Abstract
Cytotoxic T lymphocyte-associated antigen 4 (CTLA-4) is important for downregulation of T-cell activation, and CTLA-4 gene polymorphisms have been implicated as risk factors for rheumatoid arthritis (RA). Previous studies of the association between the +49 polymorphism of the CTLA-4 gene in RA have provided conflicting results. In order to determine association of the CTLA-4 gene with RA in Chinese Han population, we used denaturing gradient gel electrophoresis (DGGE) to genotype polymorphisms of four SNPs (MH30, +49, CT60 and JO31) of the CTLA-4 gene in 326 RA patients and 250 healthy controls. Furthermore, meta-analysis of all available studies relating +49 polymorphism to the risk of RA was performed to confirm the disease association. Among the SNPs examined, the genotype frequencies of CTLA-4 +49 and CT60 in RA patients differed significantly from controls (P=0.028 and 0.007). In addition, the distribution of four haplotypes constructed by these two SNPs was significantly different between patients and controls (χ2=10.58, d.f. =3, P=0.014). The meta-analysis also revealed that in both European and Asian populations, the CLTA-4 +49 G allele was associated with the risk of RA. These results suggested that the CTLA-4 gene might be involved in the susceptibility to RA in the Chinese Han population and both +49 and CT60 of CTLA-4 gene might be the causal variants in RA disease.
Similar content being viewed by others
Introduction
Rheumatoid arthritis (RA) is a complex autoimmune disorder, characterized by a chronic T-cell response that has evaded normal control mechanisms.1 Therefore, the genes involved in the regulation of T-cell responses may be primary determinants of susceptibility to RA. CTLA-4 is a key negative regulator of T-cell activation2 and is considered a candidate gene for autoimmune diseases including RA.
It has been reported that CTLA-4 polymorphisms are associated with several T-cell mediated autoimmune diseases, such as Graves' disease, type 1 diabetes mellitus (T1D) and multiple sclerosis.3, 4, 5 However, the investigations of CTLA-4 and RA have yielded variable and inconsistent results, some indicating association,6, 7, 8, 9 while others have not.10, 11, 12 Thus, the relationship between the CTLA-4 polymorphism and clinical features of RA remains unclear.
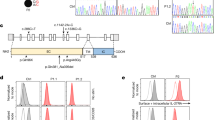
Most of the previous studies investigating the association of the CTLA-4 gene with RA were limited to the SNP +49, which located in CTLA-4 exon 1. The 6.1 kb in the 3′UTR and the 24 kb upward 5′ head of the CTLA-4 gene were reported to be significantly associated with Graves' disease (GD) in Caucasians.13 In order to determine the attribution of CTLA-4 gene polymorphisms for RA in the Chinese Han population, we investigated the polymorphisms of +49, CT60 and JO31. The latter two SNPs were in the head and the tail of the CTLA-4 3′UTR, respectively. Another SNP, MH30, which is near the CTLA-4 5′ head and reported to be highly associated with GD,13 was also genotyped (Figure 1).
Materials and methods
Patients
A total of 326 unrelated Chinese RA patients were recruited from the Outpatient Departments of Rheumatology in Guanghua Hospital, the Shanghai Second Medical University. They were composed of 91 men and 235 women, whose mean age was 48.4 (SD=±8.13). The diagnosis of RA was based on the 1987 American Rheumatism Association revised criteria for RA.14 The healthy control group consisted of 250 unrelated ethnically matched healthy subjects (73 men, 177 women, mean age 47.8 years, and SD=±11.05), who were randomly selected from community volunteers in the same district, free of any clinical evidence of autoimmune diseases or familial history of RA. The medical ethical community of Fudan University and Guanghua Hospital approved this study.
PCR-DGGE genotyping
Genomic DNA was extracted from peripheral blood by standard proteinase-K digestion and phenol/chloroform extraction and then amplified using specific DGGE primer sets15 (Table 1), which were designed to attain the optimal PCR-DGGE fragments. For optimal DGGE analysis, PCR was followed by a heteroduplex step, which involved denaturation at 96°C for 5 min and renaturation at 50°C for 1 h. DGGE was performed using the BIO-RAD Decode system. Optimized DGGE conditions were achieved according to the methods for the improvements of broad-range DGGE analysis.16 After electrophoresis, the gels were stained with ethidium bromide and photographed under an UV transilluminator. Amplified products with special DGGE banding patterns were purified and sequenced using a corresponding non-GC-clamped primer. The commonly occurring SNPs were recognized by DGGE pattern identification.
