Abstract
We test for the contribution of five strong candidate genes for obesity to quantitative variation for fatness in mice. The candidate loci are known through their major mutant phenotypes. We propose a randomization test for overall contribution of candidate genes, based on the empirical distribution of LOD scores from a quantitative trait locus (QTL) genome scan. The test is applied to data on body fat content and male gonadal fatpad weight from a QTL genome scan with an F2 population of C57BL/6J and DBA/2J inbred mice. The test is nonsignificant in this experiment for overall body fat content. QTLs detected at an experiment-wide significance level on chromosome 4, 6, 13 and 15 have effects on mean fatness of up to 19% between the homozygotes, but map to locations where there is no strong candidate gene. The test is significant for gonadal fat pad weight in males, and gives weak support for an association with the diabetes gene.
Similar content being viewed by others
Introduction
One of the goals of QTL mapping studies is to identify genes responsible for the ‘normal’ variation in quantitative phenotypes (Falconer & Mackay, 1996). Having identified regions of the genome influencing a quantitative trait or disease phenotype, it has become standard to propose one or more candidate genes that map in or near the regions a posteriori. For complex traits the number of potential candidate genes, based on the biological understanding of the traits, is very large, and QTLs identified as broad regions of the genome in a coarse-scale QTL genome scan may well contain several candidates (Kearsey & Farquhar, 1998). Furthermore, the functions of only a small fraction of genes are currently understood. Unless the confidence interval for the location of a QTL is very narrow, it is likely that an initial candidate gene proposition will be incorrect. The approach of mapping QTLs, then proposing candidate genes is different from proposing a candidate, then performing an association test. This latter approach has led to the identification of genes responsible for susceptibility to certain human diseases (e.g. Corder et al., 1993; Davies et al., 1994). In these cases very strong candidates were chosen a priori. One version of the candidate gene hypothesis is that loci known through major mutant phenotypes may have alleles with smaller quantitative effects (Mackay, 1995). In mammals, obesity is a promising trait for which to test the hypothesis. In mice, mutations at five loci (agouti (a), diabetes (db), obese (ob), tubby (tub), fat) result in grossly obese phenotypes (Chua, 1997). All five genes have been cloned, and their map positions are accurately known. These loci are strong candidates to harbour QTLs responsible for part of the normal variation for fatness. In humans the ob gene has recently been implicated in cases of severe obesity (Montague et al., 1997), but association tests of ob have proved negative (Bray et al., 1996). Here, we propose a specific candidate gene–QTL test, and apply it to fatness level in a population of 927 F2 mice derived from the C57BL/6J and DBA/2J inbreds. These strains are widely separated in the laboratory inbred mouse phylogenetic ‘tree’, and carry many molecular and phenotypic differences (Atchley & Fitch, 1993). Using tightly linked markers, we tested whether the named candidate genes show a higher than expected level of association with fatness differences, based on the empirical distribution of LOD scores from a marker-based genome scan.
Materials and methods
Mapping population and quantitative traits
The mapping population was an F2 bred at the Roslin Institute (Edinburgh), derived from inbred strains C57BL/6J and DBA/2J, which were obtained from the Jackson Laboratory in 1985. Fatness-related traits were measured in a population of 927 individuals from 104 first and second parity litters. Animals were killed at 10 weeks of age, and their body weights recorded (wt10), along with weights of gonadal fat pads in males. The trait fat pad % was 100×fat pad weight/wt10. Fat pads were replaced in the body cavity and carcasses freeze dried after removal of spleens to be used as a source of DNA for genotyping. The ratio of dry matter content to total body weight (fdw/wt10) has been shown previously to be a good predictor of fat content (Hastings & Hill, 1989). The trait predicted fat percentage (pred. fat %) was estimated as pred. fat %=100×(1.08×(fdw/wt10)−0.268), which was empirically derived from the regression of fdw/wt10 on fat content measured by ether extraction from 10 individual mice (five each from DBA/2 and C57BL/6). The correlation between predicted and measured fat was 0.99, and agrees with high correlations previously observed (Hastings & Hill, 1989).
Selection of extreme individuals for genotyping
The top 50 and bottom 50 individuals from the distributions of predicted fat % and wt10 phenotypes within each sex, after correction for litter, litter size and parity effects, were chosen for selective genotyping (Darvasi & Soller, 1992). Eighty-eight females and 85 males were selected (19% of the population).
