Abstract
Wolbachia are intracellular maternally inherited alpha-Proteobacteria infecting a wide range of arthropods. In the common pill bug Armadillidium vulgare, the known Wolbachia strain is responsible for feminization of genetic males. We have investigated Wolbachia diversity in 20 populations of A. vulgare from west and east Europe, north Africa and north America. A new Wolbachia strain (wVulM) was identified through the variability of the wsp gene, distantly related to that previously known (wVulC) in this host species. No individual with multiple infections was detected. Inoculation experiments indicated that the new wVulM bacterial strain also induces feminization in A. vulgare. However, the wVulC strain showed a higher transmission rate than the wVulM strain and was the most geographically widespread Wolbachia in A. vulgare populations. Mitochondrial 16SrDNA gene sequencing was conducted in Wolbachia-infected individuals, revealing the occurrence of four host lineages. The comparison of bacterial strains and their respective host mitochondrial phylogenies failed to show concordance, indicating horizontal transmission of the Wolbachia strains within populations of A. vulgare.
Similar content being viewed by others
Introduction
Wolbachia are maternally inherited endosymbiotic alpha-Proteobacteria (O'Neill et al, 1992) responsible for many effects on their hosts reproduction such as cytoplasmic incompatibility, thelytokous parthenogenesis, feminization of genetic males and male killing (reviewed in Stouthamer et al, 1999). Their abilities to interact with host biology have allowed Wolbachia to invade a wide spectrum of nematodes and arthropods (reviewed in Stouthamer et al, 1999). In isopod crustaceans, Wolbachia are responsible for cytoplasmic incompatibility in two species (Legrand and Juchault, 1986; Moret et al, 2001) and feminization in many species (Bouchon et al, 1998), including Armadillidium vulgare (Rigaud et al, 1991). In case of feminization, genetic males are converted into functional females, which typically produce, in turn, female-biased progenies (Rigaud et al, 1997).
PCR-based methods have allowed extensive phylogenetic studies of Wolbachia among host species during the last decade (eg O'Neill et al, 1992; Rousset et al, 1992a; Werren et al, 1995b; Bouchon et al, 1998; Zhou et al, 1998; Cordaux et al, 2001). On the other hand, less information is available about Wolbachia at a within-host species scale. No Wolbachia variability was detected in symbionts of 11 populations of Tribolium confusum (Fialho and Stevens, 1997) or in symbionts of Trichogramma kaykai from California (Schiltuizen et al, 1998) using ftsZ gene sequences, suggesting single Wolbachia infection in these species. However, the presence of several Wolbachia strains has been reported in Drosophila simulans using bacterial 16SrDNA gene sequences (Rousset et al, 1992b). But, using the same marker, Bouchon et al (1998) could not detect any Wolbachia variability at the within-species level in isopods. The cloning of the wsp gene (Braig et al, 1998) opened a new era in the phylogenetic study of Wolbachia symbionts. This major outer surface protein-encoding gene has been shown to be about 10 times more variable than the bacterial 16SrDNA gene (Zhou et al, 1998), providing a better phylogenetic assay to investigate recent horizontal transfers. Using this marker, two male-killing Wolbachia strains were identified in the same host: the ladybird Adalia bipunctata (Hurst et al, 1999) and the butterfly Acrea encedon (Jiggins et al, 2001).
The evolutionary dynamics of the host–symbiont associations can be analysed by comparing the host mitochondrial DNA (mtDNA) with the symbiont genealogy. mtDNA is in linkage disequilibrium with the intracellular bacteria, because both are maternally inherited. The association between mitochondria and intracellular symbionts that manipulate host reproduction has already been documented in insects. In D. simulans, in which Wolbachia cause cytoplasmic incompatibility, mtDNA variants were found to hitchhike with Wolbachia strains, leading to congruence between mtDNA and Wolbachia phylogenies (Turelli et al, 1992; James and Ballard, 2000). A similar relationship was observed in the mosquito Culex pipiens (Guillemaud et al, 1997), the Asian rice gall midge Orseolia oryzae (Bethura et al, 2001) and fire ants of the genus Solenopsis (Shoemaker et al, 2001). In the two-spot ladybird beetle, in which Wolbachia induce male-killing, Schulenburg et al (2002) showed that mitochondrial haplotypes are correlated with the infection status, confirming linkage disequilibrium between mitochondria and bacteria.
