Abstract
Spermatocyte chromosomes of Melarhaphe neritoides (Mollusca, Prosobranchia, Caenogastropoda) were studied using fluorescent in situhybridization (FISH) with four repetitive DNA probes (18S rDNA, 5S rDNA, (TTAGGG)n and (GATA)n). Single-colour FISH consistently mapped one chromosome pair per spread using either 18S or 5S rDNA as probes. The telomeric sequence (TTAGGG)n hybridized with termini of all chromosomes whereas the (GATA)n probe did not label any areas. Simultaneous 18S-5S rDNA and 18S-(TTAGGG)n FISH demonstrated that repeated units of the three multicopy families are closely associated on the same chromosome pair.
Similar content being viewed by others
Introduction
Fluorescence in situ hybridization (FISH) is a powerful technique enabling the visualization of nucleic acid probes on target chromosomes, nuclei, cells and tissues. To date, this method has seldom been employed in the molluscan class Gastropoda. 18S-28S rDNA has previously been used as a probe to map major ribosomal clusters to the chromosomes of two neogastropod species, the Atlantic dogwhelk Nucella lapillus (Pascoe et al, 1996) and the Mediterranean tulip shell Fasciolaria lignaria (Vitturi et al, 2000a), whereas nothing is known about the 5S rDNA localization among gastropods.
Only two papers deal with chromosome FISH mapping of other multigene families in these organisms. The first reports on in situ hybridization with the (TTAGGG)n telomeric repeat to the chromosomes of Oxynoe olivacea (Opisthobranchia, Sacoglossa) (Vitturi et al, 2000b) and F. lignaria (Vitturi et al, 2000a), revealing the genomic occurrence of the hexanucleotide sequence only in the latter species. The second one shows that in F. lignaria (Vitturi et al, 2000a), the (GATA)n sequence is abundant and dispersed throughout the genome.
In the present paper we used single-colour FISH to spermatocyte chromosomes of the periwinkle Melarhaphe neritoides (Prosobranchia, Gastropoda, Caenogastropoda) to map repeated units of the two rDNA families (18S-28S rDNA and 5S rDNA) and to test the presence of (GATA)n and (TTAGGG)n repeats in the genome of this species. Available literature data report on the localization of major (18S-28S rDNA) and minor (5S rDNA) ribosomal clusters on different chromosome pairs as the most frequent configuration in vertebrates (see Lucchini et al, 1993; Suzuki et al, 1996). Since very little is known about rDNAs mapping in gastropods, we employed in addition to single colour FISH, simultaneous double-colour FISH to investigate about their physical relationship. Furthermore, a few works concerning the application of molecular DNA analysis were highly informative for the study of phylogeny and evolution in some gastropod genera (Ozawa and Okamoto, 1993; Collins et al, 1996; Reid et al, 1996) nevertheless, the molecular approach was never applied to chromosomes.
Mediterranean periwinkles are represented by three species, which according to the recent revision of Littorinidae (Reid, 1989), are Littorina saxatilis, Nodilittorina punctata and M. neritoides; the conventional chromosome analysis and the genome size of these species were previously reported (Vitturi et al, 1988, 1995). In particular, the M. neritoides male complement was found to consist of 33 chromosomes including 16 autosomal pairs (four mono-armed (pair nos. 4, 5, 6, and 8) and 12 bi-armed (all others)) and one unpaired metacentric X chromosome (16AA+X0).
Materials and methods
A total of 225 M. neritoides specimens were examined. The periwinkles were sampled during November and December 2000 from two geographically distinct areas: Addaura (Palermo, NW Sicily, S Italy; 180 specimens) and the Lagoon of Venice (NE Italy, 45 specimens) and identified according to the guidelines of Parenzan (1970). Most of the collected specimens were not useful for the present study because they were females and chromosome spreads were difficult to get from them, therefore data reported herein were obtained from about eighty sexually mature males. Moreover, since adult M. neritoides specimens are small in size, chromosome preparations were carried out from pooled testes of 6–8 individuals at a time, after overnight in vivo colchicine treatment (1 ng/L in sea water).