Choice of markers and genotyping
To test for associations between the fatness phenotypes and locations of candidate genes, microsatellite markers closely linked to five obesity candidate genes were genotyped in the selected populations. Markers were chosen as follows.
Agouti On the consensus linkage map of mouse chromosome 2 the a locus maps 1.5 cM proximal to D2Mit27 (Siracusa et al., 1996). This marker is not informative for the strains used in this study, but D2Mit286 maps to the same position (Dietrich et al., 1996) and is informative.
Diabetes The nearest informative microsatellite to db is D4Mit303, which is 3 cM distal to db (Mock et al., 1996).
Obese Zhang et al. (1994) mapped ob 0.2 cM distal to Pax4. The nearest microsatellite to Pax4 is D6Mit48 which is 0.6 cM proximal (Moore & Elliot, 1996), but this marker is not informative, so D6Mit159 was selected, which maps to the same position (Dietrich et al., 1996).
Tubby D7Mit53 maps to the tub locus (Brilliant et al., 1996; Chung et al., 1996) but is not informative, so D7Mit222 was chosen, as it maps to the same position (Dietrich et al., 1996).
Fat Naggert et al. (1995) mapped the fat/Cpe locus to within 0.4 cM of D8Mit131. D8Mit31, which maps to the same position (Dietrich et al., 1996), was chosen as the nearest informative marker.
The selected populations were genotyped at an additional 75 autosomal markers, approximately 20 cM apart, and included 11 markers chosen to test for associations with body weight detected in a previous experiment (Keightley et al., 1996). Analysis of variance on the selectively genotyped data was carried out on a marker-by-marker basis, and all individuals were genotyped at markers showing an association significant at below the nominal 1% level for either predicted fat % or fat pad % or 10-week weight. There were 19 such markers.
Primers for amplification of microsatellite markers were purchased from Research Genetics Inc. (Huntsville, Alabama), and amplified in 96-well microtitre plates by standard techniques. PCR products were separated on 20 cm 6% or 9% polyacrylamide gels, stained with ethidium bromide, and photographed under UV light. Photographs were scored on two occasions and ambiguous genotypes reamplified.
Linkage map
A linkage map was constructed from genotypes of selected F2 individuals with the software package CRIMAP (P. Green, pers. comm.), assuming complete crossover interference between pairs of linked markers. An initial map order from Dietrich et al. (1996) was assumed, and map distances calculated. To test the initial map order, the position of each pair of linked markers was swapped, and the likelihood of the new and original configuration compared. The initial map order give the higher likelihood in all cases except one, where the likelihood for the initial map order was slightly lower. Map distances were also in reasonable agreement, but it should be noted that linkage maps produced from selectively genotyped populations are biased near QTLs (Lin & Ritland, 1996). The average spacing between adjacent pairs of markers was 21 cM.
Interval mapping by maximum likelihood
In order to obtain unbiased estimates of QTL effects and likelihood ratios where selective genotyping has been carried out, it is necessary to include the missing data in the analysis. Under maximum likelihood (ML) it is straightforward to include missing data, so this method was chosen. Its principal drawback, however, is a high computational requirement. A preliminary analysis to compare likelihoods of models with and without litter effects (without QTL effects fitted) was carried out, and showed that both the fatness traits had highly significant effects common to full-sib litters (P<0.001), so a method modified from Knott & Haley (1992) was implemented in which litter was integrated out as a normally distributed effect. Fixed effects of sex, parity and a linear covariate for litter size were also fitted. The genetic model assumed a QTL, Q, with effects g0=+a, g1=d or g2=−a for the homozygous DBA/2 genotype (DD), heterozygous genotype (BD) and homozygous C57BL/6 genotype (BB), respectively. The QTL was situated between two markers M1 and M2 with a recombination fraction c1 between M1 and Q, and c2 between Q and M2. Under complete interference, as assumed here, the recombination fraction for the interval is c=c1+c2. The marker state of M1 and M2 could be DD, BD, BB or unknown. The likelihood equation was

where NL is the number of litters, ni is the number of individuals within litter i, bi is the random effect of the ith litter assumed to be normally distributed with mean zero and variance σ2b, yij is the observed trait value of individual j from litter i, p(gq|m1ij, m2ij, c1, c2) is the probability of QTL genotype gq given marker locus states m1ij and m2ij for individual ij, μ is the population mean, xij is the design matrix for fixed effects and the covariate for individual ij, f is the vector of fixed and covariate effect estimates, and σ2w is the residual variance.
