Abstract
S-adenosylhomocysteine hydrolase (AdoHcyase) catalyzes the hydrolysis of AdoHcy to adenosine and homocysteine. Increased levels of AdoHcy may play a role in the development of cardiovascular diseases and numerous other conditions associated with hyperhomocysteinemia. Several polymorphic isoforms named SAHH-1 to 4 may be resolved by horizontal starch gel electrophoresis from red blood cells. We have identified the genetic background of isoforms SAHH-2 and SAHH-3. SAHH-2 represents the previously described polymorphism in exon 2 of the AdoHcyase gene (112 C>T; p.R38W). Isoform SAHH-3 is based on a new polymorphism in exon 3 (377 G>A), leading to the conversion of glycine to arginine at amino-acid position 123. To shed light on the effects of these polymorphisms on the molecular and catalytic properties of AdoHcyase, we made recombinant wild-type and polymorphic R38W and G123R enzymes for a comparative analysis. The amino-acid exchanges did not bring about major changes to the catalytic rates of the recombinant proteins. However, circular dichroism analysis showed that both polymorphisms effect the thermal stability of the recombinant protein in vitro, reducing the unfolding temperature by approximately 2.6°C (R38W) and 1.5°C (G123R) compared to wild-type protein. In view of the altered thermal stability, and slightly decreased enzymatic activity of polymorphic proteins (≤6%), one may consider the analyzed AdoHcyase isoforms as risk markers for diseases caused by irregular AdoHcyase metabolism.
Similar content being viewed by others
Introduction
Mildly elevated homocysteine (Hcy) levels have been reported in multiple forms of vascular diseases,1, 2 neural tube defects, cognitive problems in elderly, abortion risk, presenile cataract and other pathologic conditions.3 Recently, there is increasing evidence that elevated S-adenosylhomocysteine (AdoHcy) may be a more important risk factor for mentioned conditions than Hcy itself.4
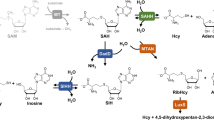
AdoHcy hydrolysis is the only source of Hcy in mammals. AdoHcy hydrolysis, catalyzed by AdoHcyase (EC 3.3.1.1), is a reversible reaction with an equilibrium favoring AdoHcy formation, but proceeds under physiological conditions in the hydrolytic direction due to the rapid removal of reaction products adenosine (Ado) and homocysteine.5
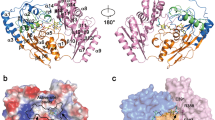
Human AdoHcyase is a tetramer consisting of chemically identical and functionally equivalent subunits, each with one molecule NAD tightly bound.6, 7 The 3-dimensional structure of human AdoHcyase has been resolved and mutational studies have allowed detailed insights into the processes important for catalytic activity.8, 9
Recently, it has been shown that missense mutations in the human AdoHcyase gene result in AdoHcyase deficiency.10 Main biochemical and clinical consequences are increased plasma creatine kinase, methionine, S-adenosylmethionine and AdoHcy, delayed brain myelination, myopathy and psychomotor retardation.10, 11, 12
The genetic frequencies of SAHH isoforms differ between populations. In the past, three polymorphic isoforms of AdoHcyase have been identified and only recently, a fourth variant has emerged.13
In this report, we elucidate the role of identified polymorphisms at amino-acid position R38 and G123 for enzymatic activity and holoenzyme formation of AdoHcyase. We used circular dichroism to determine structural aspects such as protein unfolding and stability during thermal induction. We determined the exact enzymatic capabilities of isoforms SAHH-2 (R38W) and 3 (G123R) to find clues whether aberrant AdoHcy metabolism might occur as a result of these polymorphisms.
Material and methods
Evaluation of AdoHcyase isoform 2, 3 and 4
Previously analyzed random blood samples from the hospital laboratory13 exhibiting the electrophoretic behavior for SAHH 2, 3 and 4 were tracked down to the particular individuals. None of them had neither clinical symptoms reported so far in AdoHcyase deficiency, nor elevated plasma AdoHcy. A sum of five SAHH-2, two SAHH-3 and four SAHH-4 samples was used for DNA extraction and sequence analysis as described previously.10
Cloning and overexpression of recombinant wild-type and polymorphic forms of AdoHcyase for expression in Escherichia coli
Expression vectors harboring the wild-type, R38W and G123R polymorphisms were constructed as described previously.14 In brief, single oligonucleotides (R38W: 5′-gagtaccactcccgcata-3′ and G123R: 5′-aggggcctgtccttgaagta) were used to introduce the desired amino-acid exchange into the AdoHcyase coding region. Both exchanges were confirmed by sequencing.
