Abstract
Nasopharyngeal carcinoma (NPC) occurs with high frequency in Southeast Asian populations. The high prevalence and familial clustering of NPC in these populations suggest that genetic factors may contribute to the increased cancer risk by affecting susceptibility. The aim of the present study was to map chromosomal loci linked to susceptibility genes predisposing for NPC. We carried out a genome-wide scan by multipoint affected-only allele-sharing methods in 15 Chinese NPC families with two to six affected members per family. The families were from the Guangdong province in the south of China, where the highest risk of NPC is documented. These samples were genotyped using 800 microsatellite markers covering all autosomal chromosomes with an average marker distance of 5 cM. Using multipoint linkage analysis, four loci (2q, 5p, 12p, and 18p) showed LOD scores above 1.5. After genotyping additional markers in these four regions, only one locus on 5p13 showed an increased LOD of 2.1. In further haplotype analysis, affected individuals in six families shared three marker haplotypes between D5S674 and D5S418. In conclusion, a region on 5p13 may harbor a susceptibility gene for NPC.
Similar content being viewed by others
Introduction
Nasopharyngeal carcinoma (NPC) is one of the most common malignancies in certain geographical regions with population of Mongoloid origin such as Southern China (42 cases/105 inhabitants/year) and Greenland (31 cases/105 inhabitants/year).1 It also shows an intermediate incidence in North Africa, while it is rare in Caucasian populations of European countries (eg 0.6 cases/105 inhabitants/year in Sweden).2 It is believed that environmental risk factors, including the EBV infection, cooperate with a susceptible genetic background in the high-risk populations. Thus, NPC provides an interesting model for the understanding of the interaction of a viral infection with other environmental and genetic factors in oncogenesis.
Epidemiological studies have shown that familial clustering of NPC in Southern China is significantly higher than that of low-risk areas. Approximately 5–10% of NPC patients have a familial history in this endemic region.3 The incidence of NPC is still high in the South Chinese population even after emigration from China. Offspring of admixture of southern-origin Chinese with non-Chinese groups showed an intermediate incidence.2 The high rate of familial clustering of this cancer strongly suggests that genetic components contribute to the high risk for this disease. Several candidate chromosomal regions for susceptibility have been studied.4, 5, 6 These studies, so far, have not arrived at consistent conclusions. This could be due to heterogeneity of susceptibility or modest, combined effects of several genes contributing to NPC development.
In this study, we used a genome-wide scan to search for susceptibility loci for NPC. To carry out the scan, fifteen multiplex Chinese families with two to six patients with defined NPC were recruited from the Guangdong province. The patients and their family members were genotyped with 800 markers. Our results suggest a linkage of NPC susceptibility to a region on 5p13.
Materials and methods
Families and patients
The families with at least two affected individuals with NPC were recruited from Guangdong province, China. The average age of the cases at diagnosis was 39 years old, which is younger than the average age of sporadic cases reported by the Cancer Center, Sun Yat-Sen University, being 46.6 years.3 Most of the families from Guangdong province were Cantonese-speaking. The patients were diagnosed having NPC based on clinical pathological examinations as well as clinical records by local doctors. Except for NPC, other cancers were also found in five of these families such as liver cancer, gastric cancer, and Hodgkin's lymphoma (HL). HL is also associated with EBV infection. Three to five milliliters of peripheral blood was collected from NPC family members with informed consent. Tumor paraffin blocks from some affected individuals were obtained for DNA extraction. Studies were approved by the Local Ethical Committee and the Karolinska Institutet (no. 00-302). DNA from 49 healthy donors was also used for estimation of marker-allele frequencies in this population.