Statistical analysis
Genotype and allele frequencies were calculated on patients and control subjects by manual counting. Hardy–Weinberg equilibrium (HWE) was confirmed with the χ2 test.17 The χ2 test was also used to examine the statistical difference of the genotype or the haplotype distribution between patients and controls. Odds ratios (ORs) and 95% confidence intervals (CIs) for the association of allele/haplotype with the risk of RA were calculated using the STATA statistical package (Version 7.0). The PyPop program was used to estimate the haplotype frequency and for testing pairwise linkage disequilibrium (LD).18 Meta-analysis was performed using Review Manager software (version 4.2) (http://www.cc-ims.net/RevMan/). A P-value <0.05 was considered statistically significant.
Results
A total of four SNPs were successfully genotyped in 326 RA patients and 250 healthy controls. Table 2 shows the genotype distribution of these four SNPs. Genotype frequencies of these SNPs in both patients and healthy control subjects were in HWE. Among the four SNPs, the genotype and allele distributions of MH30 and JO31 did not differ significantly between RA patients and controls, while the genotype distribution of +49 and CT60 differed significantly between RA patients and healthy controls (χ2=7.18, d.f.=2, P=0.028; χ2=9.89, d.f. =2, P=0.007, respectively). And the +49 allele G and CT60 allele G were associated with an increased risk of RA (OR=1.36, 95% CI=1.06–1.74, P=0.012; OR=1.41, 95% CI=1.10–1.82, P=0.005, respectively) (Table 3).
We also analyzed the possible haplotypes constructed by +49 and CT60. The haplotype frequencies were estimated with an expectation-maximization (EM) algorithm of the HyPop program. This analysis revealed that there were four sets of haplotypes and two SNPs in healthy control subjects were in linkage disequilibrium (D′=0.6, P<0.0001). The haplotype +49 G: CT60 G and +49 A: CT60 A were more frequent than the haplotype +49 A: CT60 G and +49 G: CT60 A. Comparing cases with controls, the distribution of four haplotypes was significantly different (χ2=10.58, d.f.=3, P=0.014). Using the haplotype +49 A: CT60 A as reference, the frequencies of the haplotype +49 G: CT60 G and +49 A: CT60 G were significantly different between cases and controls (P=0.0015 and P=0.0425, respectively). The OR of the haplotype +49 G: CT60 G was 1.57 (95% CI=1.18–2.10), which was similar with that of the haplotype +49 A: CT60 G (OR=1.54, 95% CI=0.99–2.39) (Table 4).
For meta-analysis, we searched Medline database and the China National Knowledge Infrastructure (CNKI) database and checked the reference lists of the retrieved articles. In all, 12 case–control studies about association of +49 with RA were collected; six studies were on Europeans,6, 7, 9, 10, 19, 20 five were on Asians8, 11, 12, 21, 22 and other one was on Africans.23 (Table 5). For genotype frequencies of SNP +49 in both patients and healthy controls were in HWE, we investigated the association of CTLA-4 +49 allele G with the risk of RA in two race groups respectively. In both Europeans and Asians, there was no heterogeneity between studies (P=0.42, I2=0.6%; P=0.19, I2=33.2%, respectively). For the European group, the pooled OR was 1.11 (95% CI=1.00–1.23) for both fixed effects and random effects and to Asian group, the pooled OR was 1.18 (95% CI=1.04–1.35) for fixed effects and 1.16 (95% CI=0.98–1.37) for random effects. Funnel plot with 14 studies listing in Table 5 was drawn to test publication bias (Figure 2). None of small studies with statistically significant associations were found.
