Abstract
The phylogeography of an endangered salmonid, European grayling (Thymallus thymallus), was studied based on analysis of 17 nuclear microsatellite DNA loci. In agreement with earlier mitochondrial DNA (mtDNA) studies, phylogenetic relationships of the populations suggested that northern Europe was colonized from two distinct Pleistocene refugia. Furthermore, microsatellites revealed highly supported grouping of mainland Swedish, Norwegian, Danish, German and Slovenian populations, suggesting that grayling from the northwestern and central Europe have descended from their southern conspecifics. The level of divergence between populations was substantial, even across short geographical distances. Although this was in part due to postglacial colonization patterns and contemporary barriers for gene flow, the high divergence estimates between hydrologically connected sampling locations implied efficient interpopulation reproductive isolation. Microsatellites revealed that the populations exhibited, on average, only 3.5 (±2.2) alleles per locus, indicating that T. thymallus has strikingly low levels of intrapopulation genetic diversity as compared with other freshwater fish species. Accordingly, as indicated by analysis of molecular variance (AMOVA), only 49.1–58.0% of the total grayling microsatellite diversity resided within populations. A latitudinal genetic diversity gradient, potentially resulting from glaciation-mediated founder events, was not evident. Alternatively, it is possible that grayling display limited dispersal behaviour/capability, leading to low long-term effective population sizes and, consequently, depauperate intrapopulation polymorphism. These findings have implications for conservation of T. thymallus. Importantly, they exemplify that microsatellites can be highly informative for intraspecific phylogeography studies dealing with substantial divergence scales.
Similar content being viewed by others
Introduction
Recent theoretical and technical advances in the field of molecular genetics have facilitated the assessment of evolutionary genetic relationships within and between conspecific natural populations. Studies of DNA polymorphisms have revealed that postglacial colonization events have markedly shaped the contemporary distribution of genetic variation in the wild (Avise, 2000). In addition, by identifying genetically diverged populations, such studies have had direct implications for development of appropriate conservation and management guidelines for numerous endangered or threatened species (Beaumont and Bruford, 1999), including salmonid fishes (Angers and Bernatchez, 1998).
European grayling, Thymallus thymallus (Salmonidae), is distributed across a large part of the European continent. Similarly as many other salmonids, T. thymallus exhibits diverse life history strategies; the majority of populations complete their entire life cycle in rivers or lakes (reviewed in Northcote, 1995), some grayling are adapted to a brackish environment and inhabit parts of the Baltic Sea (Nykänen and Huusko, 1999), and some limited anadromy has been reported (Müller and Karlsson, 1983). Although less important for commercial fisheries than many other salmonids, T. thymallus is considered culturally important as a target for sport fisheries. For instance in Finland, both the recreational catch quantity and financial value of European grayling exceed that of Atlantic salmon, Salmo salar (Nylander, 2000). Especially due to locally declining population sizes, grayling is listed among the ‘Threatened fishes of Europe’ (Lelek, 1984) and large-scale stocking practices of the species are now commencing.
Phylogeography of European grayling was recently assessed based on mitochondrial DNA (mtDNA) polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) analysis across ND 5, ND 6, cyt b and D-loop regions, and sequencing of 529 bp from the 5′ end of the ND 5 gene (Koskinen et al, 2000). The observed T. thymallus mtDNA haplotypes divided into three main assemblages exhibiting nucleotide sequence divergence estimates of 1.11–3.59%, implying that divergence of the three lineages clearly pre-dates the late Pleistocene period (defined as 130 000–10 000 years before present; ybp). The geographic distribution of the haplotypes indicated separation of the populations into distinct northeastern (NE), northwestern/central (NW/C) and southern (S) European groups. Consequently, it was concluded that northern Europe was colonized from at least two ancient refugia, and that southern Europe did not contribute markedly to the contemporary genetic diversity of North-European grayling (Koskinen et al, 2000).
A great majority of phylogeography studies have relied solely on assessing mitochondrial variation (reviewed in Avise, 2000). Although mitochondrial DNA has often been informative due to its haploid and non-recombining mode of inheritance and relatively rapid mutation rate, mtDNA is not without its limitations. Commonly mitochondrial studies are based on a small number of genes, and always on just one independently segregating locus, potentially leading to erroneous phylogeographic inference (eg, Pamilo and Nei, 1988). In addition, some empirical studies have shown that selection may severely complicate mtDNA patterns (Hey, 1997). In order to draw firm phylogeographic conclusions, it would therefore seem very important to study the genetic relationships of T. thymallus populations at a European-wide scale also utilizing nuclear DNA loci. Furthermore, as the evolutionary information provided by mtDNA within the T. thymallus main lineages was very limited (Koskinen et al, 2000), assessment of highly polymorphic nuclear markers could potentially provide considerable additional resolution for studying European grayling population structuring at finer geographical scales.
Microsatellites are a class of nuclear DNA loci, consisting of tandemly repeated sequence motifs of two to six base pairs in length. Because of their extremely high level of polymorphism, microsatellites have been widely used for studying genetic differentiation among closely related populations (eg, Bowcock et al, 1994). However, empirical studies have later suggested that the evolutionary informativeness of microsatellites could be hindered after only 3000–30 000 years since divergence, arguably due to frequent occurrence of allele homoplasy (Paetkau et al, 1997). Accordingly, microsatellites have also been directly criticized as phylogeographic markers (Beaumont and Bruford, 1999). In contrast however, some recent evidence implies that microsatellites may be used for resolving phylogenies at substantial divergence scales (Angers and Bernatchez, 1998; Harr et al, 1998; Richard and Thorpe, 2001), indicating that the issue could benefit from further study.