In Crustacea, the association between feminizing Wolbachia and mtDNA has been investigated in two isopod species. Michel-Salzat et al (2001) detected three different Wolbachia strains within the species complex of the isopod Porcellionides pruinosus, but no congruence between Wolbachia gene (wsp gene) and mtDNA (mitochondrial 16SrDNA gene) gene genealogies. In A. vulgare, Rigaud et al (1999) investigated mtDNA variation through RFLP according to the presence of two sex ratio distorters, a Wolbachia strain and a non-mendelian feminizing factor (f element). The symbiont was associated with two closely related mitochondrial types, whereas no link was found between the f element and any mitochondrial lineage. However, the history of infection in this host species needs further investigations, since Rigaud et al (1999) sampled populations of A. vulgare on a small geographical scale (limited to France) and identified only one Wolbachia strain. The aims of the present study were: (i) to evaluate the diversity of Wolbachia strains infecting populations of the isopod A. vulgare on a large geographical scale, and (ii) to compare it to host cytoplasmic genealogy, in order to reconstruct the evolution of Wolbachia/A. vulgare relationships. The wsp gene was used to detect Wolbachia strains and the mitochondrial 16SrDNA gene was chosen as host genetic marker. The latter appears to be well suited for the present investigation because it has previously been shown to be variable enough for population genetics analyses (Michel-Salzat and Bouchon, 2000; Michel-Salzat et al, 2001).
Materials and methods
Sampling
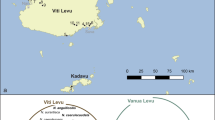
In all, 20 populations of the isopod A. vulgare were sampled (Table 1): 15 from Europe (France, Germany, Greece, Hungary, Spain and The Netherlands), one from north Africa (Tunisia) and four from north America (USA). Two different populations were sampled in Pittsford (USA) and Barcelona (Spain). The infection status of each population was assessed by testing several individuals for the presence of Wolbachia, using a PCR assay (see below).
Molecular data and phylogenetic analyses
Infected individuals randomly chosen from all Wolbachia-infected populations were sequenced for both theWolbachia wsp gene and the mitochondrial 16SrDNA gene. In addition, uninfected individuals randomly chosen from uninfected populations were analysed for their mitochondrial sequences in order to compare infected vs uninfected lineages. DNA extraction, PCR amplification and sequencing of the wsp region of the Wolbachia DNA and of the 3′ end of the 16SrDNA region of the host mtDNA were performed as described in Michel-Salzat et al (2001). Wsp and mtDNA sequences were deposited in EMBL, Genbank and DDBJ databases under accession numbers AJ419984-AJ419987 and AJ419988-AJ419997, respectively.
Wsp sequences were manually aligned taking into account the coding structure of the gene with 11 previously determined wsp sequences from isopod Wolbachia (Cordaux et al, 2001), using the Culex pipiens symbionts (Zhou et al, 1998) as an outgroup. mtDNA sequences from A. vulgare were manually aligned with the sequence of A. nasatum, previously shown to be an adequate outgroup (Michel-Salzat and Bouchon, 2000). Regions including gaps and hypervariable portions that could not be aligned with confidence were removed from the analyses. Neighbour-joining analyses were conducted using MEGA 2.1 (Kumar et al, 2001) based on Kimura two-parameter distances. Bootstrap analyses were carried out with 1000 replicates. For wsp sequences, ratios of nonsynonymous to synonymous substitutions and tests for neutrality were computed using the Nei-Gojobori method implemented in MEGA 2.1 (Kumar et al, 2001). Standard errors (SE) were estimated via a bootstrap method (1000 replicates). Deviations from the expected standard variation under the neutral hypothesis were tested by a Z-test. Pairwise uncorrected distances between infected populations were computed for Wolbachia wsp sequences and host mtDNA sequences. These distances were then used to perform a Mantel test as implemented in GENETIX 4.01 (Belkhir, 2000), in order to test the correlation between host and Wolbachia genetic distances (10 000 permutations).
Effects of Wolbachia on host phenotypes
To investigate the effects of Wolbachia from the infected population of Mery-sur-Cher, bacteria were inoculated into 36 undifferentiated juveniles (20 days of postembryonic development after releasing from the marsupium; Suzuki, 1999) from an asymbiotic lab strain of A. vulgare. Experiments were conducted as described in Bouchon et al (1998). Recipient specimens were reared and their sexual phenotype was checked every 2 weeks until their secondary sexual characters could be identified (120 days on average; Suzuki, 1999). Sexual phenotypes were defined as in Bouchon et al (1998). At the end of the experiment, the presence of Wolbachia in the surviving individuals was tested using a PCR assay.