Discussion
CTLA-4 is an important costimulatory molecule, which can block IL-2 production and expression of IL-2 receptors and inhibit the function of cytotoxic T lymphocytes.24 It has been reported that blockade of the CTLA-4: B7 interaction exacerbates autoimmune diseases in an animal model of diabetes.25 Moreover, there is a native soluble form of CTLA-4 (sCLTA-4), encoded by an alternative transcript product of CTLA-4,26 and the level of sCLTA-4 expression is associated with autoimmune diseases, such as Autoimmune Thyroid Disease, Systemic Lupus Erythematosus, and Systemic Sclerosis.27, 28, 29 Our present study showed that CTLA-4 +49, CT60 and the haplotype constructed by these two SNPs had a susceptible effect to developing RA in Chinese Han population, while MH30 and JO31, which were reported to be most significantly associated with GD,13 had no such association.
The CTLA-4 +49 polymorphism has been genotyped extensively in several autoimmune diseases. In rheumatoid arthritis, Vaidya has found in Caucasian the +49 G allele was positively associated,9 and Lee et al,21 has also reported similar results in Gaoshan people. Our study showed that +49 G allele was a risk factor of RA in Chinese Han population, which was in accord with above reports in other populations. Since approximately 10–30% of RA patients have evidence of thyroid dysfunction and we had no further clinical data on these patients, we cannot rule out the possibility that the associations reported here might be due to thyroiditis, even though this possibility was very small. In order to confirm the disease association and to minimize the influence of study bias, we performed meta-analysis of all comparable genetic association study data sets, the importance of which has been recently advocated.30 To this end, we combined the data of European populations with 1400 RA patients and 1958 healthy controls and Asian populations with 1209 RA patients and 970 healthy controls, respectively. In the European group, the OR of +49 G allele was 1.11, which was similar to that in Asian group (OR=1.18). This result supported with Ioannidis's conclusion that the biological impact of genetic markers on the risk for common diseases may usually be consistent between different races.31 In European and Asian populations, the meta-analysis results suggested that CTLA-4 +49 allele G might be associated with RA. From the funnel plot, we found none of the small studies with statistically significant association. There might be two reasons, one is that studies published in languages other than English and Chinese, such as Korean and Japanese, could not be found; the other is that small studies with statistically significant association could not be published for small samples.
The CT60 G allele has recently been found to be highly associated with several autoimmune diseases, such as Graves' disease,13 celiac disease and systemic lupus erythematosus.32, 33 Our study demonstrated that CT60 G allele were positively associated with RA in Chinese Han population. This finding was at odds with a report by Barton et al.20 This might be due to the fact that in Barton's study, all patients and controls were Caucasians living in the UK, whereas we studied the Chinese Han population, and in different populations the same SNP might have different effects. Furthermore, our study also demonstrated that using the haplotype +49 A: CT60 A as reference, the OR of the haplotype +49 G: CT60 G was similar with that of the haplotype +49 A: CT60 G. And the haplotype +49 G: CT60 G (OR=1.57) had a little more effects than CT60 G allele (OR=1.41) or +49 G allele (OR=1.36) did. These results indicated that CT60 was the independent causal variant in RA disease and the effect of +49 G allele depended on CT60 G allele, and when +49 G allele combined with CT60 G allele, the effect of the haplotype +49 G: CT60 G increased. According to previous reports, the +49 G allele was considered to be related to reduced CTLA-4 expression and increased proliferation of autoreactive T cells,34 and the CT60 G allele was associated with producing lower mRNA levels of sCTLA-4,13 it was possible that the haplotype +49 G: CT60 G might alter production of CTLA-4 protein both in membrane and in soluble form, and might interfere with regulation of T-cell activation and proliferation leading to RA. This speculation requires further work to be confirmed at the cellular level. In addition, the functional meanings of these SNPs and haplotypes need to be elucidated with a suitable gene dose regulation model.
In conclusion, the CTLA-4 gene might be involved in the susceptibility to RA in Chinese Han population and both +49 and CT 60 of CTLA-4 might be the causal variants in RA disease.
References
Jör JG, Cornelia MW : T-cell regulation in rheumatoid arthritis. Curr Opin Rheumatol 2004; 16: 121–217.
Kremer JM, Westhovens R, Leon M et al: Treatment of rheumatoid arthritis by selective inhibition of T-cell activation with fusion protein CTLA4Ig. N Engl J Med 2003; 349: 1907–1915.
Kouki T, Sawai Y, Gardine CA et al: CTLA-4 polymorphisms at position 49 in exon 1 reduces the inhibitory function of CTLA-4 and contributes to the pathogenesis of Graves' disease. J Immunol 2000; 165z: 6606–6611.