Here, we examined genetic variation at 17 microsatellite loci in order to study the phylogeography of European grayling across a large part of its natural range. Because we used primarily the same individuals as utilized in the mtDNA investigation of Koskinen et al (2000), the complementary data sets allowed us to compare the informativeness of microsatellites with mtDNA a priori expectations.
Materials and methods
Samples and DNA analyses
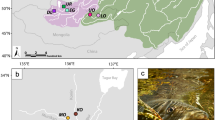
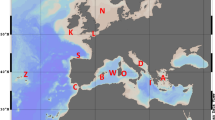
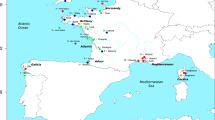
The 623 T. thymallus individuals included in the study originated from a total of 18 sampling locations distributed across Europe (Table 1; Figure 1). The specimens were caught from the wild by electrofishing, nets, ice fishing or fly fishing between 1994 and 1999, or kindly provided by researchers listed in the acknowledgements. Most of the populations are in areas considered to be still unaffected by stocking of grayling. Exceptions were the Lake Saimaa Pielinen population, which has been supplemented with individuals originating from central parts of Lake Saimaa (Makkonen et al, 2000), and the Slovenian Tolminka River, stocked with individuals originating from a Danubian drainage (Susnik et al, 1999). Three hundred and fifty-three of the individuals from 14 sampling locations were also used in the mtDNA study of Koskinen et al (2000).
A total of 17 microsatellite loci were utilized (Figure 2). Twelve of the microsatellites (BFRO004-BFRO018) were previously isolated from T. thymallus and reported to consist of either di- (11 loci) or tetranucleotide (BFRO013) repeat sequences. The five additional loci (Cocl23, Ogo2, One2, Str73INRA and Str85INRA) were derived from other salmonids and cross-amplify polymorphic alleles in grayling. Original references for the microsatellites and details of PCR procedures are outlined in Koskinen and Primmer (2001).
Allele frequencies and size distributions of 17 microsatellite loci in European grayling (Thymallus thymallus, Salmonidae) populations. Areas of the bubbles correspond to frequencies of the respective alleles in given populations, allele lengths are indicated in base pairs and shades of grey distinguish the adjacent loci. Na indicates that, despite numerous attempts, a given locus could not be amplified in a given population. The loci have been ordered as they were electrophoresed in multicolor gel lanes (Koskinen and Primmer, 2001). The horizontal bars denote the main population assemblages, as revealed using mtDNA analyses (see Figure 3 and Koskinen et al, 2000).
Genetic diversity, Hardy-Weinberg equilibrium and linkage equilibrium
Pop100gene program (available at: http://www.ensam.inra.fr/URLB/pop100gene/pop100gene.html) was used to calculate microsatellite allele numbers (A), observed heterozygosities (HO) and expected gene diversities (HE) within the T. thymallus populations. The observed allele numbers within populations were corrected for interpopulation sample size variation (corrected values were termed AC) using Ewens’ correction (Ewens, 1972). Some studies have reported latitudinal diversity declines in intrapopulation diversity, arguably caused by postglacial colonization patterns (see Discussion). To assess whether a declining diversity gradient northwards was evident in T. thymallus, Pearson's correlation was computed between the AC estimates and latitudes of the populations. Probability estimates for the correlation were obtained by permutation using program Permute version 3.4 (available at: http://www.fas.umontreal.ca/biol/legendre/).
Genepop version 3.2a (Raymond and Rousset, 1995) was employed to estimate deviations from Hardy-Weinberg (H-W) equilibrium across populations (within loci) and across loci (within populations) using the probability test, and to estimate deviations from genotypic linkage equilibrium (L-E) across all populations. Corrections for multiple significance tests were performed using Fisher's method, as implemented in Genepop, and applying a sequential Bonferroni type correction (Rice, 1989).
Genetic divergence and relationships between T. thymallus populations
Genepop was applied to test for statistically significant genic differentiation between populations (corrections for multiple tests were performed as indicated above) and to estimate interpopulation FST values, ie variance in allele frequencies (Weir and Cockerham, 1984). Because it has been suggested that comparison of variance in allele frequencies (FST) and variance in allele sizes (rhoST) may provide valuable insights to the relative effects of genetic drift and mutations in shaping patterns of genetic differentiation (Slatkin, 1995), we also utilized Genepop to compute pairwise rhoST estimates (Michalakis and Excoffier, 1996). Differences in the two estimators were assessed using Wilcoxon's signed rank test. For computations of the rhoST estimates and further stepwise mutation model (SMM) based estimators (see below), the allele sizes were converted into repeat units by dividing the observed allele sizes by the reported repeat motif lengths of the microsatellites. The microsatellite Cocl23 was originally published as a dinucleotide locus. However, in our European-wide T. thymallus data set Cocl23 always exhibited alleles five base pairs apart, and its repeat numbers were computed accordingly. The locus BFRO013 was published as a tetranucleotide marker. However, we observed some intermediate allele sizes in BFRO013, ie, alleles two base pairs apart. Consequently, this locus was excluded from the analyses that assume SMM.