Single-colour FISH was performed on fixed spermatocyte chromosomes as described by Vitturi et al (2000b) using four different probes: (a) a sea urchin (Paracentrotus lividus) rDNA probe consisting of sequences of the 18S rDNA; (b) a PCR-obtained 5S rDNA probe using as primers F (5′-TGCACGTAGTGTTCCCAAGC) and R (5′-ACGACCATACCACGTTGAATAC) deduced from the 5S coding sequence of insects available in Genbank, according to the protocol described by Mandrioli et al (2000); (c) a PCR-obtained telomeric hexanucleotide (TTAGGG)n; and (d) a PCR-obtained (GATA)n sequence. Both telomeric and (GATA)n sequences were generated by PCR (PCR DIG-Probe Synthesis Kit: Roche) in the absence of a template (Ijdo et al, 1991) using (TTAGGG)5 and (CCCTAA)5 and (GATA)7 and (TATC)7 as primers, respectively. Nick translation labelling with digoxigenin of 18S rDNA was performed according to manufacturer's (Boehringer Mannheim) instructions, while the remaining three probes were DIG-labelled following the manufacturer's protocol (Roche).
For multiple fluorescent in situ hybridization the 18S rDNA probe was labelled with biotin (Biotin-Nick Translation Mix: Roche) whereas both 5S rDNA and (TTAGGG)n probes were the same as those employed in single-colour experiments. Chromosomes were denatured for 4 min in 70% formamide/2 × SSC at 72°C. The mixed probe solution (4 ng/μl of each probe) was denatured for 5 min at 80°C. Hybridization was allowed to proceed in a moist plastic chamber at 37°C overnight. Slides were washed twice in 50% formamide/2×SSC at 37–38°C (5 min each), twice in 2×SSC at 37–38°C (5 min each), once (5 min) in 4×SSC/0.1% Tween at room temperature (RT) and finally, once (5 min) in PBS/0.1% Tween/0.5% skimmed milk powder at RT. Probe hybridization sites were detected using both Anti-DIG-Fluorescein, Fab Fragments and Streptavidin Texas Red conjugate according to the manufacturers’ (Boehringer Mannheim, Molecular Probes) instructions. Slides were mounted in an antifade solution containing DAPI (3 μg/ml) and viewed under a Leica three-colour filter set (B/G/R) which allowed the simultaneous visualization of fluorescein- and Texas red-labelled hybridization sites (green and red, respectively) and chromosomal DNA (blue).
Chromosomes were observed with a Leica microscope and photographed with Kodak Ektacolor 1000 ASA film.
Results
With respect to all techniques here employed, specimens of both populations did not show any inter-populational differences. For this reason they are not further distinguished.
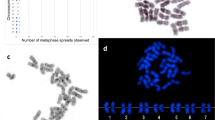
18S rDNA FISH on 15 spermatogonial metaphases of three different specimens always showed two small metacentric elements bearing a hybridization signal in the short arm (Figure 1a). Homology of these chromosomes was conclusively demonstrated by the presence of a single hybridized bivalent in 25 examined spreads at metaphase I (Figure 1b).
Figure 1 Spermatocyte chromosomes of Melarhaphe neritoides after rDNA FISH treatment with a heterologous 18S rDNA probe from the sea urchin Paracentrotus lividus: spermatogonial metaphase (a), diakinetic bivalents (b). (Arrows indicate hybridization signals).
Figure 2 Diakinetic bivalents after rDNA FISH treatment with a PCR obtained 5S rDNA probe. (Arrow indicates the hybridized bivalent.)
Figure 3 Pachytene chromosomes of Melarhaphe neritoides after FISH treatment with a PCR obtained (TTAGGG)n sequence.
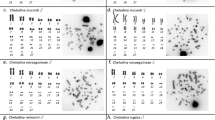
Figures 4 and 5 Diakinetic bivalents after FISH treatment with a PCR obtained (GATA)n sequence of Melarhaphe neritoides (4) and Milax nigricans (5).
Figure 6 Spermatogonial chromosomes of Melarhaphe neritoides after simultaneous double-colour FISH treatment with 18S and 5S rDNAs as probes. (Arrows indicate co-localization of hybridization signals.)
Figure 7 Diakinetic bivalents of Melarhaphe neritoides after simultaneous double-colour FISH treatment with 18S rDNA and telomeric (TTAGGG)n sequence as probes. (Arrow indicates the hybridized bivalent.)
Due to the scantiness of spermatogonial plates in chromosome preparations, FISH using 5S rDNA, (TTAGGG)n and (GATA)n probes was performed only on meiotic spreads. 5S rDNA FISH showed one hybridized bivalent per spread at metaphase I (Figure 2); the telomeric (TTAGGG)n repeat hybridized to the termini of each bivalent (Figure 3); the microsatellite did not show any labelled areas on diakinetic spreads (Figure 4). During the same experiment, the (GATA)n probe hybridized strongly to the chromosomes of F. lignaria (Vitturi et al, 2000a) chosen as a positive control, and to the slug Milax nigricans (Mollusca, Gastropoda, Pulmonata) (Figure 5).