Microsatellite marker genotyping
DNA was prepared from the total blood by phenol-based method and the purified DNA was directly used for linkage study. Genome-wide scan was performed at deCode Genetics Inc., Iceland, using fluorescently labeled primers with one of the three fluorescent dyes, FAM, HEX, and NED (PE Biosystems). A panel of 800 polymorphic microsatellite markers covering 22 autosomal chromosomes with an average marker interval of 5 cM was used in this study. The 800-microsatellite-marker set contained markers from the ABI Linkage Marker (version 2) in combination with 400 custom-made markers. The average heterozygosity of the markers selected for the study was 0.75.
All markers were extensively tested for robustness, ease of scoring, and efficiency in multiplex PCR. Marker positions were obtained from deCODE genetic map.7 PCR amplifications were set up, run, and pooled on Gilson Cyberlab robots. The reaction volume was 5 μl, and for each PCR, 20 ng of genomic DNA was amplified in the presence of 2 pmol each primer, 0.25 U AmpliTaq Gold, 0.2 mM dNTPs, and 2.5 mM MgCl2 (buffer was supplied by the manufacturer (Applera)). Cycling conditions were as follows: 95°C for 10 min, followed by 37 cycles of 94°C for 15 s, annealing at 55°C for 30 s, and extension at 72°C for 1 min. The PCR products were supplemented with the internal size standard, and the pools were separated and detected on ABI Prism 3700 sequencers by using Genescan (version 3.0) peak-calling software (Applera). Alleles were automatically called using DAC, an allele-calling program developed at deCode Genetics8 and the program DecodeGT was used to fractionate the called genotypes according to quality and to edit when necessary.9
The data were confirmed and elaborated by adding more markers to some regions, reducing the distance between them to less than 5 cM, and was also manually checked separately at deCode Genetics Inc., CMM and MTC of Karolinska Institutet.
Statistical methods for linkage analysis
The complete data set was checked for Mendelian inconsistencies by using the Pedcheck program (http://watson.hgen.pitt.edu/register). Inconsistencies were removed from analysis. We used multipoint, affected-only allele-sharing methods to assess the evidence for linkage. All results, including LOD and non-parametric linkage scores, were obtained using the program Allegro.10 We used the Spairs scoring function11, 12 and an exponential allele-sharing model13 to generate the relevant 1-df statistics. When combining the family scores to obtain an overall score, instead of weighting the families equally12 or weighting the affected pairs equally, we used a weighting scheme that is halfway between the two in the log scale; our family weights are the geometric means of the weights of the two schemes. To increase our confidence in the evidence for linkage to a particular region, additional microsatellite markers were genotyped to increase the information content on IBD sharing to >85% in that region. The marker order and positions used in linkage analysis are from our high-resolution genetic map.7 All reported locations are in Kosambi centimorgans. Allele frequencies of all markers were estimated from 49 population-matched healthy individuals. We used the GENEHUNTER-PLUS (version 1.2)13 for haplotype reconstruction.
Power simulations
To estimate power of the 15 families collected to detect linkage, we used the SLINK14 program to generate 100 replicates using two linked marker loci with an interval of 5 cM, assuming a dominant model with 85% penetrance, no phenocopies, θ=0.025, and the marker with four equal frequency allele (PIC 70%). The simulated genotype data were analyzed using the GENEHUNTER-PLUS program.13
Results
To map loci for NPC, we collected 15 multiplex families from Guangdong province, China, in the collection period from 1996 to 1999. Detailed information is shown in Table 1. The total number of affected individuals was 59 and DNA from 40 affected individuals was available for the scan study. Six of the families had more than three affected individuals. The distribution of affected individuals within families was compatible with dominant inheritance. The average onset age for patients with NPC was 39 years. We have also collected a number of third-generation individuals who have not reached onset age.
To estimate the power of 15 families to detect linkage, we simulated two linked markers based on the real pedigree structures using multipoint non-parametric linkage analysis. The 15 families collected from the Guangdong province had 46% power to find linkage with NPL of 3.0 (P<0.002) and 90% of power with NPL of 2 (P<0.02).