In addition, infected females originating from four populations (Grebbeberg, Angouleme, Mery-sur-Cher and Barcelona-2) were reared in the lab at 20°C under a natural photoperiod. These females were tested positive for Wolbachia infection after they laid their clutches. The proportion of males and females of their progenies were recorded. Biases in the masculinity rate were tested according to the rate of 0.5, expected from the genetic sex determination known in A. vulgare (Juchault and Legrand, 1972). The statistical significance of differences in sex ratios was assessed using Likelihood-ratio (L-R) χ2 tests as implemented in JMP version 3.2.2 (SAS Institute Inc., Cary, NC, USA).
Results
Wolbachia diversity
The wsp sequences were obtained from the Wolbachia of infected individuals from each of the 16 populations found to harbour symbionts (Table 1). Each infected individual only harboured a single Wolbachia strain, as shown by the direct sequencing of the purified PCR products, which revealed no ambiguity or discrepancy within and between the analysed strands of each wsp sequence. To test for recombination events that may occur in the wsp gene (Werren and Bartos, 2001), dot-plot analysis of the wsp sequences was performed. No recombination signal was detected. In total, four different sequences were identified, which can be grouped into two Wolbachia strains.
The first strain was found in 12 populations from west Europe, north America and north Africa (Table 1), which corresponds to 75% of Wolbachia-infected populations investigated. Within this strain, Wolbachia from 11 populations (ie Celles-sur-Belle, Amou, Angoulême, Grebbeberg, Pittsford-1, Pittsford-2, Ribadeo, Rivesaltes, Rochester, Slater-Creek, Tunis) exhibited identical wsp sequences. This strain was named wVulC (for Wolbachia A. vulgare type Celles-sur-Belle) according to the nomenclature of Zhou et al (1998). The wsp sequence from Saint-Cyr differed from the wsp sequence from Celles-sur-Belle by only one substitution (based on the comparison of 493 positions of the wsp gene), and was therefore assigned to wVulC.
The second Wolbachia strain was found in four populations from Europe (Table 1), which corresponds to only 25% of Wolbachia-infected populations investigated. Within this strain, Wolbachia from three populations (ie Mery-sur-Cher, Athens, Budapest) exhibited identical wsp sequences. This strain was named wVulM (for Wolbachia A. vulgare type Mery-sur-Cher). The wsp sequence from Barcelona-2 differed from the wsp sequence from Mery-sur-Cher by only one substitution at a silent site and was therefore assigned to wVulM.
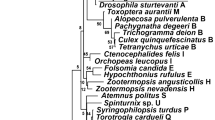
The pairwise distance between the two Wolbachia strains in A. vulgare was 5.3%. The estimated number of nonsynonymous substitutions (dN=mean number of differences±SE) between the two sequences was 9.5±2.9, whereas the estimated number of synonymous substitutions (dS=mean number of differences±SE) was 12.5±3.1. The phylogenetic relationships of wVulC and wVulM with other isopod Wolbachia strains were investigated (Figure 1). This analysis revealed that wVulC and wVulM belong to two different subclades (Vul and Ase, respectively) within the isopod-specific Wolbachia cluster of the B-supergroup of Wolbachia (Cordaux et al, 2001). These two subclades are separated by a high bootstrap score of 95% (Figure 1). Overall pairwise comparisons of the wsp sequences showed a significant deviation from the neutrality hypothesis (Z=3.69, P<0.01). The same analysis of the sequences clustering in the Vul subclade provided indication of evolution under selection (Z=2.19, P=0.03), whereas there was a neutral evolution in the Ase subclade (Z=0.67, P=0.51). Within the four, Armadillidium Wolbachia tests indicated a strong selection (Z=2.80, P<0.01) as well as between the wVulM and wVulC sequences (Z=2.82, P<0.01). Therefore, wVulC and wVulM are distantly related within the Wolbachia clade-infecting isopods.
Phylogenetic tree of isopod Wolbachia strains. The tree is based on the comparison of 493 bases of the wsp gene, using the neighbor-joining method and rooted with C. pipiens symbionts (AF020061). Numbers at nodes are bootstrap scores above 50%. Strains are identified by the host from which they were isolated. The two Wolbachia strains from A. vulgare (wVulC and wVulM) are shown in bold. Accession numbers of wsp sequences from other isopod symbionts are AJ276599–AJ276601, AJ276605–AJ276612.