Following Knott & Haley (1992), numerical integration with Gaussian quadratures was employed for integration over the distribution of litter effects. A 16-point Legendre polynomial (Press et al., 1992)was employed in the integration procedure. The results from numerical integration with this method were compared with the generalized Romberg numerical integration procedure QROMO (Press et al., 1992), which has an arbitrary convergence criterion, but is much slower. The two procedures gave results that agreed closely with one another. Likelihood maximization was performed with the simplex algorithm (Press et al., 1992). The QTL mapping procedure was tested extensively with simulated data, including cases with selective genotyping. Mean parameter estimates were found to be in very good agreement with the simulated parameter values. The method could be extended to include multiple linked or unlinked QTLs, but would be very demanding computationally.
Determination of significance levels
Likelihood ratios for models with a QTL vs. no QTL were calculated at positions within each marker bracket, and at the marker positions themselves. Because of a high computational requirement, the analysis was performed with 10 cM steps within each interval. Experiment-wide significance thresholds were determined empirically using the permutation procedure of Churchill & Doerge (1994). For each trait, 1000 interval mapping analyses over the whole genome at the marker positions and at 10 cM steps were performed with the complete genotype data permuted among individuals. The highest LOD score at any point in the genome was recorded. These highest LOD scores were ranked. The experiment-wide 5% significance threshold was the 50th highest LOD score. A ‘suggestive’ level, corresponding to a 50% chance of a false positive per genome scan, was taken as the 500th highest LOD scores. To obtain peak LOD scores where experiment-wide or suggestive significance level QTLs were detected, and to estimate QTL positions and effects, more detailed runs using the unpermuted data with a 2 cM step size were carried out.
Results
Means and variances for fatness traits in the F2
Phenotypic means and within-family phenotypic variances for pred. fat % and fat pad % in the F2 population are shown in Table 1. No measures of these traits were available for the inbred grandparents or the F1, as the traits could not be measured on live animals. The F2 population ranges in body fat content from 5% to 26% at 10 weeks of age. The phenotypic correlation between predicted fat % and fat pad % is 91%.
QTL genome scan for fatness traits
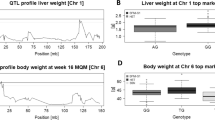
We selectively genotyped the extreme 19% of the F2 population based on phenotype at 80 microsatellite markers approximately evenly distributed in the genome, including markers tightly linked to the five obesity candidate genes (Table 2). LOD score plots from the ML interval mapping analysis with empirically derived experiment-wide and ‘suggestive’ significance levels are shown in Fig. 1 for predicted fat % and gonadal fat pad % in males. QTLs significant at the experiment-wide level are detected on chromosomes 4, 6, 13 and 15 for predicted fat %. No experiment-wide significant QTLs for gonadal fat pad % are detected, but QTLs significant at the suggestive level are on chromosomes 4, 6 and 7. At positions where selective genotyping gave the suggestion of a QTL, we augmented the data by genotyping all the individuals in the F2 population. This resulted in substantially higher LOD scores (Fig. 1), indicating that the QTL effects are repeatable. The experiment-wide significant QTLs for predicted fat explain 20% of the phenotypic variance. QTL effects detected are of the order of one-half to two-thirds of a phenotypic standard deviation between the homozygotes, and in some cases produced changes in mean fat level of the order of 20% between the homozygotes (Table 3).
LOD score plots for (a) predicted fat % and (b) gonadal fat pad % in mice obtained by interval mapping for the 19 autosomes. The plot with a broken line shows LOD score for analysis of the data from selectively genotyped individuals only. The solid line is the LOD score plot from analysis of all the available data. Horizontal solid line: experiment-wide 5% significance threshold; horizontal broken line: suggestive significance threshold (both from analysis of all available data). The experiment-wide LOD score thresholds are 4.4 and 5.3 for predicted fat % and fat pad %, respectively, and the suggestive thresholds are 2.5 and 3.1, respectively. Diamonds mark estimated positions of the five obesity candidate genes.