Genetic distances between populations were estimated using the infinite allele model (IAM) based DA distance (Nei et al, 1983), chord distance (DCE) of Cavalli-Sforza and Edwards (1967), the SMM based DSW distance (Shriver et al, 1995) and the DAS distance (Bowcock et al, 1994). These metrics were chosen in order to have a range of evolutionary assumptions covered, which can be prudent for microsatellite based phylogenetic inference (Slatkin, 1995). The resulting genetic distance matrices were used to construct neighbor-joining (N-J) phylograms and confidence estimates on the tree topologies were attained by resampling over loci with 1000 bootstrap replicates. The genetic distance estimation and bootstrapping procedures were carried out using computer programs MsatBoot version 1.1, kindly provided by Pierre-Alexandre Landry (University of Helsinki, Finland), and Microsat (Minch et al, 1996). N-J phylograms and consensus trees were constructed using programs Neighbor and Consense, respectively. Branch lengths of the consensus trees were obtained by importing the distance matrices and consensus trees into the program Fitch, and applying the ‘use user trees from input file’ option. Neighbor, Consense and Fitch are implemented in Phylip version 3.57c program (Felsenstein, 1995).
Analysis of molecular variance
Hierarchical analysis of molecular variance (AMOVA), based on allele frequency information (Excoffier et al, 1992), was carried out using the program Arlequin version 2.0 (Schneider et al, 1997). Variance components were extracted for the (i) among group; (ii) between sampling sites within groups; and (iii) within sampling sites hierarchical levels. The populations were partitioned into groups (Table 2) following earlier mtDNA based findings (Koskinen et al, 2000), and in order to shed further light on the genetic affinities of the hypothesized T. thymallus lineages (see Table 2; Koskinen et al, 2000).
Results
Microsatellite DNA diversity within and between T. thymallus populations
The mean number of alleles per locus (across the 17 microsatellite loci) within populations ranged from 1.8 to 5.0 (Table 1). The mean number of alleles per locus within populations was 2.5–5.4 after applying the Ewens’ (1972) correction for unequal sample sizes (Table 1). The lowest allele number within populations (at any single locus) was one and the highest 25 (Table 1). Average observed heterozygosity within populations varied from 0.18 to 0.59 and expected gene diversity within populations from 0.19 to 0.61 (Table 1). No significant association was revealed between the intrapopulation AC estimates and the latitude of the populations (Pearson r = 0.11; P = 0.65). The level of polymorphism per locus across all populations was much higher, the total number of alleles averaging 15.7 and ranging from five to 58 at any single locus (Figure 2). Despite numerous attempts, three of the microsatellites could not be successfully amplified in a small number of populations (Figure 2) and were coded as missing data for the subsequent analyses.
Hardy-Weinberg and linkage equilibrium
Hardy-Weinberg probability tests across populations suggested significant (P < 0.05) disequilibrium in one locus (One2) after correcting for multiple tests. The deviation of One2 appeared to be caused by the Pielinen (Pie) population. After exclusion of Pie individuals, H-W disequilibrium was not significant in any of the microsatellites. Accordingly, probability tests across loci revealed one population (Pie) to deviate significantly (P < 0.05) from H-W equilibrium after correcting for multiple tests (Table 1). The deviations in Pie occurred at five of the loci analysed and were always due to a heterozygote deficiency, observations suggesting a Wahlund effect. Individual genotype based analyses have later implied that this may be due to genetically differentiated individuals stocked into Pielinen from the central parts of Lake Saimaa (Koskinen et al, 2002). Because of its significant H-W disequilibrium, the Pielinen population was excluded from estimation of genetic differentiation and phylogenetic relationships. Genotypic linkage equilibrium tests across populations suggested significant (P < 0.05) linkages only between the loci BFRO007 and BFRO013, and loci BFRO011 and BFRO018 after correcting for multiple tests. Removal of any one of these loci did not markedly affect the obtained T. thymallus population relationships.
Genetic differences and phylogenetic relationships among T. thymallus populations
Many microsatellite alleles were geographically confined in the European grayling populations investigated (Figure 2). For instance, all of the populations originating from northeastern Europe (Russia, Finland and Bothnian Bay) displayed an allele BFRO012*133, and the allele was not observed in any other population. Similarly, all of the populations originating from southern Europe (southern Germany and Slovenia) were fixed for, or exhibited a high frequency of, an allele Str85INRA*178, but this allele was absent in all other populations, except the central German Eger sample where it occurred at a frequency of 4% (Figure 2).
Accordingly with the geographically distinct allele size distributions, highly significant genic differentiation (P < 0.001) was revealed between all populations and, consequently, the null-hypothesis of random mating could be rejected for all interpopulation comparisons. A substantial level of genetic differentiation was generally observed between the sampling locations, with FST estimates ranging from 0.03 to 0.74 (mean = 0.40) and rhoST estimates from 0.01 to 0.89 (mean = 0.41) (see Appendix). Clear differentiation was often observed also across short geographical distances and, importantly, also within areas where populations should have had possibilities to exchange migrants postglacially; for instance, the Ulkokrunnit and Tornionjoki populations (Figure 1) exhibited pairwise FST and rhoST distances of 0.19 and 0.09, respectively (Appendix). The allele frequency variance and allele size variance based estimators (FST and rhoST, respectively) were generally concordant in characterizing genetic differentiation between populations (Appendix), ie correlation between the two matrices was high (0.76). The rhoST/FST ratio between the northwestern/central and northeastern populations exceeded 1.0 in 28/45 of the pairwise comparisons (Appendix; mean = 1.13). A two-tailed Wilcoxon's signed rank test indicated that the difference in the pairwise rhoST and FST values was highly significant (T− = 190; T0.001, 45 = 233; P < 0.001).