Marron MP, Zeidler A, Raffel LJ et al: Genetic and physical mapping of a type-1 diabetes susceptibility gene (IDDM12) to a 100-kb phagemid artificial chromosome clone containing D2S72-CTLA4-D2S105 on chromosome 2q33. Diabetes 2000; 49: 492–499.
Kantarci OH, Hebrink DD, Achenbach SJ et al: CTLA-4 is associated with susceptibility to multiple sclerosis. J. Neuroimmunol 2003; 134: 133–141.
Seidl C, Donner H, Fischer B et al: CTLA4 codon 17 dimorphism in patients with rheumatoid arthritis. Tissue Antigens 1998; 51: 62–66.
Gonzalez-Escribano MF, Rodriguez R, Valenzuela A et al: CTLA-4 polymorphisms in Spanish patients with rheumatoid arthritis. Tissue Antigens 1999; 53: 296–300.
Yanagawa T, Gomi K, Nakao EI et al: CTLA-4 gene polymorphism in Japanese patients with rheumatoid arthritis. J Rheumatol 2000; 27: 2740–2742.
Vaidya B, Pearce SH, Charlton S et al: An association between the CTLA-4 exon 1 polymorphism and early rheumatoid arthritis with autoimmune endocrinopathies. Rheumatology (Oxford) 2002; 41: 1213.
Barton A, Myerscough A, John S et al: A single nucleotide polymorphism in exon 1 of cytotoxic T-lymphocyte-associated-4 (CTLA-4) is not associated with rheumatoid arthritis. Rheumatology (Oxford) 2000; 39: 63–66.
Matsushita M, Tsuchiya N, Shiota M et al: Lack of a strong association of CTLA-4 exon 1 polymorphism with the susceptibility to rheumatoid arthritis and systemic lupus erythematosus in Japanese: an association study using a novel variation screening method. Tissue Antigens 1999; 54: 578–584.
Lee YH, Choi SJ, Ji JD et al: No association of polymorphisms of the CTLA-4 exon 1(+49) and promoter (−318) genes with rheumatoid arthritis in the Korean population. Scand J Rheumatol 2002; 31: 266–270.
Ueda H, Joanna MM, Howson, Laura E et al: Association of the T-cell regulatory gene CTLA-4 with susceptibility to autoimmune disease. Nature 2003; 423: 506–511.
Arnett FC, Edworthy SM, Bloch DA : The American rheumatism association 1987 revised criteria for the classification of the rheumatoid arthritis. Arthritis Rheum 1988; 31: 315–324.
Ying Wu, Hayes VW, Jan Osinga et al: Improvement of fragment and primer selection for mutation detection by denaturing gradient gel electrophoresis. Nucleic acids Res 1998; 26: 5432–5440.
Vanessa MH, Ying W, Jan O et al: Improvements in gel composition and electrophoretic conditions for broad-range mutation analysis by denaturing gradient gel electrophoresis. Nucleic acids Res 1999; 27: e29.
Tang G, Xie H, Xu L, Hao Y, Lin D, Ren D : Genetic study of apolipoprotein E gene, alpha-1 antichymotrypsin gene in sporadic parkinson disease. Am J Med Genet 2002; 114: 446–449.
Alex L, Mark PN, Richard MS et al: PyPop: a software framework for population genomics: analyzing large-scale multi-locus genotype data; in Altman R B, et al (eds):: Pacific Symposium on Biocomputing. Singapore: World Scientific, 2003, pp. 514–525.
Milicic A, Brown MA, Wordsworth BP : Polymorphism in codon 17 of the CTLA-4 gene (+49 A/G) is not associated with susceptibility to rheumatoid arthritis in British Caucasians. Tissue Antigens 2001; 58: 50–54.
Barton A, Jury F, Eyre S et al: Haplotype analysis in simples families and novel analytic approaches in a case-control cohort reveal no evidence of association of the CTLA-4 gene with rheumatoid arthritis. Arthritis Rheum 2004; 50: 748–752.
Lee CS, Lee YJ, Liu HF et al: Association of CTLA-4 gene A-G polymorphism with rheumatoid arthritis in Chinese. Clin Rheumatol 2003; 22: 221–224.