Candidate gene–QTL test applied to fatness
Chromosomes 2, 4, 6 and 7, which contain candidate genes (Table 2), harbour QTLs for fatness detected at least at the suggestive level (Table 3). A randomization procedure was employed to test whether the QTLs detected for the fatness traits are in significant overall association with the candidate genes. The selectively genotyped data for the fatness traits were analysed by ML interval mapping. The LOD scores at the positions of the five candidate gene markers were added together, giving an overall observed LOD score. One million sets of five random positions in the mapped region of the genome were sampled and LOD score totals for each set were calculated. The distribution of these sums is the distribution of expected LOD scores under the null hypothesis that candidate genes have an equal probability of occurring at any position in the mouse genome. The resampling will occasionally sample LOD scores at linked positions, but this is justified because the model assumes that the five candidate genes could occur anywhere, including linked positions. If the observed LOD score for the candidate genes is in the top 5% of the distribution of expected scores, then there is a significant overall association.
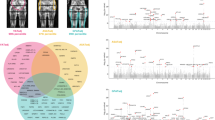
The distribution of sums of LOD scores from the 106 sets of five randomly chosen positions in the mapped region of the genome for the predicted fat % genome scan is shown in Fig. 2a. The overall LOD score at the candidate gene markers is not higher than expected on the basis of the distribution of LOD from the randomly chosen positions (P=0.38). There is therefore no statistical support for the contribution of these five candidate genes to genetic variation in predicted fat %. For gonadal fat pad %, the candidate gene–QTL randomization test showed that the sum of LOD scores is significantly higher than expected (P=0.04, Fig. 2b.
Distribution of sums of LOD scores from five randomly chosen positions in the mapped region of the mouse genome from interval mapping with selectively genotyped data of (a) predicted fat % and (b) fat pad %. Arrows indicate the observed LOD score totals at the positions of the markers closely linked to the five fatness candidate genes.
The overall significant result was largely caused by the db gene on chromosome 4, with a peak LOD score of 1.77, which is the highest LOD score at any point in the genome for the selectively genotyped data.
Discussion
The major motivation for this study was to test whether part of the ‘normal’ quantitative variation for fatness level is associated with variation in candidate genes. We perform a randomization test to compare the LOD scores from a QTL interval mapping analysis at markers linked to a set of strong candidate genes with LOD scores elsewhere in the mapped region of the genome. For the test, it would be preferable to use markers within the candidate genes themselves, but it seems reasonable to use the information from the mouse genome mapping project, and to use tightly linked markers. Linkages are sufficiently tight in the F2 that effects of candidate genes themselves will be underestimated by a few per cent, at most. The test was negative for overall body fat content. QTLs with moderate effects on overall body fat content were detected at the experiment-wide significance level on chromosomes 4, 6, 13 and 15, and a QTL significant at the suggestive level was detected on chromosome 5. In crosses involving C57BL/6J, QTLs for fatness have been previously detected on chromosomes 6 (Warden et al., 1995) and 15 (Warden et al., 1995; York et al., 1996), so it is possible that these are the same loci as detected here. Chromosomes 13 and 15 harbour the largest QTL effects detected, but do not contain the selected candidate genes. The a, db, tub and fat genes are at locations where there is no statistical support at the experiment-wide level for QTLs. Close to the ob gene, a LOD score peak is above the experiment-wide significance threshold, but not at the marker closest to ob. The result of the test applied to the fat pad % data was positive. However, no position in the genome had a LOD score above the suggestive signficance level. Futhermore, the candidate gene–QTL test was negative for predicted fat %, a highly correlated trait measured on both males and females. Subsequent typing of all the individuals in the F2 population showed that the map locations of the db and the ob markers have LOD scores above the suggestive significance level. Thus, the data do not rule out the possibility of the involvement of these candidate genes for fatness, but the evidence seems rather weak. Strong evidence that the candidate genes are indeed involved could come from introgression experiments with small regions of the genome including these genes.
