Abstract
According to classical markers, France has been reported to be regionally heterogeneous. Here, we propose to test the homogeneity of the French mitochondrial gene pool by analysing D-Loop and coding regions polymorphisms in 210 individuals stemming from five regions. The data set obtained was also used to test the ability of mitochondrial DNA to detect well historically established admixtures (admixtures between British/Irish people and native Breton people in our case). For these purposes, the sampling procedure was subject to special care, concerning the individuals' geographical origin and maternal pedigree. The mtDNA analysis revealed some regional specificities in haplogroup distribution, which is discussed in terms of successive settlements of France. Statistical analyses were conducted to investigate mtDNA diversity and structure within and between British, Irish and French groups. They tended to show affinities between Morbihan region and Britain plus Ireland. Furthermore, genetic evidences were in line with the fact that Morbihan region results from an admixture event, agreeing with historical evidences of successive migrations from Britain and Ireland into Brittany. These results also tended to outline the fact that two geographically very adjacent samples (Morbihan and Finistère), sharing a cultural and linguistic area, can present a distinct genetic pattern. Although mtDNA analyses were able to identify a historically reported admixture event, we point out here the high influence of the sampling procedure and representativeness over the migrations hypothesis. We also underline the importance of regional sampling for studies on the spread and/or origin of specific European haplogroups (here U5a1a and U8).
Similar content being viewed by others
Introduction
The mitochondrial DNA has revealed to be a useful tool for studying the human settlement of different European regions,1 as well as for studying population migrations inside the subcontinent.2 Nevertheless, the successive Europe settlements have often been deduced from imprecise samples, usually collected from geographically broad regions, without taking into account regional diversities. This is particularly the case for the French regions that our paper deals with. The French territory is centrally located in Western Europe: particularly, it links Northern Europe to the Mediterranean and Iberian areas. Certainly, this peculiar geographic localisation had a strong impact on the human settlement processes as well as on the genetic structure of settled human populations. Nevertheless, the French mitochondrial gene pool remains poorly described and has almost solely been explored with a forensic point of view.3,4 In this way, although previous studies concerned relatively numerous individuals (n=185 in all), those were not regionally localised, and were only surveyed for polymorphisms in their D-Loop. Also, diagnostic RFLPs from coding regions were neglected, albeit crucial for haplogroup discussion. These lacking elements make these published data difficult to be used for further studies of the human settlements of France. Moreover, this lacking information could represent a potential distorting factor in research on the settlement of Europe, especially considering that studies of blood genetic markers have revealed a great inter-regional variability of French populations,5 and complex migratory Prehistory and History.6 The results of these studies outlined the importance of precise geographical information for samples analysed through genetic studies, in order to map the migratory patterns precisely. In order to point out the importance of sampling strategies information in human settlement of France, we report mitochondrial data that can be more properly used in this prospect. Features of the sampling procedure allowed us (i) to test the inter-regional genetic homogeneity of the French mitochondrial gene pool and its implication on potential ancient population movements; (ii) to test the ability of mtDNA in detecting admixture events that are unquestionably established by History and previously detected by classical markers data analyses.6
Material and method
Subjects and sampling procedure
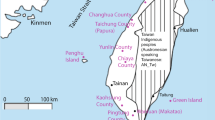
Our sample consisted of 210 maternally unrelated volunteers, originating from five French locations (Figure 1). The sampled area in each region was about 2000 km2. Most of the selected subjects were born in the first part of the 20th century, and pedigree investigations were conducted over the last three (for about 50% of the subjects) or more generations to confirm the regional maternal origin of sampled individuals. These considerations should minimise genetic disruptions related to population movements inside the French territory, movements that became important on and after the 19th century (but that can be neglected for earlier periods).7 Moreover, because big towns and coastal belts are known to have been more implicated in population movements than the rural ones, we verified that the oldest known ancestor of selected subjects originated from rural and noncoastal parts of the region. Then, disruptions related to potential gene flows, anterior to the oldest known ancestor birth, should also be minimised. In order to test the impact of historical migrations on the mtDNA pool of Brittany, we included previously published European data (see Figure 1).