A logistic regression testing the effects of both the geographical origin of populations and the symbiont strains on the prevalence of Wolbachia in females (Table 1) revealed only a significant effect of their interaction (whole model: χ2=53.5, d.f.=15, P<0.001; effect of the Wolbachia strain: L-R χ2=0, d.f.=1, P=1; effect of the geographical origin: L-R χ2=5.9, d.f.=11, P=0.88; interaction: L-R χ2=17.4, d.f.=3, P<0.001). The proportion of infected females was similar (56% in average), whatever the geographical origin when infected by the wVulC Wolbachia strain (L-R χ2=17.5, d.f.=11, P=0.09), whereas there were differences in the infection pattern in females infected by the wVulM Wolbachia strain (L-R χ2=32.9, d.f.=3, P<0.001). However, the sample size for wVulC is relatively modest compared to wVulM, making it hard to resolve the issue of this heterogeneity in this infection.
Wolbachia-induced phenotype on hosts
The feminizing effect of the wVulC Wolbachia strain in A. vulgare has already been demonstrated (Rigaud et al, 1991). To investigate whether wVulM also induces feminization in A. vulgare or not, it was artificially inoculated into 36 asymbiotic undifferentiated juveniles as described in Bouchon et al (1998). Secondary sexual characters could be seen after 128 days of rearing: three males, 14 females and five intersexes were identified among the 22 surviving individuals. Time for secondary sexual differentiation and survival rate were similar to those recorded by Suzuki (1999). The intersex phenotype was defined as sterile males with genital female apertures and hypertrophied androgenic glands (Bouchon et al, 1998). The results were identical after 230 days and the PCR assay showed that all intersexes carried Wolbachia, whereas no male did. The intersex phenotype is therefore interpreted as resulting from a partial feminization of genetic males. The masculinity rate was 0.14 in the surviving individuals, which is significantly lower than the expected masculinity rate of 0.5 without inoculation (χ2=11.6, d.f.=1, P<0.001). Altogether, these results demonstrate the feminizing effect of wVulM in A. vulgare. PCR assay in females showed that 10 of them harboured Wolbachia, whereas four lacked symbionts. According to the demonstration of the feminizing effect of wVulM in males, females lacking bacteria are genetic females (ie heterogametic females WZ) whereas females harbouring Wolbachia could be either feminized genetic males (ie homogametic females ZZ) or genetic females (Juchault and Legrand, 1972).
We also investigated possible differences in the feminization effects induced by wVulM and wVulC in A. vulgare. Females harbouring either wVulC or wVulM produced the same proportion of lines with biased sex ratios (χ2=0.06, d.f.=1, P=0.81). As expected, female-biased progenies were the most common (66.7% for females harbouring wVulC and 67.5% for those harbouring wVulM, Table 2). However, a small proportion of females produced male-biased progenies (11.1% for females harbouring wVulC and 15.0% for those harbouring wVulM). Male-biased progenies are most likely due to a lack of transmission of feminizing Wolbachia in homogametic females (ZZ), perhaps following the selection of host genes limiting Wolbachia transmission (Rigaud and Juchault, 1992). Lineages with tables 1:1 sex ratio can be explained by the presence of: (i) heterogametic females (WZ) lacking Wolbachia (Juchault and Legrand, 1972) and/or (ii) homogametic females (ZZ) with a Wolbachia transmission rate of 50%. Given that heterogametic females (WZ) rapidly disappear from Wolbachia-infected progenies (Taylor, 1990), explanation (ii) is most likely. Thus, the presence of males in the progenies of infected females suggests that both wVulC and wVulM are not successfully transmitted to all offspring.
In addition, we examined the efficiency of feminization of wVulC and wVulM within progenies. This efficiency was defined by two components: the transmission rate and the penetrance. The transmission rate corresponds to the proportion of individuals in a progeny which inherit Wolbachia from an infected mother. The penetrance refers to the levels of feminization expressed by the Wolbachia present in infected individuals. For example, all descendants of an infected mother might inherit Wolbachia (high transmission rate), but these might not express feminization in the descendants (low penetrance) because of a too small number of bacteria transmitted by the mother. However, all Wolbachia-infected individuals of A. vulgare analysed so far showed female (or at least intersexed) phenotype whatever the infecting strain (Juchault et al, 1980; Bouchon et al, 1998; this study), suggesting consistently high penetrance of both wVulC and wVulM. The comparison of the transmission rates of wVulC and wVulM was performed by measuring the proportion of males within the progenies of infected females, since individuals carrying Wolbachia are females whereas those lacking Wolbachia are males. Comparisons of progenies harbouring either wVulC or wVulM (Table 2) showed that the mean masculinity rate in wVulC-infected progenies (0.18) was significantly lower (χ2=20.7, d.f.=1, P<0.001) than the mean masculinity rate in wVulM-infected progenies (0.27). This indicated that wVulC has a higher vertical transmission rate than wVulM, or wVulC exhibits a more efficient feminizing effect than wVulM. Moreover, significant differences in the primary masculinity rates were recorded according to both Wolbachia strain and host populations (Table 2; whole model test: χ2=33.3, d.f.=3, P<0.001; effect of the Wolbachia strain: L-R χ2=11.9, d.f.=1, P<0.001; effect of the host population: L-R χ2=9.9, d.f.=1, P<0.01; interaction: L-R χ2=0.08, d.f.=1, P=0.77).