A detailed protocol for expression and purification of recombinant AdoHcyases in E. coli BL21 (DE3) RIL is given in Belužić et al.14
Enzymatic assays
AdoHcyase activity in purified enzyme preparations was assayed according to Takata et al15 by using 1–2 μg recombinant protein.
The synthetic activity of the purified AdoHcyase was determined by the rate of disappearance of adenosine according to Belužić et al.14
Quantitation of enzyme bound NAD+ and NADH was performed using a fluorescence technique described by Hohman et al16 using 200 μg of recombinant protein.
Polyacrylamide gel electrophoresis
SDS-polyacrylamide gel electrophoresis (PAGE) and determination of protein concentrations followed standard laboratory procedures. Additionally, purity and electrophoretic behavior of the recombinant AdoHcyase protein was analyzed using native-PAGE as described previously.14
Circular dichroism analysis
Circular dichroism (CD) measurements were performed on Jasco J-715 spectropolarimeter (Jasco Europe S.R.L, Cremella) as described previously.14 The protein concentrations used for measurements were 3.2 mg/ml for the wild-type, 0.57 mg/ml for the p.R38W and 0.5 mg/ml for the p.G123R protein.
Besides the CD analysis, we performed heat inactivation experiments with wild-type and p.R38W and p.G123R proteins according to Belužić et al.14
Bioinformatics
We used software applications nnPredict (Donald Kneller, University of California) for secondary structure prediction. Deepview/Swiss-PdbViewer17 was used for in silico mutation and 3D-structural analysis of the crystal structure of the human AdoHcyase (1A7A; Turner et al8). PeptideMass (http://www.expasy.ch/tools) was used for molecular weight prediction of recombinant AdoHcyase protein.
Results
Purification and characterization of recombinant R38W and G123R AdoHcyases
DNA sequencing confirmed a polymorphism in exon 2 of the AdoHcyase gene (112 C>T; p.R38W) for SAHH-2, and in exon 3 (377 G>A; p.G123R) for SAHH-3 samples, respectively. SAHH-4 samples did not reveal any polymorphism at all. Cloning, expression and purification of recombinant R38W and G123R enzyme was achieved as described previously.14 Approximately 7–10 mg of homogeneous enzymes were obtained from a 1-l culture that are indistinguishable from each other as judged by SDS-PAGE (Figure 1a).
Recombinant AdoHcyases have an increased molecular weight (63.9 kDa) due to the additional 152 amino-acid residues contributed by the plasmid expression vector tag region. Cleavage with thrombin reduced the size of the recombinant proteins by 130 residues (∼14 kDa), leaving only 22 additional amino acids fused to the AdoHcyase proteins. SDS-PAGE analysis of cleaved or uncleaved protein showed their mobility is in agreement with the predicted molecular weight (Figure 1a).
Firstly, the overall catalytic activity was determined in the directions of AdoHcy synthesis or hydrolysis. Table 1 summarizes the catalytic properties of wild-type and polymorphic enzymes. Both p.R38W and p.G123R enzymes retained a substantial amount of the overall catalytic activity in both directions (≥94% of the wild-type).
Further, quantitation of the NAD and NADH content in the recombinant p.R38W and p.G123R enzymes showed approximately 36% of the coenzyme bound to p.R38W as NADH, whereas pG123R shows a value of 50% NADH. These results are in accordance to previously published data from bovine liver AdoHcyase.18 The coenzyme content of recombinant AdoHcyases was determined to be 3.5 molecules NAD per tetramer for p.R38W and four molecules NAD per tetramer for p.G123R in agreement with previously published data.7
Further, native-PAGE analysis shows an electrophoretic mobility for the analyzed recombinant proteins in accordance to the mobility of AdoHcyase isoforms from red blood samples as resolved by starch gel electrophoresis.13 Thus, the p.R38W protein migrates slightly ahead of wild-type protein, whereas the p.G123R protein moves slowest (Figure 1b). The same behavior is observed for thrombin-digested AdoHcyase proteins, indicating that the fusion part of the recombinant protein does not interfere with holoenzyme formation (Figure 1b).