Relatively clear phylogenetic relationships were observed among the T. thymallus populations. The DCE, DA and DAS metrices indicated grouping of the southernmost grayling (Ramsach, Obrh and Tolminka) with bootstrap support estimates of 94% to 97% (Figure 3). The DCE, DA and DAS distances further revealed grouping of all populations from mainland Sweden, Norway, central Europe and those from southern Europe with bootstrap estimates of 86%, 85% and 74%, respectively. Thereby, these populations were confidently separated from the northeastern T. thymallus populations (Figure 3). A topology very similar to that presented in Figure 3 was recovered also with the DSW distance, however, the bootstrap estimates for the nodes defining the above-mentioned major clusters were not as high (Figure 3). In addition to defining the major population clusters, all of the distance metrices indicated several internal subgroups among the NW/C and NE populations (Figure 3). For example, populations from northern Finland (Tenojoki, Lätäseno and Tornionjoki) formed a distinct group, as did the populations from Kola Peninsula (Varzuga and Juzija) (Figure 3). The genetic T. thymallus population relationships, as revealed using microsatellites, were strikingly similar to those summarized by mtDNA data (Figure 3).
Genetic relationships in European grayling (Thymallus thymallus, Salmonidae), as resolved by mtDNA and microsatellite analyses. The mtDNA gene tree (re-drawn from Koskinen et al 2000) is based on Wagner parsimony analysis of pooled PCR-RFLP site data, covering the ND 5, ND 6, cyt b and D-loop regions, and 529 bp of ND 5 sequence data. The haplotype numbers and bootstrap support values for the three main grayling lineages correspond to Koskinen et al (2000). The microsatellite N-J population phylogram is based on the analysis of 17 loci and the DCE genetic distance. The first values beside the branches of the tree represent DCE bootstrap support over 1000 replicates of resampled loci. The second, third and fourth values indicate bootstrap estimates as revealed by the DA, DAS and DSW distances, respectively. The vertical bars of the mtDNA and microsatellite trees denote the northeastern (NE; white), northwestern/central (NW/C; black) and southern (S; grey) genetic lineages. The pie charts of the map (re-drawn from Koskinen et al 2000) indicate the distribution of the resolved mtDNA haplotype lineages in European grayling populations (note that each one of the populations exhibited mtDNA haplotypes from only a single lineage). The shades of grey of the pie charts correspond to those of the vertical bars denoting the main mtDNA and microsatellite phylogram lineages.
Analysis of molecular variance
The AMOVA analysis revealed that 49.1–58.0% of the total T. thymallus genetic variance resided within populations, a substantial proportion (42.0–50.9%) of the variance being at the within-group and among-group hierarchical levels (Table 2). When the populations were partitioned into three groups according to the three main mtDNA lineages (Koskinen et al, 2000), 19.7% of the total variance resided between the assemblages (P < 0.001) (Table 2). This supported the distinctiveness of the three assemblages. When the NE populations were grouped with the NW/C populations, 26.2% of the variance resided within groups. When the NE and the NW/C populations were grouped with the S grayling, 35.4% and 30.1% of the variance was at the within-group level, respectively (Table 2). Hence, these observations implied that (i) the NE and NW/C populations are more closely related to each other than they are to the S populations; and (ii) the NW/C and S populations are more closely related than the NE and S populations.
Discussion
Postglacial dispersal of T. thymallus
Microsatellite data provided high resolution of T. thymallus evolutionary relationships. Interestingly, the northwestern, central European and the South-European grayling populations grouped together with high bootstrap support (Figure 3). Earlier mtDNA analyses had implied an equidistant split between the southern, central/northwestern and northeastern populations, leading to the suggestion that the southern populations did not contribute to the genetic diversity of North-European regions (Koskinen et al, 2000). The microsatellite phylograms, however, now open the possibility that southern European grayling were the ancestors of the NW/C populations. A feasible northward colonization route would have been via the Danube into the Elbe during ancient times (until the early Pleistocene, circa 1.8 million ybp), when the headwaters of the Elbe drained into the upper Danube (Hantke, 1993). An ancient Danube-Elbe dispersal route was also supported by the fact that the Eger River population (originating from a tributary of the Elbe) grouped together with the South-European populations, two of which (Ram and Obr) originated from tributaries of the Danube (Figure 3). Although this connection has not been previously observed in grayling, other fish species inhabiting central and northern Europe have been argued to originate from their South-European conspecifics (Nesb et al, 1999; Englbrecht et al, 2000). Furthermore, the Danube-Elbe colonization route has been recently discussed to explain mtDNA haplotype patterns in bullhead, Cottus gobio (Englbrecht et al, 2000).
The southern populations (Ram, Obr and Tol) formed a distinct subgroup (Figure 3). The genetic distinctiveness of this group was evident also when looking at the allelic size distributions, eg at the locus Str85INRA (Figure 2). It was sensible that also the German Ramsach population was placed within this group, because it originates from a Danubian drainage, similarly as the Slovenian Obrh population (Figure 1). Highly supported clustering of the Obrh and Tolminka populations may be a reflection of extensive supplementation of the Tolminka river using individuals originating from the Danube (Susnik et al, 1999).
A clear result from the N-J dendrogram was the resolution of the genetic relationships between the central German, Danish, Norwegian and mainland Swedish populations (Figure 3). Relationships among these populations provided evidence for the colonization of the NW grayling from a central European origin, which likely acted as a source also for the colonization of the German Eger and Danish Skjern rivers, perhaps via the Elbe drainage. Similar observations have also been reported in whitefish, Coregonus sp., based on mitochondrial analyses (Hansen et al, 1999). The phylogenetic trees (Figure 3), as well as the AMOVA analyses (Table 2), implied that the Baltic Sea region is currently inhabited by populations that originated from two diverged sources. Namely, the northeastern populations appeared very distinct from the lineage that arguably colonized North-West from a central European refugium (see above). Exact dating of the coalescence of the NE and NW/C lineages requires further study. However, AMOVA grouping of the NE and NW/C populations resulted in a lower within-group component of molecular variance than grouping of the NE with S or NW/C with the S populations (Table 2). Consequently, it appears possible that separation of the North-European population assemblages (NE and NW/C) post-dates their divergence from the southern populations.