Host cytoplasmic types associated to Wolbachia in A. vulgare
The mitochondrial 16SrDNA sequences were obtained from the same individuals whose Wolbachia wsp gene was sequenced, as well as in additional individuals from uninfected populations for comparison. The aim was not to check the whole mtDNA diversity in these populations, but rather to track the history of Wolbachia-host co-evolution. Four mitochondrial lineages named AvI, AvII, AvIII and AvIV were identified (Figure 2). These monophyletic clusters were strongly supported by high bootstrap scores ranging from 97 to 100%. Lineage AvIV was the representative of the most basal lineage of the A. vulgare clade. Lineage AvI was the most common, while lineage AvII was the least common among the studied populations. The inferred tree also indicated a monophyletic origin of Athens and Mery-sur-Cher lineages, which could be considered as a subgroup of AvI lineage. Pairwise distances ranged from 0.0 to 1.0% within lineages, and from 3.0 to 6.3% between lineages (based on the comparison of 399 positions of the mitochondrial 16SrDNA gene). These distance ranges are similar to those observed for the same marker within other species of terrestrial isopods (Michel-Salzat and Bouchon, 2000; Michel-Salzat et al, 2001). The wVulC Wolbachia strain was found in association with the four host lineages. In contrast, the wVulM Wolbachia strain was only associated with AvIII and a subgroup of AvI. A Mantel test showed that the genetic distances of hosts and symbionts from the 16 infected populations were not correlated (r2=0.017; P=0.39). mtDNA sequences from uninfected individuals were found in the two most distantly related lineages AvI and AvIV. Therefore, genetic differentiation of host mtDNA could not be related to the presence of different Wolbachia symbionts in A. vulgare assemblage, suggesting that the association of the two partners does not follow a strict co-evolutionary process.
Cytoplasmic genealogy of A. vulgare. The tree is based on the comparison of 399 bases of the mitochondrial 16SrDNA gene, using the neighbor-joining method and rooted with A. nasatum (AJ388098). Numbers at nodes are bootstrap scores above 50%. Populations harbouring the Wolbachia strain wVulC are identified by the symbol ○ and those harbouring wVulM are identified by the symbol •. Other populations of A. vulgare are uninfected.
Discussion
Based on the wsp sequences, two distantly related Wolbachia strains (wVulC and wVulM) were detected among A. vulgare symbionts sampled from three different continents. Similar to the first described Wolbachia strain (wVulC) in A. vulgare (Rigaud et al, 1991), the newly identified wVulM strain has a feminizing effect. The wVulC strain is widely spread, being associated to all host mitochondrial lineages (Figure 2). On the contrary, wVulM is restricted to two mitochondrial lineages (AvI and AvIII). Despite the large geographical distances between collection sites, mtDNA diversity was not found correlated with geographical origin, suggesting long-distance genetic exchanges between A. vulgare populations. No correlation between hosts and symbionts genetic distances was detected. Moreover, phylogenetic analysis showed that the divergence between the two Wolbachia strains does not result from strict co-evolution with their host, in which case we would have expected the two Wolbachia strains to be sister groups and/or at least some degrees of congruence between symbiont and host phylogenetic trees. These results suggest that wVulC and wVulM were acquired independently by A. vulgare, and strongly support the occurrence of horizontal transmission of Wolbachia within this host.
Inferring the evolutionary steps leading to such Wolbachia distribution in A. vulgare therefore requires gains and losses of symbionts. Gains of symbionts are the result of successful horizontal transfers to a new host. These successes crucially depend upon the propagation potential of the Wolbachia strain involved in the transfer. Looking to results of artificial transfers between isopod hosts (Rigaud and Juchault, 1995; Rigaud et al, 2001), Wolbachia seem to have a weak propagation potential, probably because strains are specialized to their natural hosts and are therefore maladapted to new ones. However, at the within-host species level, such maladaptation is not so strongly expected. Consequently, horizontal transfers within host species are more likely to be successful (Rigaud and Juchault, 1995; Rigaud et al, 2001) and our data in A. vulgare fully support this expectation.