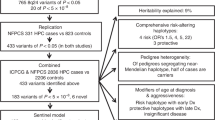
Using multipoint affected-only allele-sharing methods to assess the evidence for linkage, the LOD scores for the 22 autosomal chromosomes were first calculated from 15 pedigrees (Figure 1). Four chromosome regions showed an LOD score higher than 1.5. They were found in chromosome region 2q at D2S2382 (LOD=1.5), 5p at D5S2021 (LOD=2.1), 12p at D12S85 (LOD=1.5), and 18p at D18S1163 (LOD=2.0). No locus reached genome-wide significance. The LOD of 2.1 in this study almost meet genome-wide suggestive linkage (LOD 2.2). To analyze these potential susceptibility loci in more detail, we genotyped additional markers in these four regions and conducted the fine mapping in MTC and CMM, Karolinska Institutet. After genotyping of additional flanking markers, only the LOD peak at 5p region remained (Figure 2). For the 5p locus, we also examined the genotype data of the nine markers using parametric linkage analysis. Multipoint analysis showed a maximum HLOD score of 1.73 (α=0.08) at position of D5S2021, assuming a dominant model with 50% penetrance and 2% phenocopy. Using the same parameters, two-point analysis resulted in a LOD score of 1.9 (α=0) at the same position. The LOD scores at other chromosomal loci decreased to less than 0.5 (data not shown).
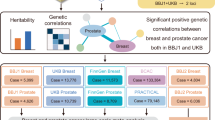
Through individual family evaluation, we observed six families contributing increased LOD. To narrow down the most likely location of this putative locus, we constructed haplotypes of the affected families between markers in the region 5p13.1 using minimum recombination by the GENEHUNTER-PLUS program. Six families from Guangdong province with at least three patients with cancer shared disease-associated haplotypes (Figure 3). The minimum-shared region shared by affected members in these families was located between markers D5S426 and D5S2021. The candidate region could be narrowed down to a segment of 3 Mb (Figure 3). This result suggested that an NPC susceptibility locus might be localized in this region. Further fine mapping is necessary using more families as well as sporadic trios, to verify and narrow down the region. To obtain further evidence supporting genetic heterogeneity in the six families, we checked clinical data as well as pathologic results in all the 15 families. No difference of pathologic changes was found between the six families and the remaining nine families. We also found no significant difference of mean age of diagnosis between the six families with haplotypes shared and the remaining nine families with haplotypes unshared.
Pedigrees and haplotypes of family F, G, K, T, W2, and W3. Haplotypes were inferred with a minimum recombination between markers by GENEHUNTER-PLUS. Filled symbols (black) represent affected individuals with NPC and filled symbols (gray) represent individuals with other forms of cancers. Boxes indicate the chromosome region shared by affected members of the pedigrees. The markers used for genotyping are listed in pedigree F.
Discussion
Genetic factors, which for example may influence the susceptibility of nasopharyngeal epithelium to EBV infection, exert a major impact on the disease risk of NPC. The relative risk of the first-degree relatives of the NPC patients is eight times in Greenland and 2.1 times in Cantonese population of that of the general population in the respective endemic regions, while it drops down as the degree of relationship decreases.15, 16 One characteristic of familial cancers is early age of onset, as observed in, for example, breast cancer.17 We also found a lower age of onset (39 years) in these families studied, compared to that reported for sporadic cases (46.6 years).3
Several studies have reported chromosomal regions that confer susceptibility to NPC, either using linkage analysis or association strategies. The HLA region on 6p21 is the most frequently reported region associated with NPC. Certain HLA haplotypes have already been found to be associated with increased NPC risk. These findings were further strengthened by linkage studies using affected sib pairs, suggesting a gene closely linked to the HLA locus at D6S1624 with an increased risk of NPC.4, 18, 19 However, so far no predisposing genes with causal mutations have been identified among families. In this study, no marker showing an LOD score higher than 0.5 was ever found in the HLA region, suggesting no linkage within this region. In one genome-wide scan of NPC, evidence of linkage to a 14-cM region on 4p15 with a maximum LOD score of 3.1 at D4S405 was reported.5 Surprisingly, we could not find evidence of a susceptibility locus in that region in the present study, although several NPC families originated from the same Guangdong area. This discrepancy may reflect genetic heterogeneity between subsets of families and/or that there are several susceptibility genes for NPC, even in the same high endemic area. Another possibility is that the linkage to chromosome 4p locus exists only in a few large families without susceptibility genes that occur in most other NPC families. In our scan study based on only 15 families, on the other hand, possibility of a false-negative LOD in the locus is hardly excluded.