When populations have been isolated for more than 2000 generations (see Estoup and Angers, 1998), novel microsatellite mutations arguably lead to rhoST:FST ratio >1.0 (Slatkin, 1995). This was exactly the case in the rhoST/FST comparison between the northwestern/central and northeastern grayling populations (P < 0.001). Assuming a T. thymallus generation interval of 5 years, the result suggests that separation of the NW/C and NE populations exceeds 10 000 years. As the major ice sheets receded from northern Europe at approximately 10 000 ybp (Svendsen et al, 1999), the observation reinforces that northern Europe was colonized from at least two isolated Pleistocene refugia.
The differentiated NE and NW/C population groups met at a sharp border zone located between the Holmön and Vindelälven sampling sites in northern Sweden. Northern Sweden has been previously noted to be a probable suture zone for various European mammals based on mtDNA evidence (Taberlet et al, 1998). Microsatellite data from European grayling now exemplify, that such a pattern may also be common for North-European fish species. The colonization of the Norwegian Lesjaskogsvatn water system could not be confidently resolved based on mtDNA analyses, as the population was fixed for a haplotype dominating in the NE populations, most likely due to a human-mediated founder effect (Koskinen et al, 2000). However, microsatellites disclosed the close genetic relationships between the southern Swedish Lake Vättern and Lesjaskogsvatn populations (Figure 3). This relationship most likely reflects colonization of the Glomma River system (connected to Lesjaskogsvatn) via its connection to Lake Vänern, southern Sweden, circa 10 200 years ago. Also populations of perch, Perca fluviatilis, have been suggested to have utilized this colonization route (Refseth et al, 1998).
Deep genetic divergence across a range of geographical scales: potential roles of pre- and postglacial divergence
The level of genetic divergence between the T. thymallus populations originating from across Europe was substantial. Marked differentiation was often observed also across short geographical distances. For instance, the DCE estimate between the populations Eger and Ramsach (DCE = 0.64; Figure 3), separated by a great circle distance of 260 km, was higher than the corresponding estimate reported between non-anadromous populations of Arctic char from Lake Saimaa (eastern Finland) and Lake Riasten (southern Norway), with a great circle distance of 850 km (DCE = 0.49; Primmer et al, 1999). In some cases, such extreme European grayling interpopulation divergence estimates resulted from postglacial colonization of the populations from separate ice-age refugia, ie reproductive isolation may have already existed between many of the populations at the end of the last glacial period. Furthermore, possibilities for contemporary mixing among most of the studied populations have been virtually non-existent due to geographic barriers for gene flow; in the absence of the homogenizing effect of migration, postcolonization divergence of the populations may have also been efficient. Physical migration barriers have been formed, for example, by land. For instance, populations Eger and Ramsach compared above inhabit rivers that have been isolated since the early Pleistocene (Hantke, 1993). In addition, saline environments could have restricted migration attempts, as T. thymallus is intolerant to salt concentrations above 4‰.
However, populations within the Bothnian Bay and its coastal rivers (Lät, Tor, Kru, Hol and Vin), Varzuga River (Var and Juz) and Lake Saimaa (Pie, Lie and Esa) have been in hydrological contact since recolonizing their present habitats, making postcolonization mixing feasible. Even so, the level of interpopulation divergence was substantial also within these regions. For instance, the DCE estimate between the Ulkokrunnit (Kru) and Tornionjoki (Tor) populations was similar than between the over 10 times more distantly located Arctic char populations listed above (results not shown). In such cases, there are two alternative explanations for the observed level of European grayling divergence: (i) contemporary populations within each of the above water systems have originated from more than one already reproductively isolated glacial refugium and persisted in isolation also in postglacial times, despite the potential for physical contact. This has been recently argued to have occurred, for example, between some lake whitefish (Coregonus clupeaformis) populations in St. John River basin, northeastern North America (Lu et al, 2001); or (ii) the populations have originated from the same panmictic ancestral refugium, but postglacial interpopulation migration has been sufficiently limited to enable genetic drift and/or mutations to cause the present divergence. Rigorous distinction between these alternatives may benefit from denser sampling across the natural range of T. thymallus. At present, however, there is no evidence to suggest that populations within each of the above continuous water bodies would originate from more than one ice-age refugium, leaving alternative ii) as the most parsimonious explanation.
Informativeness of microsatellites in phylogeography studies
An important feature of this study was, that the microsatellite results could be directly compared with mtDNA based a priori expectations (Figure 3; Koskinen et al, 2000). The two complementary data sets exemplified that microsatellites were not only able to provide the same inference than the mtDNA data, but also to provide additional phylogeographic resolution across the natural range of a widely distributed and highly structured species. This finding was noteworthy given that many of the grayling populations have most likely diverged >400 000 years ago (Koskinen et al, 2000). Based on some empirical evidence, the utility of microsatellites for resolving evolutionary relationships across more than 3000–30 000 generations has been questioned (Paetkau et al, 1997), however, the T. thymallus data exemplify that such conclusions may be tenuous. In line with this suggestion are some recent data from brook charr (Salvelinus fontinalis; Angers and Bernatchez, 1998), fruitfly (Drosophila melanogaster; Harr et al, 1998) and the Western Canary Island Lizard (Gallotia galloti; Richard and Thorpe, 2001), implying that microsatellites have potential for resolving intraspecific phylogenies even across >200 000 years since divergence. It remains to be determined whether the evolutionary informativeness of microsatellites relates to the biological features of the species being studied, properties of the loci, or combination of these two factors. In the case of European grayling, it appears that the larger than average number of loci assessed was a key factor (Koskinen et al, submitted).