Simultaneous 18S-5S rDNA FISH to the chromosomes of the same plate, allowed individuation of the two probes as partial distinct signals (red and green, respectively) because they were closely joined to each other (Figure 6).
Simultaneous in situ hybridization using 18S rDNA and (TTAGGG)n as probes showed that ribosomal (red spot) and telomeric (green spot) sequences were located adjacently at the terminal region of the rDNA-bearing bivalent (Figure 7).
Discussion
Major ribosomal genes of specimens of M. neritoides from the Lagoon of Venice were previously localised using silver staining. They were on the short arm of a variable number of chromosomes, most frequently three, of which two were small metacentrics and the third a large subtelocentric (Vitturi et al, 1995). In the present study, single-colour FISH with 18S rDNA demonstrates that the silver positive area previously observed in the large subtelocentric chromosome does not contain any ribosomal clusters, since NORs were constantly mapped to one small metacentric chromosome pair in all three examined periwinkles.
In the same manner as 18S, the 5S rDNA probe hybridized to a single metaphase I bivalent. Single-colour FISH, however, was unable to determine whether the hybridized bivalents corresponded in the two experiments.
By means of simultaneous double-colour FISH, it was possible to establish that 18S-28S rDNA and 5S rDNA were co-localized and, presumably, interspersed, due to overlapping of the two hybridization signals.
18S and 5S rDNA co-localization is rare in vertebrates (Liu and Fredga, 1999). Among invertebrates, the 18S-28S and 5S rDNAs were found in the same chromosome in the nematode Meloidogyne arenaria (Vahidi et al, 1991) and in seven out of 11 species of calanoid copepods so far analyzed.
In situ hybridization using the (GATA)n probe revealed a lack of evident labelling of the genome of M. neritoides, suggesting that this sequence might be absent in the periwinkle or, if present, is in a very low amount. Any technical shortcoming can be ruled out about this finding because, during the same experiment, the (GATA)n probe hybridized strongly to the chromosomes of F. lignaria (Vitturi et al, 2000a) chosen as a positive control, as well as to the chromosomes of the slug Milax nigricans of which FISH chromosome analysis was undertaken at the same time. These results show a heterogeneous distribution of the microsatellite (GATA)n among gastropods.
FISH using the (TTAGGG)n probe demonstrates that the hexanucleotide is present at the chromosomal ends of the periwinkle examined here. It is worth remarking that, among the Mollusca, this sequence has also been found in the neogastropod F. lignaria (Vitturi et al, 2000a) and in two species of the class Bivalvia (Guo and Allen, 1997; González-Tizón et al, 1998), whereas it is absent in another gastropod species, the sacoglossan Oxynoe olivacea (Opisthobranchia) (Vitturi et al, 2000b). Other telomeric sequences, considered to be derivatives of this, have been found, including, among others, the pentamere (TTAGG)n repeat in insects (Okazaki et al, 1993; Sahara et al, 1999), the (TTGGGG)n in ciliates (Liu and Fredga, 1999) and the Arabidopsis-type (TTTAGGG)n in plants (Görtner et al, 1998).
Double colour FISH has been performed to examine the relationship between repeated units of major rDNA and telomeric sequences. A close association between repeated units of the two multigene families could be observed. Since results of this study also demonstrated a co-localization between 18S-28S and 5S, it can be concluded that, in the periwinkle, three multicopy families are linked on the same chromosome pair.
Some authors (Dover, 1986; Liu and Fredga, 1999) claim that co-localization or interspersion of repeated units of different multigene families may be a selective advantage. In fact, an adjacent disposition of such genes might cause unequal crossing-over with consequent heteromorphism which would play an important role in the maintenance of a conserved and multiple array.
Data from the literature indicate that this assumption may be true only for some species. In fact, among vertebrates, an adjacent disposition between NORs and telomeres is reported to be an unusual finding (see Liu and Fredga, 1999) and a co-localization of 18S-28S and 5S rDNAs is not considered the rule, but rather the exception (Sola et al, 2000; Mandrioli et al, 2000). For example, in fish, a co-localization of repeated units of ribosomal DNAs has been described only in four out of 20 species so far analyzed (Pendás et al, 1994; Morán et al, 1996; Mandrioli et al, 2000).