Loss of tumor suppressor gene activity is associated with the pathogenesis of multistage carcinogenesis. High frequency of loss of heterozygosity was found in chromosome 3p in NPC. RASFF1A, FHIT, and BLUE genes in this region may be involved in pathogenesis of NPC and may be useful markers for diagnosis and prognosis of NPC.20, 21 It is worthwhile to identify susceptibility genes on pedigree-based and/or familial case-based analyses to NPC. Although a Chinese group reported a strong linkage to the chromosome 3 region,6 no region with significantly increased allele sharing within 3p was found in our data.
In this study, we showed that an NPC susceptibility locus might be localized to chromosome 5p. A marker D5S2021 on 5p13.1 yielded a maximum LOD score of 2.1, which almost reaches the genome-wide suggestive linkage of LOD score of 2.2.22 The haplotype analysis of our families indicated that an NPC susceptibility gene was likely to be located to the region between D5S1986 and D5S1969, which may narrow down the region to a 17.3-cM segment (DeCode map), or 16.6 cM (Genethon map), or 16.2 cM (Marshfield map). About 111 genes are present in this region according to a search in the human genome database, including 30 coding for hypothetical proteins, 16 similar or homologous to known genes/proteins, and 65 known genes such as zinc-finger RNA-binding protein, C1q, tumor necrosis factor-related protein-3, interleukin 7-receptor, oncostatin M–receptor, and caspase-recruitment domain family member 6.
In summary, a susceptibility locus on 5p13.1 may account for a subset of hereditary risk for NPC. Our results should encourage further studies to confirm the finding and identify the gene.
References
Ung A, Chen CJ, Levine PH et al: Familial and sporadic cases of nasopharyngeal carcinoma in Taiwan. Anticancer Res 1999; 19: 661–665.
McDermott AL, Dutt SN, Watkinson JC : The aetiology of nasopharyngeal carcinoma. Clin Otolaryngol Allied Sci 2001; 26: 82–92.
Zeng YX, Jia WH : Familial nasopharyngeal carcinoma. Semin Cancer Biol 2002; 12: 443–450.
Lu SJ, Day NE, Degos L et al: Linkage of a nasopharyngeal carcinoma susceptibility locus to the HLA region. Nature 1990; 346: 470–471.
Feng BJ, Huang W, Shugart YY et al: Genome-wide scan for familial nasopharyngeal carcinoma reveals evidence of linkage to chromosome 4. Nat Genet 2002; 31: 395–399.
Xiong W, Zeng ZY, Xia JH et al: A susceptibility locus at chromosome 3p21 linked to familial nasopharyngeal carcinoma. Cancer Res 2004; 64: 1972–1974.
Kong A, Gudbjartsson DF, Sainz J et al: A high-resolution recombination map of the human genome. Nat Genet 2002; 31: 241–247.
Fjalldal JB, Benediktsson K, Sigurdsson J, Ellingssen LM : Automated genotyping: combining neural networks and decision trees to perform robust allele calling. Proc Int Joint Conf Neural Networks 2001; pp A1–A6.