The occurrence of Wolbachia transfers between populations supposes that a strain can occasionally be transferred to a population already harbouring another Wolbachia strain. This implies that, at a certain point, bi-infected individuals may exist. Wolbachia bi-infected and even tri-infected individuals have already been reported in numerous insects (Werren et al, 1995a; Vavre et al, 1999; Werren and Windsor, 2000). By contrast, bi-infected individuals have never been reported so far in isopod host species, including the present study, suggesting bi-infection should be unstable. Furthermore, minimum models of feminization suggest that coexistence of several feminizers in the same population could be unstable at equilibrium (Taylor, 1990). Such instability could result from the rapid loss of the Wolbachia strain with the lower transmission efficiency whenever two strains are in competition within the same host. Our results in A. vulgare support this hypothesis since 75% of the infected samples harbour wVulC, which precisely happens to be associated with a higher transmission rate than wVulM.
Considered in combination with two studies by Rigaud et al (1999, 2001), our results may provide further insight into the infection dynamics of the two Wolbachia strains infecting A. vulgare. Rigaud et al (1999) analysed populations of A. vulgare from western France. Without regard to their geographical origin, all individuals harbouring Wolbachia exhibited closely related mtDNA haplotypes, suggesting an ancestral infection by a single Wolbachia strain. Moreover, Rigaud et al (1999) suggested the possible occurrence in these populations of an unknown Wolbachia strain prior to the arrival of the present-day strain. Our data reveal that the Wolbachia-infected populations in Rigaud et al (1999) study (ie Angoulême, Celles-sur-Belle and Saint-Cyr) harbour the wVulC strain and belong to the AvI mitochondrial lineage. In addition, experimental transfers of Wolbachia in different isopod hosts (Rigaud et al, 2001) revealed that the Wolbachia strain from Celles-sur-Belle (wVulC in the present study) has a wide potential for successfully infecting new hosts, compared to other Wolbachia strains present in isopods. This view is consistent with the results of the present study showing that wVulC has better transmission efficiency than wVulM. Taking into account these data, a possible scenario for the evolution of Wolbachia infection within mitochondrial lineages AvI and AvIII of A. vulgare can be proposed, in which the ancestor of AvI and AvIII harboured wVulM (the ‘resident’). Later, wVulC (the ‘intruder’) invaded independently lineages AvI and AvIII, replacing the ‘resident’ wVulM in most populations, thanks to a better spread potential.
In the alternative scenario, where wVulC is the ‘resident’ and wVulM the ‘intruder’, wVulM would have replaced wVulC in some populations only. This would have required a better propagation potential for wVulM in order to replace wVulC, which is in contradiction with our data indicating a current weaker transmission rate and feminization ability of wVulM compared to wVulC. It is possible that the spread efficiency of a Wolbachia strain can vary over time and one might speculate that the propagation potential of wVulM strongly decreased since its successful competition with wVulC when initially invading A. vulgare populations. However, finding potential reasons for such decrease is not straightforward, especially when an already efficiently transmitted strain is facing competition. Therefore, given the present evidence, the scenario in which wVulM is the ‘resident’ seems more likely.
In conclusion, the discovery of two distantly related feminizing Wolbachia strains infecting the isopod A. vulgare opens new perspectives in the study of both hosts/symbionts and symbionts/symbionts interactions with regard to feminizing effects. Further experimental studies are in progress to examine the survival and spread of Wolbachia strains artificially injected into new host strains, according to the Wolbachia strain, the host mitotype and the previous infection status of the new host. They will provide a better understanding of the evolution of feminizing Wolbachia in isopods.
References
Belkhir K (2000) GENETIX, logiciel sous Windows™ pour la génétique des populations. Laboratoire Génome et Populations, CNRS UPR 9060, Université de Montpellier II, Montpellier, France.
Bethura SK, Sahu SC, Mohan M, Nair S (2001). Wolbachia in the Asian rice gall midge, Orseolia oryzae (Wood-mason): correlation between host mitotypes and infection status. Insect Mol Biol 10: 163–171.
Bouchon D, Rigaud T, Juchault P (1998). Evidence for widespread Wolbachia infection in isopod crustaceans: molecular identification and host feminization. Proc R Soc London B 265: 1081–1090.