Finally, among invertebrates, a high degree of rDNA polymorphism occurs (eg, Vitturi et al, 1999, 2000c), and terminal NORs are often observed. This implies that major rDNA clusters are physically associated to telomeric sequences. Among these organisms next to nothing or very little is known about the physical relationship between repeated units of the 18S-28S and 5S rDNA because, at the present time, this feature has seldom been investigated.
References
Collins, TM, Frazer, K, Palmer, AR, Vermeij, GJ, Brown, WM (1996). Evolutionary history of northern hemisphere Nucella (Gastropoda, Muricidae): molecular, morphological, ecological, and paleontological evidence. Evolution, 50: 2287–2304.
Dover, GA (1986). Linkage disequilibrium and molecular drive in the rDNA gene family. Genetics, 122: 249–252.
Gonzáles-Tizón, A, Martínez-Lage, A, Marias, L, Feire, R, Cornudella, L, Méndez, Y (1998). Cytogenetic characterization of Donax trunculus (Mollusca, Bivalvia). 13th International Chromosome Conference. Cytogenet Cell Genet Abstracts, 109.
Gortner, G, Nenno, M, Weising, K, Zink, D, Nagl, W, Kahl, G (1998). Chromosomal localization and distribution of simple sequence rooeats and the Arabidopsis-type telomere sequence in the genome of Cicer arietinum I. Chrom Res, 6: 97–104.
Guo, X, Allen, SK Jr (1997). Fluorescence in situ hybridization of vertebrate telomere sequence to chromosomes ends of the Pacific oyster, Crassostrea gigas Thunberg. J Shellfish Res, 16: 87–89.
Ijdo, JW, Wells, RA, Baldini, A, Reeders, ST (1991). Improved telomere detection using a telomere repeat probe (TTAGGG)n generated by PCR. Nucl Acids Res, 19: 4780
Liu, WS, Fredga, K (1999). Telomeric (TTAGGG)n sequences are associated with nucleolus organizer regions (NORs) in the wood lemming. Chrom Res, 7: 235–240.
Lucchini, S, Nardi, I, Barsacchi, G, Batistoni, R, Andronico, F (1993). Molecular cytogenetics of the ribosomal (18S + 28S and 5S) DNA loci in primitive and advanced urodele amphibians. Genome, 36: 762–773.
Mandrioli, M, Colomba, MS, Vitturi, R (2000). Chromosomal analysis of repeated DNAs in the rainbow wrasse Coris julis (Pisces, Labridae). Genetica, 108: 191–195.
Morán, P, Martínez, JL, García-Vázquez, E, Pendás, AM (1996). Sex chromosome linkage of 5S rDNA in rainbow trout (Oncorhynchus mykiss). Cytogenet Cell Genet, 75: 145–150.
Ozawa, R, Okamoto, K (1993). Integrated palaeontological and molecular phylogenetic approaches to the study of phylogeny: a case study of Umbonium (Gastropoda). Chikyu Month, 15: 589–595. (In Japanese).
Okazaki, S, Tsuchida, K, Maekawa, M, Ishikawa, M, Fujiwara, H (1993). Identification of a pentanucleotide telomeric sequence (TTAGG)n in the silkworm Bombix mori and in other insects. Mol Cell Biol, 13: 1424–1432.
Parenzan, P (1970). Carta d’identità delle conchiglie del Mediterraneo (Gasteropodi). Bios Taras (ed), vol 1, pp 161–215.
Pascoe, PL, Patton, SJ, Critcher, R, Dixon, DR (1996). Robertsonian polymorphism in the marine gastropod Nucella lapillus: advances in karyology using rDNA loci and NORs. Chromosoma, 104: 455–460.
Pendás, AM, Morán, P, Freije, JP, García-Vazquez, E (1994). Chromosomal mapping and nucleotide sequence of two tandem repeats of Atlantic salmon 5S rDNA. Cytogenet Cell Genet, 67: 31–36.
Reid, DG (1989). The comparative morphology, phylogeny and evolution of the gastropod family Littorinidae. Phil Trans R Soc Lond B, 324: 877–895.
Reid, DG (1996). Systematic and Evolution of Littorina, The Ray Society n. 164 The Dorset Press: Dorchester, UK.
Reid, DG, Rumbak, E, Thomas, RH (1996). DNA, morphology and fossils: phylogeny and evolutionary rates of the gastropod genus Littorina. Phil Trans R Soc Lond B, 351: 877–895.