The far-UV CD spectra of wild-type, p.R38W and p.G123R proteins in water at room temperature were almost identical (not shown). A significant difference was observed in the thermal stability of the recombinant proteins. The p.R38W protein undergoes an unfolding process between 45.2 and 52.6°C with a Tm of 48.9±0.7°C, while unfolding of p.G123R is similar to wild-type, that is from 43.7 to 56.3°C with a Tm of 50.0±1.3°C.
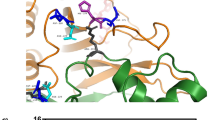
Results of 10-min heat-inactivation experiments showed a 77% loss of enzymatic activity for p.R38W at 50°C, whereas neither wild-type nor p.G123R protein were effected. p.R38W protein was fully inactivated above 53°C. Wild-type and p.G123R proteins retained approximately 10% activity even after incubation at 57°C. Analysis with the software package DeepView suggests that the R38W polymorphism results in the loss of a single hydrogen bond existing between R38 and Y39 (Figure 2a, b). The G123R polymorphism does not bring about changes to the H-bond network (not shown). Secondary structure prediction does not indicate changes in the regions containing polymorphic residues R38 or G123, respectively.
Discussion
Heterologous expression of wild-type and polymorphic human AdoHcyase cDNA yields functional holoenzymes with kinetic parameters very similar to those of AdoHcyases with other origins.19, 20, 21 AdoHcyase isoform 4 remains enigmatic as we have not been able to detect any polymorphism in the coding region of the gene. Therefore, SAHH-4 might occur as a result of post-translational modification of yet unknown character.
Reduction in the relative enzymatic activity is not observed for either polymorphic protein at physiological temperature of 37°C (data not shown). This indicates, that the hydrogen bond between R38 and Y39 is not essential for enzymatic activity, but plays a role for subunit folding and protein integrity at rising temperature (>40°C). This is confirmed by circular dichroism analysis, which showed a reduced unfolding temperature for the p.R38W enzyme compared to wild-type protein. Although the hydrogen-bond network in p.G123R protein is not disturbed, a slighly lower Tm of 1.5°C compared to wild-type indicates some sensitivity to thermal induction.
Under optimal laboratory conditions the analyzed polymorphisms seem to have no major effect on the enzymatic activity of AdoHcyase. However, our previous results show that significant differences exist between enzymatic activities of AdoHcyase measured in patients' tissues and under in vitro conditions.14 Similar results have been shown by Corbo et al,22 describing differences in the Km/Vmax ratios between AdoHcyase isoforms from red blood cells and, kinetic differences upon treatment with inhibitors. Therefore, slightly, but significantly decreased hydrolytic activity in vitro in combination with altered thermal stability, may result in greater differences in vivo and should be investigated further. Accordingly, we suggest both AdoHcyase isoforms 2 and 3 as possible risk markers for irregular AdoHcyase biochemistry. In conclusion, individuals with AdoHcyase isoforms SAHH-2, and SAHH-3 should be monitored for changes in their levels of plasma AdoHcy and other related compounds in an effort to assess the relative risk confered by the polymorphisms on aberrant AdoHcyase metabolism and numerous pathologic conditions previously associated with hyperhomocysteinemia.
References
Gellekink H, den Heijer M, Kluijtmans LAJ et al: Effect of genetic variation in the human S-adenosylhomocysteine hydrolase gene on total homocysteine concentrations and risk of recurrent venous thrombosis. Eur J Hum Genet 2004; 12: 942–948.
Castro R, Rivera I, Blom HJ et al: Homocysteine metabolism, hyperhomocysteinaemia and vascular disease: An overview. J Inherit Metab Dis 2006; 29: 3–20.
Prigge ST, Chiang PK : S-adenosylhomocysteine hydrolase. in Carmel R, Jacobsen DW (eds): Homocysteine in Health and Disease. USA: Cambridge University Press, 2001, pp 79–90.
Loscalzo J : Homocysteine trials – clear outcomes for complex reasons. N Engl J Med 2006; 354: 1629–1632.
Dela Haba G, Cantoni GL : The enzymatic synthesis of S-adenosyl-L-homocysteine from adenosine and homocysteine. J Biol Chem 1959; 234: 603–608.
Coulter-Karis DE, Hershfield MS : Sequence of full length cDNA for human S-adenosylhomocysteine hydrolase. Ann Hum Genet 1989; 53: 169–175.
Palmer JL, Abeles RH : Mechanism for enzymatic thioether formation: mechanism of action of S-adenosylhomocysteinase. J Biol Chem 1976; 251: 5817–5819.