Localisation of the analysed samples. Our samples include individuals from the central ‘department’ of Var (VR, n=37), the region of Périgord-Limousin (PL, n=72) straddling three western-central ‘departments’ (northern Dordogne, western Corrèze and southern Haute-Vienne), the regions of Caux and Bray located in Normandy (CB, n=39) and the ‘departments’ of Morbihan (BM, n=40) and Finistère (BF, n=22), both located in Brittany. Previously published data were also included in the analyses, and they concern 139 individuals from England2 (ENG), 70 individuals from Wales (WA), 50 individuals from Cornwall8 (CNW), 101 individuals from western Ireland9 (IRL), 47 individuals from north-eastern France9 (NEF) and 185 French individuals who remain regionally unlocalised3,4,9 (FRA).
mtDNA extraction and genotyping
DNA was extracted from peripheral blood mononuclear cells or hair roots as described elsewhere.10Hypervariable segments (HVS-1 and HVS-2) of the control region were simultaneously amplified by PCR using either L16025 and HV2AS,10 or L15832 (light chain, nps 15838–15858) and HV2AS. Amplifications were performed with standard PCR conditions, except that dTTPs were replaced by dUTPs in the PCR reaction mix (as in forthcoming coding regions amplifications) in order to avoid potential contamination from earlier amplifications (especially in routine PCR laboratory). Indeed, dUTPs allow the use of uracile-N-glucosidase (UNG) that destroys potential PCR products arising from earlier amplifications (by cutting DNA with uracile). The purification of PCR products and sequencing conditions followed are those described in Mogentale-Profizi et al.10 The light strand of HVS-1 and HVS-2 was sequenced, permitting the achievement of HVS-1 and HVS-2 sequences, respectively, between at least nucleotide 16055 and 16410 (precision in Supplemental Material), and between nucleotide 00037 and 00222. Since the traces were of excellent quality and unambiguous, the obtention of only one strand was sufficient. The typing of coding regions polymorphisms was performed as in Mogentale-Profizi et al10 except six additional relevant polymorph sites that were typed by sequencing: nps 4216 (for haplogroup J), 4646 (for U4), 10238 (for N1b), 11719 (for pre-HV), 15904 (for pre-V and V), 15907 (for U2), according to Richards et al9 Moreover, we report here a new method to check 4580 and 12308 positions state, based on allele-specific PCR (primers and conditions reported in Table 1). Our method was tested on samples previously typed by enzymatic restriction and sequencing, and the results were strictly reliable between the three methods. It was strictly reproducible, cheaper than sequencing and faster than enzymatic restriction method; moreover, it does not depend on specific endonucleases. Thus, this method could be extended advantageously to other diagnostic RFLP sites.
As recommended,12 polymorphisms appearing incongruent were double-checked by amplification, sequencing and/or enzymatic restriction. In this way, we amplified and sequenced HVS-1 and HVS-2 twice in both senses in the case of individuals included in haplogroup H and not presenting 00073 G to A transition, or included in haplogroup U with a 00073 A transition. In these cases, the state of sites 7028 and 12308 was also double-checked. Also, deletions and/or insertions, and haplotypes with unusual polymorphism patterns were double-checked.
Statistical analysis
The DNA sequences alignments and haplotype identification were conducted as in Mogentale-Profizi et al10using CLUSTAL X 1.8113 and MEGA 2.0.14 To maximise the number of British populations in comparison with French data, statistics and analysis were conducted on HVS-1 sequence between sites 16090 and 16365. Statistical analysis focused on historical movements that led to the constitution of the present-day Breton mtDNA gene pool. The population genetic structure of the potentially implicated regions was analysed through AMOVA15 using Arlequin 2.000 package.16 Taking into account the historical framework, AMOVA analyses were applied to different – but plausible – population groupings. Gene diversity (H) and nucleotide diversity (πn)17 were estimated. As samples from Britain and Ireland perceptibly were less diverse than French ones, these statistics were used in a bivariate analysis in order to detect a potential effect on Breton diversities. We also report the estimated mean number of pairwise differences (MNP).18
Results
Molecular data
The mtDNA polymorphisms found in the D-Loop analysis of the French samples, along with the status of the samples at nps 16519 and at the typed diagnostic restriction sites, are reported in Supplemental Material. Seven individuals were found to be heteroplasmic. Six of them present one heteroplasmy at nps 16093. This observation is the somatic manifestation of the particularly fast mutation rate of this site, previously demonstrated in human pedigrees studies and evolutionary analyses.19,20
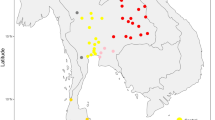