Low levels of genetic diversity in T. thymallus populations across Europe
In a recent meta-analysis of microsatellite polymorphism displayed by freshwater, anadromous and marine fishes, DeWoody and Avise (2000) reported freshwater fish populations to display, on average, 9.1 (±6.1) alleles and expected gene diversities of 0.54 (±0.25). With mean allele number and HE within populations of only 3.5 (±2.2) and 0.41 (±0.27), respectively (Table 1), T. thymallus undoubtedly exemplifies a species with a low level of within-population microsatellite diversity. Because the overall diversity of the 17 microsatellites (across all populations) was relatively high (Figure 2), the low level of intrapopulation variation appeared not to be an attribute of the loci, but rather of the individual populations. Accordingly, the AMOVA analyses demonstrated that a substantial proportion of grayling molecular variation resided between populations (Table 2).
What could explain the low intrapopulation allele numbers and heterozygosity estimates in T. thymallus? Dramatic population size reductions, such as those mediated by the Pleistocene glaciations, may potentially cause decreased variation in populations inhabiting formerly glaciated regions. For instance, glaciations have been argued to contribute to the low intrapopulation diversity observed in another freshwater salmonid, bull trout, Salvelinus confluentus (Taylor et al, 2001). An important consequence of this theory is, that because many northern regions were formerly completely covered with ice (including the whole of northern Europe; Svendsen et al, 1999), the contemporary northern populations should display lower levels of molecular variation than southern populations (reviewed in Bernatchez and Wilson, 1998). In T. thymallus a latitudinal genetic diversity gradient was by no means evident: eg, the southernmost populations Obrh and Tolminka from Slovenia exhibited diversity indices below or close to the overall average (Table 1) and no significant correlation was revealed between allele numbers and latitudes of the populations.
Alternatively, the low microsatellite diversity might relate to the life history characteristics of European grayling. One possibility is, that grayling individuals display limited dispersal behaviour/capability from their natal sites. This could result in low long-term effective population sizes and, consequently, depauperate levels of within-population T. thymallus microsatellite diversity. Mark-recapture and radio-telemetry studies support this hypothesis, as they have revealed grayling to be far less mobile than autumn spawning salmonids (Peterson, 1968; Northcote, 1995). Limited dispersal of grayling was also suggested by a recent microgeographic study within Lake Saimaa, eastern Finland, where microsatellite-based individual assignment tests were applied to investigate the genetic distinctiveness of three populations (Koskinen et al, 2001). It was shown that 100% of the spawning individuals could be assigned to their correct source population, while being confidently (P ≤ 0.05) excluded from all alternative reference populations. Consequently, it was suggested that interpopulation dispersal was extremely limited (Koskinen et al, 2001). Finally, it is noteworthy, that grayling juveniles have been reported to favour shallow habitats, presumably due to their relatively poor swimming performance (Nykänen and Huusko, 1999). It has been suggested that this tendency may result in massive juvenile mortality when water levels fluctuate, because grayling juveniles have poor capability to switch habitat into deeper waters, unlike S. trutta or S. salar (Peterson, 1968; Nykänen and Huusko, 1999). Such year class ‘drop-out’ events could decrease effective population sizes and genetic diversity.
Conservation implications
One alternative for direct application of genetic results in the development of appropriate conservation guidelines involves recognition of so-called management units (MU). Distinct MUs have been defined as populations connected by little or no contemporary gene flow, but not separated historically for very long periods of time (Ricker, 1972; Waples, 1991; Moritz, 1994). All of the European grayling populations included in this study would appear to fulfill such criteria for being recognized as separate MUs. Furthermore, it was noteworthy that a large proportion of the total T. thymallus genetic variance resided between populations, intrapopulation diversity being atypically low (Tables 1 and 2; Figure 2). Genetic data based on this and also earlier studies (Koskinen et al, 2000, 2001, 2002) suggest that European grayling populations should be supplemented from distinct broodstocks even across relatively short geographical distances.
References
Angers, B, Bernatchez, L (1998). Combined use of SMM and non-SMM methods to infer fine structure and evolutionary history of closely related brook charr (Salvelinus fontinalis, Salmonidae) populations from microsatellites. Mol Biol Evol, 15: 143–159.
Avise, JC (2000). Phylogeography. Harvard University Press: Cambridge, Massachusetts.
Beaumont, M, Bruford, MW (1999). Microsatellites in conservation genetics. In: Goldstein DB, Schlötterer C (eds) Microsatellites Evolution and Applications. Oxford University Press: New York. pp 165–182.
Bernatchez, L, Wilson, CC (1998). Comparative phylogeography of Nearctic and Palearctic fishes. Mol Ecol, 7: 431–452.
Bowcock, AM, Ruiz-Linares, A, Tomfohrde, J, Minch, E, Kidd, JR, Cavalli-Sforza, LL (1994). High resolution of human evolutionary trees with polymorphic microsatellites. Nature, 368: 455–457.
Cavalli-Sforza, LL, Edwards, AWF (1967). Phylogenetic analysis: models and estimation procedures. Am J Hum Genet, 19: 233–257.
