseling T, Weigel D . Plant development: from cell fate to organ formation (meeting report). Plant Cell 2001; 13:222.
Morelli G, Ruberti I . Shade avoidance responses. Driving auxin along lateral routes. Plant Physiol 2000; 122:621–6.
Sieburth LE . Auxin is required for leaf vein pattern in Arabidopsis. Plant Physio 1999; 121:1179–90.
Mattsson J, Sung ZR, Berleth T . Response of plant vascular systems to auxin transport inhibition. Development 1999; 126:2979–91.
Nemhauser JL, Feldman LJ, Zambryski PC . Auxin and ETTIN in Arabidopsis gynoecium morphogenesis. Development 2000; 127:3877.
Acknowledgements
Studies were supported by the National Natural Sciences Foundation of China (No. 30070073, 95-Yu-29-7) and State Key Project of Basic Research (No. G1999011604). We greatly thank Dr. Klaus Palme for providing the Atpin1 nucleotide sequences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
NI, W., CHEN, X., XU, Z. et al. A Pin gene families encoding components of auxin efflux carriers in Brassica juncea. Cell Res 12, 247–255 (2002). https://doi.org/10.1038/sj.cr.7290131
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290131
Keywords
This article is cited by
-
Can Translation Equivalents in L1 Activated by L2 Produce Homophonic Interference: An Eye Movement Study of Cross-Language Lexical Activation in Chinese English Learners
Journal of Psycholinguistic Research (2023)
-
Application of three prediction models in pesticide poisoning
Environmental Science and Pollution Research (2022)
-
The comparison of the wisdom view in Chinese and Western cultures
Current Psychology (2022)
-
Compositionality of the Constituent Characters in Chinese Two-Character-Word Recognition by Adult Readers of High and Low Chinese Proficiency
Journal of Psycholinguistic Research (2022)
-
Cross-cultural adaptation and validation of the Groningen Frailty Indicator in Chinese nursing home residents
Aging Clinical and Experimental Research (2020)