The haplogroup frequencies observed in each sample are summarised in Table 2. Most of them fall into the variability of European populations (see Simoni et al21) and cannot differentiate the studied samples (although appearing quite heterogenous). However, peculiar frequency distributions across our French sample can be noted. First, haplogroup K frequency appears higher in Morbihan (17.5%) and in Périgord-Limousin (15.3%) than the highest observed European frequency (13.3% in Norwegians and Bulgarians, the average of European K frequency being 5.6% – 21 populations have been taken into account; data from Simoni et al21). Second, haplogroup V appears particularly infrequent, and specially absent from Périgord-Limousin. Third, haplogroup U8 (as defined in Finnilä et al22) is solely present in the sample from Var, and encompasses three different haplotypes in this locality. It is interesting to note that haplogroup U8 is rare in Europe, and absent from Northern Caucasus and Near East.9 Moreover, it has rarely been observed with associated polymorphisms (only three Finnish individuals presenting the same transition at the hypervariable site 16093 were described),22 while 2/3 haplotypes belonging to haplogroup U8 presented associated polymorphisms in Var. Fourth, in north-western France (Normandy and Morbihan), we typed three haplotypes being phylogenetic intermediates between two clusters differentiated by two substitutions: cluster U5a1 and its phylogenetically related subcluster U5a1a (Figure 2). For convenience, we thus grouped these three haplotypes as an intermediate cluster: U5a1a#. And fifth, Périgord-Limousin also presents a high frequency (about 15%) of individuals belonging to haplogroups [T1, J, and (pre-HV)1] considered as introduced in Europe during the Neolithic.1 For comparison, the sample from Var, located in the Mediterranean basin, presents a lower frequency of this type of haplogroups (2.7%; Fisher's exact test: P=0.04)
Median-joining network23 of French Haplogroup U5a1. Two Norman haplotypes (CB26 and CB33) and one haplotype from Morbihan (BM39) present the derived state at nps 16399, but lack the reversion at nps 16192. Although position 16192 is known to exhibit a high substitution rate, the occurrence of three mutations at this site, on the same lineage and over 24 300 years (the lower limit of the Confidence Interval of the estimated age of haplogroup U5a1)9 is unlikely. They thus have been grouped into haplogroup U5a1a#.
Statistical analysis
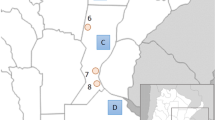
AMOVA analyses (Table 3) showed a genetic homogeneity between French, British and Irish populations. However, it has to be noted that combination D (Table 3) produces the highest variation among groups (0.46%) and the lowest variation within groups (0.26%), associated with the strongest significance (respectively, P=0.00489 and 0.00098). Gene diversity, nucleotide diversity and MNP of the studied samples are reported in Table 4. Correlation tests demonstrated that the sample sizes have no influence either on the nucleotide diversity or on the gene diversity (data not shown). On the other hand, we could demonstrate a strong correlation between both diversities (R2Pearson=0.7586, P=0.001; Figure 3). The sample from Morbihan exhibits an intermediate position between samples from the British/Irish Isles and samples from the continent, especially according to its gene diversity. Moreover, its nucleotide diversity appears relatively higher compared with its gene diversity.
Bivariate analysis of H and πn (abbreviations as in Figure 1).
Discussion
Although our French samples could not be statistically differentiated by their haplogroup frequencies, each one harbours some qualitative and quantitative peculiarities that may reflect a rather different history. Albeit located in a region considered to have been largely affected during Neolithic,24 the sample from Var exhibits a pattern that does not appear to have been genetically influenced by potential ‘Neolithic waves’ from the Near East. This observation could be the consequence of human demographical and ecological peculiarities of the Mediterranean coast at the end of the Neolithic period, as proposed by Simoni et al21 or could also reflect a sexual-specific pattern of migration (in favour of men) for this period, as suggested by a study by Semino et al.25 On the contrary, the Périgord-Limousin sample exhibits a quite large Neolithic component (15%), estimated from the frequency of Neolithic-specific haplogroups.1 Then, according to Richards et al9 all European regions experienced a large immigration at the end of the Upper Palaeolithic. In western France (represented here by samples from Brittany and Périgord-Limousin), the high observed frequency of haplogroup K, introduced in Europe from the Near and/or Middle East, suggests that the majority of Late Upper Palaeolithic immigrants belonged to this haplogroup. Finally, two previous and successive studies26,27 detected a re-expansion of populations from the south-western Europe to north-eastern Europe after the last glacial maximum (LGM). Albeit the postglacial re-expansion undoubtedly concerned French regions because of their immediate proximity, haplogroup V appeared to be very rare in our samples and absent from Périgord-Limousin (although this region is included in the northern part of the refugium formed during the LGM). Thus, we postulate that the traces of the re-expansion have been at least partially erased by posterior migrations, on and/or after the Mesolithic period. The relatively great Neolithic component (see above) in this region could partially explain the erasure of immediate postglacial pattern.