Dewoody, JA, Avise, JC (2000). Microsatellite variation in marine, freshwater and anadromous fishes compared with other animals. J Fish Biol, 56: 461–473.
Englbrecht, CC, Freyhof, J, Nolte, A, Schliewen, U, Tautz, D (2000). Phylogeography of the bullhead Cottus gobio (Pisces: Teleostei: Cottidae) suggests a pre-Pleistocene origin of the major central European populations. Mol Ecol, 9: 709–722.
Estoup, A, Angers, B (1998). Microsatellites and minisatellites for molecular ecology: theoretical and empirical considerations. In: Carvalho GR (ed), Advances in Molecular Ecology, IOS Press: Amsterdam. pp55–86.
Ewens, WJ (1972). The sampling theory of selectively neutral alleles. Theor Popul Biol, 3: 87–112.
Excoffier, L, Smouse, P, Quattro, JM (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics, 131: 479–491.
Felsenstein, J (1995). Phylip (Phylogeny Inference Package) version 3.57c. Department of Genetics, University of Washington, Seattle, WA.
Hansen, MM, Mensberg, K-LD, Berg, S (1999). Postglacial recolonization patterns and genetic relationships among whitefish (Coregonus sp.) populations in Denmark, inferred from mitochondrial DNA and microsatellite markers. Mol Ecol, 8: 239–252.
Hantke, R (1993). Flugeschichte Mitteleuropas. Ferdinand Enke-Verlag: Stuttgart.
Harr, B, Weiss, S, David, JR, Brem, G, Schlötterer, C (1998). A microsatellite-based multilocus phylogeny of the Drosophila melanogaster species complex. Curr Biol, 8: 1183–1186.
Hey, J (1997). Mitochondrial and nuclear genes present conflicting portraits of human origins. Mol Biol Evol, 14: 166–172.
Koskinen, MT, Piironen, J, Primmer, CR (2001). Interpopulation genetic divergence in European grayling (Thymallus thymallus, Salmonidae) at a microgeographic scale: implications for conservation. Conserv Genet, 2: 133–143.
Koskinen, MT, Primmer, CR (2001). High throughput analysis of 17 microsatellite loci in grayling (Thymallus spp. Salmonidae). Conserv Genet, 2: 173–177.
Koskinen, MT, Ranta, E, Piironen, J, Veselov, A, Titov, S, Haugen, TO et al (2000). Genetic lineages and postglacial colonization of grayling (Thymallus thymallus, Salmonidae) in Europe, as revealed by mitochondrial DNA analyses. Mol Ecol, 9: 1609–1624.
Koskinen, MT, Sundell, P, Piironen, J, Primmer, CR (2002). Genetic assessment of spatiotemporal evolutionary relationships and stocking effects in grayling (Thymallus thymallus, Salmonidae). Ecol Letters. (in press).
Lelek, A (1984). Threatened fishes of Europe. In: European Committee for the Conservation of Nature and Natural Resources-Council of Europe (ed), The Freshwater Fishes of Europe 9. AULA-Verlag: Wiesbaden. pp 93–96.
Lu, G, Basley, DJ, Bernatchez, L (2001). Contrasting patterns of mitochondrial DNA and microsatellite introgressive hybridization between lineages of lake whitefish (Coregonus clupeaformis); relevance for speciation. Mol Ecol, 10: 965–985.
Makkonen, J, Westman, K, Pursiainen, M, Heinimaa, P, Eskelinen, U, Pasanen, P, Kummu, P (2000). Viljelykantarekisteri Riista-ja kalatalouden tutkimuslaitoksen kalanviljelylaitoksissa ja maitipankissa säilytyksessä olevat kalalajit ja kannat. Riista-ja kalatalouden tutkimuslaitos: Helsinki (In Finnish).
Michalakis, Y, Excoffier, L (1996). A generic estimation of population subdivision using distances between alleles with special interest to microsatellite loci. Genetics, 142: 1061–1064.
Minch, E, Ruiz-Linares, A, Goldstein, D, Feldman, M, Cavalli-Sforza, LL (1996). Microsat (version 1.5): A computer program for calculating various statistics on microsatellite allele data. Stanford University Medical Center, Stanford, CA.
Moritz, C (1994). Defining ‘evolutionary significant units’ for conservation. Trends Ecol Evol, 9: 373–375.
Müller, K, Karlsson, L (1983). The biology of the grayling, Thymallus thymallus L., in coastal areas of the Bothnian Bay. Aq Ser Zool, 22: 65–68.
Nei, M, Tajima, F, Tateno, Y (1983). Accuracy of estimated phylogenetic trees from molecular data. J Mol Evol, 19: 153–170.
Nesb, CL, Fossheim, T, Vllestad, LA, Jakobsen, KS (1999). Genetic divergence and phylogeographic relationships among European perch (Perca fluviatilis) populations reflect glacial refugia and postglacial colonization. Mol Ecol, 8: 1387–1404.
Northcote, TG (1995). Comparative biology and management of Arctic and European grayling (Salmonidae, Thymallus). Rev Fish Biol Fish, 5: 141–194.
Nykänen, M, Huusko, A (1999). Harjuksen elinympäristövaatimukset virtavesissä. Riista- ja kalatalouden tutkimuslaitos, kalatutkimuksia, 156. Helsinki (in Finnish).
Nylander, E (2000). Finnish Fisheries Statistics. Finnish Game and Fisheries Research Institute: Helsinki.
Paetkau, D, Waits, LP, Clarkson, PL, Craighead, L, Strobeck, C (1997). An empirical evaluation of genetic distance statistics using microsatellite data from bear (Ursidae) populations. Genetics, 147: 1943–1957.