Sahara, K, Marec, R, Traut, W (1999). TTAGG telomere repeats in chromosomes of some insects and other arthropods. Chrom Res, 7: 449–460.
Sola, L, De Innocentiis, S, Gornung, E, Papalia, S, Rossi, AR, Marino, G, De Marco, P, Cataudella, S (2000). Cytogenetic analaysis of Epinephelus marginatus (Pisces: Serranidae), with the chromosome localization of the 18S and 5S rRNA genes and of the (TTAGGG)n telomeric sequence. Mar Biol, 137: 47–51.
Suzuki, H, Sakurai, S, Matsuda, Y (1996). Rat rDNA spacer sequences and chromosomal assignement of the genes to the extreme terminal region of chromosome 19. Cytogenet Cell Genet, 72: 1–4.
Vahidi, H, Purac, A, Leblanc, JM, Honda, BM (1991). Characterization of potentially functional 5S rRNA-encoding genes within ribosomal DNA repeats of the nematode Meloidogyne arenaria. Gene, 108: 281–284.
Vitturi, R, Catalano, E, Macaluso, M, Zava, B (1988). The karyology of Littorina neritoides (Linnaeus, 1758) (Mollusca: Proso-branchia). Malacologia,, 29: 319–324.
Vitturi, R, Colomba, MS, Barbieri, R, Zunino, M (1999). Ribosomal DNA location in the scarab beetle Thorectes intermedius (Costa) (Coleoptera: Geotrupidae) using banding and fluorescent in situ hybridization. Chrom Res, 77: 255–260.
Vitturi, R, Colomba, MS, Gianguzza, P, Pirrone, AM (2000a). Chromosomal location of ribosomal DNA (rDNA), (GATA)n and (TTAGGG)n telomeric repeats in the neogastropod Fasciolaria lignaria (Mollusca: Prosobranchia). Genetica, 198: 253–257.
Vitturi, R, Gianguzza, P, Colomba, MS, Jensen, KR, Riggio, S (2000b). Cytogenetics in the sacoglossan Oxynoe olivacea (Mollusca: Opisthobranchia): karyotype, chromosome banding and fluorescent in situ hybridization. Mar Biol, 137: 577–582.
Vitturi, R, Libertini, A, Panozzo, M, Mezzapelle, G (1995). Karyotype analysis and genome size in three Mediterranean species of periwinkles (Prosobranchia: Mesogastropoda). Malacologia, 37: 123–132.
Vitturi, R, Ramella, L, Colomba, MS, Caputo, V, Sella, G (2000c). NOR regions of Polychaete worms of the genus Ophryotrocha studied by chromosome banding techniques and FISH. J Hered, 91: 18–23.
Acknowledgements
We gratefully acknowledge Dr R Barbieri for providing the sea urchin rDNA probe and Prof AM Pirrone for supplying telomeric sequence and labelling all probes. This work was supported by a MURST grant (ex 60%) to R Vitturi.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Colomba, M., Vitturi, R., Castriota, L. et al. FISH mapping of 18S-28S and 5S ribosomal DNA, (GATA)n and (TTAGGG)n telomeric repeats in the periwinkle Melarhaphe neritoides (Prosobranchia, Gastropoda, Caenogastropoda). Heredity 88, 381–384 (2002). https://doi.org/10.1038/sj.hdy.6800070
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800070
Keywords
This article is cited by
-
Evolutionary dynamics of 5S rDNA location in acridid grasshoppers and its relationship with H3 histone gene and 45S rDNA location
Genetica (2011)
-
Chromosomal mapping of repetitive DNAs in the beetle Dichotomius geminatus provides the first evidence for an association of 5S rRNA and histone H3 genes in insects, and repetitive DNA similarity between the B chromosome and A complement
Heredity (2010)
-
Chromosomal organization of simple sequence repeats in the Pacific oyster (Crassostrea gigas): (GGAT)4, (GT)7 and (TA)10 chromosome patterns
Journal of Genetics (2008)
-
Isolation and Mapping of Telomeric Pentanucleotide (TAACC) n Repeats of the Pacific Whiteleg Shrimp, Penaeus vannamei, Using Fluorescence In Situ Hybridization
Marine Biotechnology (2006)
-
Karyotype and Chromosomal Location of 18S–28S and 5S Ribosomal DNA in the Scallops Pecten maximus and Mimachlamys varia (Bivalvia: Pectinidae)
Genetica (2006)