Braig HR, Zhou W, Dobson SL, O'Neill SL (1998). Cloning and characterization of a gene encoding the major surface protein of the bacterial endosymbiont Wolbachia pipientis. J Bacteriol 180: 2373–2378.
Cordaux R, Michel-Salzat A, Bouchon D (2001). Wolbachia infection in crustaceans: novel hosts and potential routes for horizontal transmission. J Evol Biol 14: 237–243.
Fialho RF, Stevens L (1997). Molecular evidence for single Wolbachia infection among geographic strains of the flour beetle Tribolium confusum. Proc R Soc London B 264: 1065–1068.
Guillemaud T, Pasteur N, Rousset F (1997). Contrasting levels of variability between cytoplasmic genomes and incompatibility types in the mosquito Culex pipiens. Proc R Soc London B 264: 245–251.
Hurst GDD, Jiggins FM, Schulenburg JHG vd, Bertrand D, West SA, Goriacheva II et al (1999). Male-killing Wolbachia in two species of insect. Proc R Soc Lond B 266: 735–740.
James AC, Ballard JWO (2000). The expression of cytoplasmic incompatibility and its impact on population frequencies and the distribution of Wolbachia strains in Drosophila simulans. Evolution 54: 1661–1672.
Jiggins FM, Hurst GDD, Schulenburg JHG vd, Majerus MEN (2001). Two male-killing Wolbachia strains coexist within population of the butterfly Acraea encedon. Heredity 86: 161–166.
Juchault P, Legrand JJ (1972). Croisements de néo-mâles expérimentaux chez Armadillidium vulgare Latr. (Crustacé, Isopode, Oniscoïde). Mise en évidence d'une hétérogamétie femelle. C R Acad Sci Paris III 258: 2685–2686.
Juchault P, Legrand JJ, Mocquard JP (1980). Contribution à l'étude qualitative et quantitative des facteurs contrôlant le sexe dans les populations du crustacé isopode terrestre Armadillidium vulgare Latr. I – La population de Niort (Deux-Sèvres). Arch Zool Exp Gén 121: 3–27.
Kumar S, Tamura K, Jakobsen IB, Nei M (2001). MEGA2: Molecular Evolutionary Genetics Analysis Software. Arizona State University: Tempe, AZ, USA.
Legrand JJ, Juchault P (1986). Rôle de bactéries symbiotiques dans l'intersexualité, la monogénie et la spéciation chez des crustacés oniscoïdes. Boll Zool 53: 161–172.
Michel-Salzat A, Bouchon D (2000). Phylogenetic analysis of mitochondrial LSU rRNA in oniscids. C R Acad Sci Paris Life Sci 323: 827–837.
Michel-Salzat A, Cordaux R, Bouchon D (2001). Wolbachia diversity in the Porcellionides pruinosus complex of species (Crustacea, Oniscidea): evidence for host-dependent patterns of infection. Heredity 87: 428–434.
Moret Y, Juchault P, Rigaud T (2001). Wolbachia endosymbiont responsible for cytoplasmic incompatibility in a terrestrial crustacean: effects in natural and foreign hosts. Heredity 86: 325–332.
O'Neill SL, Giordano R, Colbert AME, Karr TL, Robertson HM (1992). 16S rRNA phylogenetic analysis of the bacterial endosymbionts associated with cytoplasmic incompatibility in insects. Proc Natl Acad Sci USA 94: 2699–2702.
Rigaud T, Bouchon D, Souty-Grosset C, Raimond R (1999). Mitochondrial DNA polymorphism, sex ratio distorters and population genetics in the isopod Armadillidium vulgare. Genetics 152: 1669–1677.
Rigaud T, Juchault P (1992). Genetic control of the vertical transmission of a cytoplasmic sex factor in Armadillidium vulgare Latr. (Crustacea, Oniscidea). Heredity 68: 47–52.
Rigaud T, Juchault P (1995). Success and failure of horizontal transfers of feminizing Wolbachia endosymbionts in woodlice. J Evol Biol 8: 249–255.
Rigaud T, Juchault P, Mocquard JP (1997). The evolution of sex determination in isopod crustaceans. BioEssays 19: 409–416.
Rigaud T, Pennings P, Juchault P (2001). Wolbachia bacteria effects after experimental interspecific transfers in terrestrial isopods. J Invertebr Pathol 77: 251–257.