The analysis here has involved the fitting of one QTL at a time. Fitting multiple linked or unlinked QTLs may increase the power of detection of QTLs and the precision of the estimation of their map locations. Thus, if there are several QTLs on a chromosome, including a contribution from a candidate gene, the peak LOD score may not occur at the position of the candidate gene. However, LOD scores are expected to be highest, on average, at the positions of marker loci where information is greatest, and this further tempers the weak evidence for the involvement of the candidate genes. An alternative way to carry out the test would be to ignore the positions of candidate genes for QTL mapping purposes, and subsequently infer the LOD scores associated with each candidate gene from the LOD score plot based on map positions relative to the markers. A test of this type has been recently carried out using data from a Drosophila bristle number selection experiment, with a negative result (Nuzhdin et al., 1998). By carrying out the test in this way, there is bias against obtaining a positive result. If markers were extremely dense, the result of the test would not depend on whether or not markers were deliberately chosen to be tightly linked to candidate loci, as long as their map positions relative to the markers were accurately known.
The candidates chosen here are ‘strong’ because they are known from classical genetics to be capable of having large effects on the trait. As our understanding of the basis of the trait improves, the set of mapped candidate genes could include a much larger set of ‘physiological’ candidates. However, a randomization test of the type described would have very little power if a larger set of weaker candidates were used, because LOD score (or equivalent test statistic) is usually too high over much of the genome in an F2 experiment. To increase the power, designs leading to narrower confidence intervals for QTLs, perhaps restricted to target regions of the genome could be carried out (Darvasi, 1998). An alternative way to increase power would be to combine results from several genome-wide surveys for the same or similar traits, but for fatness in mice these have been restricted so far to individual chromosomes (Warden et al., 1995; Taylor & Phillips, 1996, 1997; York et al., 1996).
References
Atchley, W. R. and Fitch, W. (1993). Genetic affinities of inbred mouse strains of uncertain origin. Mol Biol Evol, 10: 1150–1169.
Bray, M. S., Boerwinkle, E. and Hanis, C. L. (1996). Ob gene not linked to human obesity in Mexican–American affected sib pairs from Starr County, Texas. Hum Genet, 98: 590–595.
Brilliant, M. H., Williams, R. W., Holdener, B. C. and Angel, J. M. (1996). Mouse chromosome 7. Mamm Genome, 6: s135–s150.
Chua, S. C., Jr (1997). Monogenic models of obesity. Behav Genet, 27: 277–284.
Chung, W. K., Goldbergberman, J., Powerkehoe, L. and Leibel, R. L. (1996). Molecular mapping of the Tubby (Tub) mutation on mouse chromosome 7. Genomics, 32: 210–217.
Churchill, G. A. and Doerge, R. W. (1994). Empirical threshold values for quantitative trait mapping. Genetics, 138: 963–971.
Corder, E. H., Saunders, A. M., Strittmatter, W. J., Schmechel, D. E., Gaskell, P. C., Small, G. W. et al (1993). Gene dose of apolipoprotein E type 4 allele and the risk of Altzheimer’s disease in late onset families. Science, 261: 921–923.
Darvasi, A. (1998). Experimental strategies for the genetic dissection of complex traits in animal models. Nature Genet, 18: 19–24.
Darvasi, A. and Soller, M. (1992). Selective genotyping for determination of linkage between a marker locus and a quantitative trait locus. Theor Appl Genet, 85: 353–359.
Davies, J. L., Kawaguchi, Y., Bennett, S. T., Copeman, J. B., Cordell, H. J., Pritchard, L. E. et al (1994). A genome-wide search for human type 1 diabetes susceptibility genes. Nature, 371: 130–136.
Dietrich, W. F., Miller, J., Steen, R., Merchant, M. A., Damronboles, D., Husain, Z. et al (1996). A comprehensive genetic map of the mouse genome. Nature, 380: 149–152.
Falconer, D. S. and Mackay, T. F. C. (1996). Introduction to Quantitative Genetics, 4th edn. Longman, London.
Hastings, I. M. and Hill, W. G. (1989). A note on the effect of different selection criteria on carcass composition of mice. Anim Prod, 48: 229–233.
Kearsey, M. J. and Farquhar, A. G. L. (1998). QTL analysis in plants; where are we now? Heredity, 80: 137–142.
Keightley, P. D., Hardge, T., May, L. and Bulfield, G. (1996). A genetic map of quantitative trait loci for body weight in the mouse. Genetics, 142: 227–235.
Knott, S. A. and Haley, C. S. (1992). Maximum likelihood mapping of quantitative trait loci using full-sib families. Genetics, 132: 1211–1222.