Brittany was affected by well-established historical migrations from Britain and Ireland. Earliest massive historical migrations from Britain occurred during the fourth century, involving about 40 000 individuals. Those migrations essentially concerned soldiers, women being largely in minority.28 A second migration wave occurred during the sixth and seventh centuries, involving this time a real settlement from Britain in Brittany, the immigrants becoming numerous inside native Continentals.28 The principal immigration that followed involved Irish people (during the War of Ireland, 1641–1651), bringing small Irish groups (about 35 000 persons in all) fleet to Lower-Brittany29 (present-day Finistère, most of Morbihan territory and western Côtes d'Armor). Marriage, birth and death certificates from this period indicate integration of these Irish immigrants with the native people.29 All together, historical elements show that the Breton gene pool has undoubtedly been affected by British and Irish genes. This case appeared opportune for us to test some methods that could enable the detection of such an admixture event. As inferring admixture proportions and processes from single-locus analysis does not seem feasible for only slightly differentiated human populations,30 indirect methods, which could be less dependent on the differentiation degree of parental populations, should be proposed in such a homogenous context. The results of the bivariate analysis of H and πn are compatible with the historical data of successive British and Irish migrations into Brittany. Indeed, the position of the sample from Morbihan with regard to the regression line is what can be expected from an admixed population (Figure 3): the gene diversity of population resulting from the admixture event should average gene diversities of parental populations, whereas the nucleotide diversity should tend to increase. Indeed, because the probability that two separated populations (even since a relatively short time) generated by mutation and/or fixed by drift the same alleles is low, the gene pool combination of these populations will result in a higher nucleotide diversity compared to the previous separated populations ones. Here, the nucleotide diversity of Morbihan is higher than the ones measured in insular populations and a nonlocalised French sample, and is quite high with regard to the gene diversity of the entire sample. Indeed, successive migrations of British/Irish populations (usually less diverse than the French ones, see Figure 3 and Table 4) appear to have reduced the gene diversity of the indigenous French mitochondrial gene pool in Morbihan. Moreover, AMOVA analyses showed that Morbihan is more similar to British/Irish populations than French ones (see combination D, Table 3), supporting the historically reported migrations. Considering results from the bivariate and AMOVA analyses, this Breton sample appears to stem from an admixture event. Also, if one considers that some migrations from the Britain and Ireland involved only (military) men,28,31 the detected signal (through mitochondrial diversity) would solely be an under-estimation of the real flows impact. In contrast, the Breton sample from Finistère presents a different position from the Morbihan sample. Finistère is located at the extremity of the distribution of the bivariate analysis. Neither AMOVA nor the bivariate analysis results permit to show that this population stems from the admixture between British/Irish and French populations. This observation moderates the impact of immigrations from Britain and Ireland on the mitochondrial gene pool of Brittany. These migrations should have affected the entire Lower-Brittany mtDNA pool; nonetheless, our results tend to show that mtDNA from Finistère was less, until not, affected by the historical migrations considered. Then, these results underline the importance of the sampling when micro-geographical problems are inquired. Indeed, if only people from Finistère were sampled, or if Brittany was considered as a whole and that Bretons were sampled without care of their precise regional origins, historically well-documented immigrations in Brittany would not have been detected.
Our French study has also permitted to suggest information on the spread and/or origin of specific European haplogroups. This is the case of haplogroup U5a1a, for which the regional origin remains unclear, and that is reported to have differentiated very recently (between 2200 and 12 800 BP).9 The fact that three different haplotypes belonging to U5a1a# were found in Normandy and Morbihan suggests that this haplogroup appeared in north-western Europe: it would have subsequently spread across Europe from this centre. One part of U5a1a would have reached the Near East, and certainly subsequently back-migrated into Europe, as suggested by Richards et al9 Then, haplogroup U8 was found principally in Alpine and north-eastern European regions.9,22 Although it was previously described as diversity-free in most cases, U8 exhibits diversity in our Var sample. The age estimate of this cluster is 44 400±27 010 years BP (calculated as in Saillard et al32 with Network 3.0 – http://www.fluxus-engineering.com, including all the 12 haplotypes previously described,9,22 and the three haplotypes from Var), and clearly shows that this cluster is of Palaeolithic origin. Haplogroup U8 has most probably differentiated on the west central Mediterranean coast during the Upper Palaeolithic, and has subsequently migrated into north-eastern Europe (possibly via Alpine region), during the population re-expansions that occurred after the LGM. Thus, this cluster suggests that the French Mediterranean coast acted as a refugium during the LGM, and indicates that, as well as haplogroup V, some other clusters have differentiated in Western Europe during the Upper Palaeolithic and could be taken into account to evaluate the impact of the post-LGM expansion on the present-day European gene pool.