Turner MA, Yuan CS, Borchardt RT et al: Structure determination of selenomethionyl S-adenosylhomocysteine hydrolase using data at a single wavelength. Nat Struct Biol 1998; 5: 369–376.
Yamada T, Takata Y, Komoto J et al: Catalytic mechanism of S-adenosylhomocysteine hydrolase: Roles of His 54, Asp130, Glu155, Lys185, and Aspl89. Int J Biochem Cell Biol 2002; 37: 2417–2435.
Barić I, Fumić K, Glenn B et al: S-adenosylhomocysteine hydrolase deficiency in a human: a genetic disorder of methionine metabolism. Proc Natl Acad Sci USA 2004; 101: 4234–4239.
Barić I, Ćuk M, Fumić K et al: S-Adenosylhomocysteine hydrolase deficiency: a second patient, the younger brother of the index patient, and outcomes during therapy. J Inh Metab Dis 2005; 28: 885–902.
Buist NRM, Glenn B, Vugrek O et al: S-adenosylhomocysteine hydrolase deficiency in a 26-year-old man. J Inh Metab Dis 2006; 29: 538–545.
Kloor D, Fumić K, Attig S et al: Studies of S-adenosylhomocysteine-hydrolase polymorphism in a Croatian population. J Hum Genet 2006; 51: 21–24.
Belužić R, Ćuk M, Pavkov T et al: A single mutation at tyrosine 143 of human S-adenosylhomocysteine hydrolase renders the enzyme thermosensitive and effects the oxidation state of bound co-factor NAD. Biochem J 2006; 400: 245–253.
Takata Y, Yamada T, Huang Y et al: Catalytic mechanism of S-adenosylhomocysteine hydrolase: site-directed mutagenesis of Asp-130, Lys-185, Asp-189, and Asn-190. J Biol Chem 2002; 277: 22670–22676.
Hohman RJ, Guitton MC, Veron M : Inactivation of S-adenosyl-L-homocysteine hydrolase by cAMP results from dissociation of enzyme-bound NAD+. Proc Natl Acad Sci USA 1985; 82: 4578–4581.
Guex N, Peitsch MC : SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 1997; 18: 2714–2723.
Matuszewska B, Borchardt RT : The role of nicotinamide adenine dinucleotide in the inhibition of bovine liver S-adenosylhomocysteine hydrolase by Neplanocin A. J Biol Chem 1987; 262: 265–268.
Gomi T, Date T, Ogawa H et al: Expression of rat liver S-adenosylhomocysteinase cDNA in Escherichia coli and mutagenesis at the putative NAD binding site. J Biol Chem 1989; 264: 16138–16142.
Ault-Riche DB, Yuan CS, Borchardt RT : A single mutation at lysine 426 of human placental S-adenosylhomocysteine hydrolase inactivates the enzyme. J Biol Chem 1994; 269: 31472–31478.
Abeles RH, Fish S, Lipinskas B : S-adenosylhomocysteinase: mechanism of inactivation by 2′-deoxyadenosine and interaction with other nucleosides. Biochemistry 1982; 21: 5557–5562.
Corbo RM, Ingianna R, Scacchi R, Bozzi A : Kinetic properties of the common electrophoretic variants of human S-adenosylhomocysteine hydrolase (AHCY): the effect of four nucleoside analogue inhibitors. Ann Hum Genet 1992; 56: 35–43.
Acknowledgements
This work was supported by Grants 0098086 (OV) and 0108016 (KF, MC, IB) of the Ministry of Science, Education and Sports from the Republic of Croatia and by the Austrian Science Fund (FWF) project P17885 (TP). We thank Drs Dorotea Dorčić and Danijela Petković for help in collecting the samples and information of studied individuals. We are extremely grateful to Vesna Musani for supervision of the sequencing core facility at the Ruđer Bošković Institute. We greatly appreciate the commitment of participants and their informed consent regarding this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fumić, K., Belužić, R., Ćuk, M. et al. Functional analysis of human S-adenosylhomocysteine hydrolase isoforms SAHH-2 and SAHH-3. Eur J Hum Genet 15, 347–351 (2007). https://doi.org/10.1038/sj.ejhg.5201757
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201757
Keywords
This article is cited by
-
Identification of enzymes and activity from two-dimensional gel electrophoresis
Nature Protocols (2007)