Lin, J. Z. and Ritland, K. (1996). The effects of selective genotyping on estimates of proportion of recombination between linked quantitative trait loci. Theor Appl Genet, 93: 1261–1266.
Mackay, T. F. C. (1995). The genetic basis of quantitative variation: numbers of sensory bristles of Drosophila melanogaster as a model system. Trends Genet, 11: 464–470.
Mock, B. A., Stoyle, J., Spence, J., Jackson, I., Eppig, J. T., Fiedorek, F. T. and Newmann, P. E. (1996). Mouse chromosome 4. Mamm Genome, 6: s73–s96.
Montague, C. T., Farooqi, I. S., Whitehead, J. P., Soos, M. A., Rau, H., Wareham, N. J. et al (1997). Congenital leptin deficiency is associated with severe early-onset obesity in humans. Nature, 387: 903–908.
Moore, K. J. and Elliot, R. W. (1996). Mouse chromosome 6. Mamm Genome, 6: s113–s134.
Naggert, J. K., Fricker, L. D., Varlamov, O., Nishina, P. M., Rouille, Y., Steiner, D. F. et al (1995). Hyperproinsulinaemia in obese fat/fat mice associated with a carboxypeptidase E mutation which reduces enzyme activity. Nature Genet, 10: 135–142.
Nuzhdin, S. V., Keightley, P. D., Pasyukova, E. G. and Morozova, E. A. (1998). Mapping quantitative trait loci affecting Drosophila melanogaster sternopleural bristle number using changes of marker allele frequencies in divergently selected lines. Genet Res. in press.
Press, W. H., Teukolsky, S. A., Vetterling, W. T. and Flannery, B. P. (1992). Numerical Recipes in C, 2nd edn. Cambridge University Press, Cambridge.
Siracusa, L. D., Morgan, J. L., Fisher, J. K., Abbott, C. M. and Peters, J. (1996). Mouse chromosome 2. Mamm Genome, 6: s51–s63.
Taylor, B. A. and Phillips, S. J. (1996). Detection of obesity QTLs on mouse chromosome 1 and chromosome 7 by selective DNA pooling. Genomics, 34: 389–398.
Taylor, B. A. and Phillips, S. J. (1997). Obesity QTLs on mouse chromosomes 2 and 17. Genomics, 43: 249–257.
Warden, C. H., Fisler, J. S., Shoemaker, S. M., Wen, P. Z., Svenson, K. L., Pace, M. J. and Lusis, A. J. (1995). Identification of 4 chromosomal loci determining obesity in a multifactorial mouse model. J Clin Invest, 95: 1545–1552.
York, B., Lei, K. and West, D. B. (1996). Sensitivity to dietary obesity linked to a locus on chromosome-15 in a CAST/Ei×C57BL/6J F2intercross. Mamm Genome, 7: 677–681.
Zhang, Y. Y., Proenca, R., Maffei, M., Barone, M., Leopold, L. and Friedman, J. M. (1994). Positional cloning of the mouse obese gene and its human homologue. Nature, 372: 425–432.
Acknowledgements
We thank Sara Knott and Chris Haley for helpful advice on the analysis, Margaret Murray for techical assistance, Grahame Bulfield, Lutz Bünger, Brad Frecking, Bill Hill, Simon Horvat, Trudy Mackay, and two anonymous referees for constructive comments, and the BBSRC and Royal Society for support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Keightley, P., Morris, K., Ishikawa, A. et al. Test of candidate gene–quantitative trait locus association applied to fatness in mice. Heredity 81, 630–637 (1998). https://doi.org/10.1046/j.1365-2540.1998.00450.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.1998.00450.x
Keywords
This article is cited by
-
Subcongenic analysis of tabw2obesity QTL on mouse chromosome 6
BMC Genetics (2012)
-
Multi-environment mapping and meta-analysis of 100-seed weight in soybean
Molecular Biology Reports (2012)
-
Congenic and bioinformatics analyses resolved a major-effect Fob3b QTL on mouse Chr 15 into two closely linked loci
Mammalian Genome (2010)
-
Quantitative Trait Loci for Regional Adiposity in Mouse Lines Divergently Selected for Food Intake
Obesity (2007)
-
A meta-analysis of quantitative trait loci associated with body weight and adiposity in mice
International Journal of Obesity (2007)