Palsson B, Palsson F, Perlin M, Gudbjartsson H, Stefansson K, Gulcher J : Using quality measures to facilitate allele calling in high-throughput genotyping. Genome Res 1999; 9: 1002–1012.
Gudbjartsson DF, Jonasson K, Frigge ML, Kong A : Allegro, a new computer program for multipoint linkage analysis. Nat Genet 2000; 25: 12–13.
Whittemore AS, Halpern J : A class of tests for linkage using affected pedigree members. Biometrics 1994; 50: 118–127.
Kruglyak L, Daly MJ, Reeve-Daly MP, Lander ES : Parametric and nonparametric linkage analysis: a unified multipoint approach. Am J Hum Genet 1996; 58: 1347–1363.
Kong A, Cox NJ : Allele-sharing models: LOD scores and accurate linkage tests. Am J Hum Genet 1997; 61: 1179–1188.
Ott J : Computer-simulation methods in human linkage analysis. Proc Natl Acad Sci USA 1989; 86: 4175–4178.
Friborg J, Wohlfahrt J, Koch A, Storm H, Olsen OR, Melbye M : Cancer susceptibility in nasopharyngeal carcinoma families – a population-based cohort study. Cancer Res 2005; 65: 8567–8572.
Jia WH, Feng BJ, Xu ZL et al: Familial risk and clustering of nasophar-yngeal carcinoma in Guangdong, China. Cancer 2004; 101: 363–369.
Hopper JL : Genetic epidemiology of female breast cancer. Semin Cancer Biol 2001; 11: 367–374.
Ooi EE, Ren EC, Chan SH : Association between microsatellites within the human MHC and nasopharyngeal carcinoma. Int J Cancer 1997; 74: 229–232.
Simons MJ, Wee GB, Day NE, Morris PJ, Shanmugaratnam K, De-The GB : Immunogenetic aspects of nasopharyngeal carcinoma: I. Differences in HL-A antigen profiles between patients and control groups. Int J Cancer 1974; 13: 122–134.
Hu LF, Eiriksdottir G, Lebedeva T et al: Loss of heterozygosity on chromosome arm 3p in nasopharyngeal carcinoma. Genes Chromosomes Cancer 1996; 17: 118–126.
Shao JY, Wang HY, Huang XM et al: Genome-wide allelotype analysis of sporadic primary nasopharyngeal carcinoma from southern China. Int J Oncol 2000; 17: 1267–1275.
Lander E, Kruglyak L : Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 1995; 11: 241–247.
Acknowledgements
This work has been supported by grants from the Swedish Cancer Society and Cancerföreningen in Stockholm. We are also grateful for the support by STINT, SIDA, and Karolinska Institutet. We are grateful to Johanna Skoglund and Tatjana Djureinovic, CMM, Karolinska Institute, for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hu, LF., Qiu, QH., Fu, SM. et al. A genome-wide scan suggests a susceptibility locus on 5p13 for nasopharyngeal carcinoma. Eur J Hum Genet 16, 343–349 (2008). https://doi.org/10.1038/sj.ejhg.5201951
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201951
Keywords
This article is cited by
-
Whole-Exome Sequencing of Nasopharyngeal Carcinoma Families Reveals Novel Variants Potentially Involved in Nasopharyngeal Carcinoma
Scientific Reports (2019)
-
Interplay of DNA methyltransferase 1 and EZH2 through inactivation of Stat3 contributes to β-elemene-inhibited growth of nasopharyngeal carcinoma cells
Scientific Reports (2017)
-
Prognostic impact of family history in southern Chinese patients with undifferentiated nasopharyngeal carcinoma
British Journal of Cancer (2013)
-
Gene discovery in familial cancer syndromes by exome sequencing: prospects for the elucidation of familial colorectal cancer type X
Modern Pathology (2012)
-
Characterization of Epstein-Barr virus type 1 nuclear antigen 3C sequence patterns of nasopharyngeal and gastric carcinomas in northern China
Archives of Virology (2012)