Rigaud T, Souty-Grosset C, Raimond R, Mocquard JP, Juchault P (1991). Feminizing endocytobiosis in the terrestrial crustacean Armadillidium vulgare Latr. (Isopoda): recent acquisitions. Endocyt Cell Res 7: 259–273.
Rousset F, Bouchon D, Pintureau B, Juchault P, Solignac M (1992a). Wolbachia endosymbionts responsible for various alterations of sexuality in arthropods. Proc R Soc London B 250: 91–98.
Rousset F, Vautrin D, Solignac M (1992b). Molecular identification of Wolbachia, the agent of cytoplasmic incompatibility in Drosophila simulans, and variability in relation with host mitochondrial types. Proc R Soc London B 247: 163–168.
Schiltuizen M, Honda J, Stouthamer R (1998). Parthenogenesis-inducing Wolbachia in Trichogramma kaykai originates from a single infection. Ann Entomol Soc Am 91: 410–414.
Schulenburg JHG vd, Hurst GDD, Tetzlaff D, Booth GE, Zakharov IA, Majerus MEN (2002). History of infection with different male-killing bacteria in the two-spot ladybird beetle Adalia bipunctata revealed through mitochondrial DNA sequence analysis. Genetics 160: 1075–1086.
Shoemaker DD, Ross KG, Keller L, Vargo EL, Werren JH (2001). Wolbachia infections in native and introduced populations of fire ants (Solenopsis spp.). Insect Mol Biol 9: 661–673.
Stouthamer R, Breeuwer JA, Hurst GD (1999). Wolbachia pipientis: microbial manipulator of arthropod reproduction. Annu Rev Microbiol 53: 71–102.
Suzuki S (1999). Androgenic gland hormone is a sex-reversing factor but cannot be a sex-determining factor in the female crustacean isopod Armadillidium vulgare. Gen Comp Endocrinol 115: 370–378.
Taylor DR (1990). Evolutionary consequences of cytoplasmic sex ratio distorters. Evol Ecol 4: 235–248.
Turelli M, Hoffmann AA, McKechnie SW (1992). Dynamics of cytoplasmic incompatibility and mtDNA variation in natural Drosophila simulans populations. Genetics 132: 713–723.
Vavre F, Fleury F, Lepetit D, Fouillet P, Bouletreau M (1999). Phylogenetic evidence for horizontal transmission of Wolbachia in host–parasitoid associations. Mol Biol Evol 16: 1711–1723.
Werren JH, Bartos JD (2001). Recombination in Wolbachia. Curr Biol 11: 431–435.
Werren JH, Windsor DM (2000). Wolbachia infection frequencies in insects: evidence for a global equilibrium? Proc R Soc London B 267: 1277–1285.
Werren JH, Windsor D, Guo L (1995a). Distribution of Wolbachia among neotropical arthropods. Proc R Soc London B 262: 197–204.
Werren JH, Zhang W, Guo L (1995b). Evolution and phylogeny of Wolbachia: reproductive parasites of arthropods. Proc R Soc London B 261: 55–71.
Zhou W, Rousset F, O'Neill SL (1998). Phylogeny and PCR based classification of Wolbachia strains using wsp gene sequences. Proc R Soc London B 265: 509–515.
Acknowledgements
We are grateful to Jack Werren for providing the American populations of A. vulgare and to Robert Comte for technical assistance. We thank James B Whitfield and Jack Werren for helpful comments on earlier versions of the paper. This research was supported in part by a grant from the Centre National de la Recherche Scientifique (Action incitative CNRS/NSF no 7394).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cordaux, R., Michel-Salzat, A., Frelon-Raimond, M. et al. Evidence for a new feminizing Wolbachia strain in the isopod Armadillidium vulgare: evolutionary implications. Heredity 93, 78–84 (2004). https://doi.org/10.1038/sj.hdy.6800482
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800482
Keywords
This article is cited by
-
Experiment in semi-natural conditions did not confirm the influence of malaria infection on bird attractiveness to mosquitoes
Parasites & Vectors (2022)
-
Evolutionary transition to XY sex chromosomes associated with Y-linked duplication of a male hormone gene in a terrestrial isopod
Heredity (2021)
-
Joint effects of group sex-ratio and Wolbachia infection on female reproductive success in the terrestrial isopod Armadillidium vulgare
BMC Evolutionary Biology (2019)
-
Feminizing Wolbachia influence microbiota composition in the terrestrial isopod Armadillidium vulgare
Scientific Reports (2018)
-
Microbial population analysis of the midgut of Melophagus ovinus via high-throughput sequencing
Parasites & Vectors (2017)