Liu MF, Wang CR, Chen PC, Lin TL : CTLA-4 gene polymorphism in promoter and exon-1 regions is not associated with Chinese patients with rheumatoid arthritis. Clin Rheumatol 2004; 23: 180–181.
Hadj KH, Kaddour N, Adyel FZ et al: HLA-DQB1 CAR1/CAR2, TNFa IR2/IR4 and CTLA-4 polymorphisms in Tunisian patients with rheumatoid arthritis and Sjgren's syndrome. Rheumatology 2001; 40: 1370–1374.
Krummel MF, Allison JP : CTLA-4 engagement inhibits IL-2 accumulation and cell cycle progression upon activation of resting T cells. J Exp Med 1996; 183: 2533–2540.
Lenschow DJ, Ho SC, Sattar H et al: Differential effects of anti-B7-1 and anti-B7-2mAb treatment on the development of diabetes in the NOD mouse. J Exp Med 1995a; 181: 1145–1155.
Oaks MK, Hallett KM, Penwell RT et al: A native soluble form of CTLA-4. Cell Immunol 2000; 201: 144–153.
Oaks MK, Hallett KM : Cutting edge: a soluble form of CTLA-4 in patients with autoimmune thyroid disease. J Immunol 2000; 164: 5015–5018.
Sato S, Fujimoto M, Hasegawa M et al: Serum soluble CTLA-4 levels are increased in diffuse cutaneous systemic sclerosis. Rheumatology 2004; 43: 1261–1266.
Liu MF, Wang CR, Chen PC, Fung LL : Increased expression of soluble cytotoxic T-lymphocyte-associated antigen-4 molecule in patients with systemic lupus erythematosus. J Immunol 2003; 57: 568–572.
Becker N, Nieters A, Rittgen W : Single nucleotide polymorphism–disease relationships: statistical issues for the performance of association studies. Mutat Res 2003; 525: 11–18.
John PAI, Evangelia EN, Thomas AT : Racial. differences in genetic effects for complex diseases. Nat Genet 2004; 36: 1312–1318.
Van Belzen MJ, Mulder CJ, Zhernakova A et al: CTLA4 +49 A/G and CT60 polymorphisms in Dutch coeliac disease patients. Eur J Hum Genet 2004; 12: 782–785.
Torres B, Aguilar F, Franco E et al: Association of the CT60 marker of the CTLA4 gene with systemic lupus erythematosus. Arthritis Rheum 2004; 50: 2211–2215.
Maurer M, Loserth S, Kolb-Maurer A et al: Polymorphism in the human cytotoxic T-lymphocyte antigen 4 (CTLA4) gene (exon 1 +49) alters T-cell activation. Immunogenetics 2002; 54: 1–8.
Acknowledgements
We would like to say great thanks to all subjects participating in this study, especially Dr Feng Zhang, Dr Yungang He and the well-trained doctors from Guanghua Hospital who helped us in sample collection. Fudan University and the St Luke's Medical Center of Aurora Health Care, WI, USA supported this study, and additional funds were donated by Changhai Hospital, Shanghai, China.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lei, C., Dongqing, Z., Yeqing, S. et al. Association of the CTLA-4 gene with rheumatoid arthritis in Chinese Han population. Eur J Hum Genet 13, 823–828 (2005). https://doi.org/10.1038/sj.ejhg.5201423
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201423
Keywords
This article is cited by
-
The yin and yang of co-inhibitory receptors: toward anti-tumor immunity without autoimmunity
Cell Research (2020)
-
CTLA-4 Expression Inversely Correlates with Kidney Function and Serum Immunoglobulin Concentration in Patients with Primary Glomerulonephritides
Archivum Immunologiae et Therapiae Experimentalis (2019)
-
Cytotoxic T- Lymphocyte Antigen-4 (CTLA4) Gene Expression and Urinary CTLA4 Levels in Idiopathic Nephrotic Syndrome
The Indian Journal of Pediatrics (2019)
-
The synergic effects of CTLA-4/Foxp3-related genotypes and chromosomal aberrations on the risk of recurrent spontaneous abortion among a Chinese Han population
Journal of Human Genetics (2018)
-
T‐cell exhaustion: understanding the interface of chronic viral and autoinflammatory diseases
Immunology & Cell Biology (2016)