Our results showed that the quality of the sampling procedure can act on the qualitative and quantitative results obtained from the analysis of mtDNA data. Dealing with recent migratory events, we could suggest that a sample better reflects the regional history when it is well localised geographically, and if it does take into account the individuals maternal ancestry. In this way, we can conclude that the previous nonlocalised French sample, used to represent the French population, is not suitable to reconstruct population flows into French territory. Owing to its heterogeneous migratory history, data from various regional samples are necessary to elucidate successive settlements of the French territory, and more generally, settlements of Europe. We can also conclude that precise regional sampling can permit the detection of dispersed or very localised haplotypes, and can help to better understand the phylogeographical history of some clusters (here U5a1a and U8). All together, these elements are essential for understanding the history of the European mitochondrial gene pool. Finally, studies dealing with other DNA markers would enrich and precise the picture given by mtDNA, allowing to access to paternal parameters (with Y-chromosome polymorphisms) and to the resultant of the interaction of paternal and maternal gene pools (with autosomic markers). For instance, concerning Var, it would be interesting to test whether, contrary to mtDNA, the analysis of Y-chromosome polymorphisms would permit the detection of a Neolithic impact on the French Mediterranean coast.
References
Richards M, Macaulay V, Torroni A, Bandelt HJ : In search of geographical patterns in European mitochondrial DNA. Am J Hum Genet 2002; 71: 1171–1174.
Helgason A, Hickey E, Goodacre S et al: mtDNA and the islands of the North Atlantic: estimating the proportions of Norse and Gaelic ancestry. Am J Hum Genet 2001; 68: 723–737.
Rousselet F, Mangin P : Mitochondrial DNA polymorphisms: a study of 50 French Caucasian individuals and application to forensic casework. Int J Legal Med 1998; 111: 292–298.
Cali F, Le Roux MG, D'Anna R et al: mtDNA control region and RFLP data for Sicily and France. Int J Legal Med 2001; 114: 229–231.
Ohayon E, Cambon-Thomsen A (eds) Génétique Des Populations Humaines. Paris: INSERM, vol 142, 1986.
Cavalli-Sforza LL, Menozzi P, Piazza A : The History and Geography of Human Genes. Princeton: Princeton University Press; 1994.
Biraben JN : Démographie historique et génétique. in Ohayon E, Cambon-Thomsen A (eds) Génétique Des Populations Humaines. Paris: INSERM; 1986, vol 142, pp 29–40.
Richards M, Côrte-Real H, Forster P et al: Paleolithic and Neolithic lineages in the European mitochondrial gene Pool. Am J Hum Genet 1996; 59: 185–203.
Richards M, Macaulay V, Hickey E et al: Tracing European founder lineages in the near eastern mtDNA pool. Am J Hum Genet 2000; 67: 1251–1276.
Mogentale-Profizi N, Chollet L, Stévanovitch A et al: Mitochondrial DNA sequence diversity in two groups of Italian Veneto speakers from Veneto. Ann Hum Genet 2001; 65: 153–166.
Anderson S, Bankier AT, Barell BG et al: Sequence and organization of the human mitochondrial genome. Nature 1981; 290: 457–465.
Bandelt HJ, Lahermo P, Richards M, Macaulay V : Detecting errors in mtDNA data by phylogenetic analysis. Int J Legal Med 2001; 115: 64–69.
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG : The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 1997; 24: 4876–4882.
Kumar S, Tamura K, Jakobsen IB, Nei M : MEGA2: molecular evolutionary genetics analysis software. Bioinformatics 2001; 17: 1244–1245.
Excoffier L, Smouse P, Quattro J : Analysis of molecular variance from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 1992; 131: 479–491.
Schneider S, Roessli D, Excoffier L : Arlequin Ver. 2.000: A Software for Population Genetics Data Analysis. Switzerland: Genetics and Biometry Laboratory, University of Geneva; 2000.