Pamilo, P, Nei, M (1988). Relationships between gene trees and species trees. Mol Biol Evol, 5: 568–583.
Peterson, HH (1968). The grayling, Thymallus thymallus (L.), of the Sudsvall Bay area. Report Freshw Res Inst Drottningholm, 48: 36–56.
Primmer, CR, Aho, T, Piironen, J, Estoup, A, Cornuet, J-M, Ranta, E (1999). Microsatellite analysis of hatchery stocks and natural populations of Arctic charr, Salvelinus fontinalis, from the Nordic region: implications for conservation. Hereditas, 130: 277–289.
Raymond, M, Rousset, F (1995). GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered, 86: 248–249.
Refseth, UH, Nesb, CL, Stacy, JE, Vollestad, LA, Fjeld, E, Jakobsen, KS (1998). Genetic evidence for different migration routes of freshwater fish into Norway revealed by analysis of current perch (Perca fluviatilis) populations in Scandinavia. Mol Ecol, 7: 1015–1027.
Rice, WR (1989). Analysing tables of statistical tests. Evolution, 43: 223–225.
Richard, M, Thorpe, RS (2001). Can microsatellites be used to infer phylogenies? Evidence from population affinities of the Western Canary Island lizard (Gallotia galloti). Mol Phylogenet Evol, 20: 351–360.
Ricker, WE (1972). Hereditary and environmental factors affecting certain salmonid populations. In: Simon RC, Larkin PA (eds) The Stock Concept in Pacific Salmon, University of British Columbia: Institute of Animal Resource Ecology, Vancouver. pp 19–160.
Schneider, S, Kueffer, J-M, Roessli, D, Excoffier, L (1997). Arlequin, Version 1.1. Genetics and Biometry Laboratory, Department of Anthropology, University of Geneva, Switzerland.
Shriver, MD, Jin, L, Boerwinkle, E, Deka, R, Ferrel, RE, Chakraborty, R (1995). A novel measure of genetic distance for highly polymorphic tandem repeat loci. Mol Biol Evol, 12: 914–920.
Slatkin, M (1995). A measure of population subdivision based on microsatellite allele frequencies. Genetics, 139: 457–462.
Sunsik, S, Snoj, A, Dovc, P (1999). Microsatellites in grayling (Thymallus thymallus): comparison of two geographically remote populations from the Danubian and Adriatic river basin in Slovenia. Mol Ecol, 8: 1756–1758.
Svendsen, JI, Astakhov, VI, Bolshiyanov, DYU, Demidov, I, Dowdeswell, JA, Gataullin, V et al (1999). Maximum extent of the Eurasian ice sheets in the Barents and Kara Sea region during the Weichselian. Boreas, 28: 234–242.
Taberlet, P, Fumagalli, L, Wust-Saucy, A-G, Cossons, J-F (1998). Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol, 7: 453–464.
Taylor, EB, Redenbach, Z, Costello, AB, Pollard, SM, Pacas, CJ (2001). Nested analysis of genetic diversity in northwestern North American char, Dolly Warden (Salvelinus malma) and bull trout (Salvelinus confluentus). Can J Fish Aquat Sci, 58: 406–420.
Waples, RS (1991). Pacific salmon, Oncorhynchus spp. and the definition of a ‘species’ under the Endangered Species Act. Mar Fish Rev, 53: 11–22.
Weir, BS, Cockerham, CC (1984). Estimating F-statistics for the analysis of population structure. Evolution, 38: 1358–1370.
Acknowledgements
The European grayling samples used in the study were kindly provided by numerous researchers. Especially we acknowledge Jorma Piironen, Teija Aho, Jaakko Lumme, Ari Savikko, Timo Hartikainen (Finland), Michael Hansen (Denmark), Thrond Haugen (Norway), Simona Susnik, Ales Snoj (Slovenia), Bernhard Gum and Ralph Kühn (Germany). Louis Bernatchez and anonymous reviewers are much appreciated for their constructive comments on earlier versions of this manuscript. The work was supported by grants from the University of Helsinki, the Biological Interactions Graduate School, the Finnish Game and Fisheries Research Institute, the Finnish Ministry of Agriculture and Forestry and the Finnish Academy (project number 172964).
Author information
Authors and Affiliations
Corresponding author
Appendix 1
Appendix 1
(See Table below)
Rights and permissions
About this article
Cite this article
Koskinen, M., Nilsson, J., Veselov, A. et al. Microsatellite data resolve phylogeographic patterns in European grayling, Thymallus thymallus, Salmonidae. Heredity 88, 391–401 (2002). https://doi.org/10.1038/sj.hdy.6800072
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800072
Keywords
This article is cited by
-
Large-scale genetic structure and diversity of Arctic rainbow smelt Osmerus dentex Steindachner et Kner, 1870 throughout its distributional range based on microsatellites
Polar Biology (2021)
-
Development of 21 Microsatellite Loci and Diversity Analysis of Amur Grayling in Amur River
Thalassas: An International Journal of Marine Sciences (2020)
-
Complex phylogeographic relationships among the Eurasian perch (Perca fluviatilis) populations in the eastern part of the Baltic Sea Region
Hydrobiologia (2020)
-
Phylogeography of the white-clawed crayfish (Austropotamobius italicus) in Spain: inferences from microsatellite markers
Molecular Biology Reports (2013)
-
Microsatellite markers reveal clear geographic structuring among threatened noble crayfish (Astacus astacus) populations in Northern and Central Europe
Conservation Genetics (2013)