Nei M : Molecular Evolutionary Genetics. New York: Columbia University Press; 1987.
Tajima F : Evolutionary relationship of DNA sequences in finite populations. Genetics 1983; 105: 437–460.
Forster L, Forster P, Lutz-Bonengel S, Willkomm H, Brinkmann B : Natural radioactivity and human mitochondrial DNA mutations. Proc Natl Acad Sci USA 2002; 99: 13950–13954.
Parsons TJ, Muniec DS, Sullivan K et al: A high observed substitution rate in the human mitochondrial DNA control region. Nat Genet 1997; 15: 363–368.
Simoni L, Calafell F, Pettener D, Bertranpetit J, Barbujani G : Reconstruction of prehistory on the basis of genetic data. Am J Hum Genet 2000; 66: 1177–1179.
Finnilä S, Lehtonen MS, Majamaa K : Phylogenetic network for European mtDNA. Am J Hum Genet 2001; 68: 1475–1484.
Bandelt HJ, Forster P, Rölh A : Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 1999; 16: 37–48.
Vaquer J (ed) Le Néolithique du NORD-Ouest Méditerranéen. Paris: Société Préhistorique Française; 1999.
Semino O, Passarino G, Oefner P et al: The genetic legacy of Paleolithic Homo sapiens sapiens in extant Europeans: a Y chromosome perspective. Science 2000; 290: 1155–1159.
Torroni A, Bandelt HJ, Macaulay V et al: A signal, from human mtDNA, of postglacial recolonization in Europe. Am J Hum Genet 2001; 69: 844–852.
Torroni A, Bandelt HJ, D'Urbano L et al: mtDNA analysis reveals a major late Paleolithic population expansion from southwestern to northeastern Europe. Am J Hum Genet 1998; 64: 1137–1152.
Monnier JJ : L'immigration bretonne en Armorique. in Monnier JJ, Cassard JC (eds) Toute L'histoire de Bretagne. Morlaix: Skol Vreizh; 1997, pp 97–106.
Le Noac'h A : Les immigrés irlandais en centre Bretagne au XVIIe siècle. Actes XVe Congres Natl Généal, Brest 1999; 176–181.
Dupanloup I, Bertorelle G : Inferring admixture proportions from molecular data: extension to any number of parental populations. Mol Biol Evol 2001; 18: 672–675.
Rouche M : Les mélanges de population en Gaule durant le Haut Moyen-Age. in: Ohayon E, Cambon-Thomsen A (eds) Génétique des Populations Humaines. Paris: INSERM; 1986, vol 142, pp 19–28.
Saillard J, Forster P, Lynnerup N, Bandelt HJ, Norby S : mtDNA variation among Greenland Eskimos: the edge of the Beringian expansion. Am J Hum Genet 2000; 67: 718–726.
Acknowledgements
Many thanks to André Gilles, Marie-France Deguilloux, and Alexandre Vienne for helpful, stimulating discussions and critical readings on this manuscript, and to three anonymous referees for their helpful comments. We are grateful to all volunteers who contributed their DNA to the research, and to Pascal Pouedras, Paulette Besson, Colette Barra, Andrée Dubut, Paul Clergerie and Christiane Bigot for their help with the sample collection. We express our gratitude to Vincent Macaulay, who kindly provided us previously published Irish and French HVS-1 sequences. This work was partially supported by QIAGEN France and ROCHE France.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg).
Supplementary information
Rights and permissions
About this article
Cite this article
Dubut, V., Chollet, L., Murail, P. et al. mtDNA polymorphisms in five French groups: importance of regional sampling. Eur J Hum Genet 12, 293–300 (2004). https://doi.org/10.1038/sj.ejhg.5201145
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201145
Keywords
This article is cited by
-
Soldiers fallen in the Battle of Austerlitz
Archaeological and Anthropological Sciences (2021)
-
Coevolution of genes and languages and high levels of population structure among the highland populations of Daghestan
Journal of Human Genetics (2016)
-
Analysis of ancient human mitochondrial DNA from the Xiaohe cemetery: insights into prehistoric population movements in the Tarim Basin, China
BMC Genetics (2015)
-
Fine-scale human genetic structure in Western France
European Journal of Human Genetics (2015)
-
Mitochondrial diversity patterns and the Magdalenian resettlement of Europe: new insights from the edge of the Franco-Cantabrian refuge
Journal of Human Genetics (2012)