Abstract
We report on a novel localization for a recessive form of deafness (DFNB), by linkage analysis in an Iranian consanguineous family. Affected individuals suffer from prelingual profound sensorineural hearing loss. Genome-wide analysis led to the characterization of a new locus, DFNB40, which maps to an ∼9 Mb interval between markers D22S427 and D22S1144 at chromosome 22q11.21–12.1. Maximum lod score of 3.09 was obtained with D22S1174. Since the Bronx waltzer (bv) mouse mutant, characterized by waltzing behavior, deafness, and degeneration of cochlear inner hair cells, has been mapped to the syntenic region on murine chromosome 5, we suggest that DFNB40 and bv may result from orthologous gene defects.
Similar content being viewed by others
Introduction
Congenital deafness affects more than one in 1000 children at birth1 or during early childhood, with a majority of cases (nearly 70%) having no associated features (nonsyndromic deafness). Most cases of hereditary hearing impairment are inherited on the autosomal recessive mode (DFNB). To date, 33 different DFNB loci and 20 of the corresponding genes have been identified2 (http://www.uia.ac.be/dnalab/hhh/). Here we report on a novel DFNB locus, DFNB40, at chromosome 22q11.21–12.1 in a family from the region of Isfahan. This is the first deafness locus to be identified in the hearing-impaired population in Iran.
Patients and methods
Informed consent was obtained from adult subjects and from the parents of under-aged patients.
Auditory tests
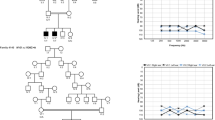
Audiometry testing was performed on all the family members. Air and bone-conduction were evaluated (with a Beltone 2000 clinical audiometer) at frequencies of 250, 500, 1000, 2000, 4000, and 8000 Hz (pure-tone audiometry). Air conduction pure-tone average (ACPTA) threshold in the conversational frequencies (0.5, 1, and 2 kHz) was calculated for each ear, and the hearing loss was classified as mild (20 dB⩽ACPTA⩽40 dB), moderate (40 dB
Genotyping
Genomic DNAs were prepared from 5 to 10 ml of blood by standard techniques. The genome-wide screening was performed with 400 microsatellite markers spaced 10–12 cM apart (ABI Prism Linkage mapping set 2, Applied Biosystems, Courtaboeuf, France).
Linkage analysis
Lod scores were calculated using the MAPMAKER/HOMOZ program package (version 0.9), a computer package based on an algorithm especially designed for homozygosity mapping.3 Deafness was assumed to be inherited in a fully penetrant autosomal recessive mode with a disease allele frequency of 10−3; changing it to 10−2 only slightly modified the lodscore value. Recombination frequencies were considered to be identical in males and females. For each polymorphic marker, allele frequencies were supposed to be equal. Finally, the equivalence between genetic and physical distances was considered to be 1 Mb=2.3 cM in the chromosomal region limited by markers D22S420 and D22S1144, located ∼10 Mb and 23 cM apart.
Results and discussion
Clinical description
In family 914-IS, three individuals (IV-1, IV-4, IV-6), born from a marriage between two first cousins (III-1 and III-2), were affected by prelingual, bilateral, and profound sensorineural deafness (Figure 1). The ages of generation IV individuals ranged from 12 to 28 years. A complete medical history was obtained for each affected individual to exclude the possibility of infectious or environmental causes of hearing impairment. No other clinical sign was associated to deafness. Audiometric tests in both parents were normal. Therefore, a recessive form of isolated hearing loss is segregating in this family.
Linkage analysis
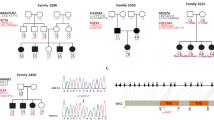
A genome-wide linkage analysis using polymorphic microsatellite markers revealed an ∼9 Mb critical region of homozygosity between markers D22S427 and D22S1144 (Figure 2). Maximum lod score of 3.09 was obtained with D22S1174, thereby defining a new deafness locus, DFNB40, at chromosome 22q11.21–12.1. We also tested 73 Iranian consanguineous families affected by recessive deafness, for which the DFNB1 locus had been excluded, and did not find any that was linked to DFNB40 (data not shown). Whereas several DFNB forms have been demonstrated to be allelic to dominant forms (DFNA) of deafness,2 no DFNA locus has been reported in the DFNB40 chromosomal region. The DFNB40 interval contains more than 80 genes. Also, this interval overlaps with the intervals for syndromes that include deafness, namely cat eye syndrome (MIM115470) and velocardiofacial/DiGeorge (MIM192430/MIM188400) syndrome. As several deafness genes have been found to underlie both isolated and syndromic deafness,2 we searched for mutations in the TBX1 transcription factor gene, which is involved in velocardiofacial syndrome4 and has been proposed, based on the phenotype of transgenic mice, to be responsible for the hearing impairment in some of the patients.5 No mutation in the TBX1 coding exons was detected in family 914-IS. We also searched for mutations in two additional candidate genes within the DFNB40 interval, namely DGCR6 (DiGeorge syndrome critical region gene 6) that is expressed in the inner ear6 and CRYBB1 (β crystallin B1 gene), since another crystallin gene has been shown to underlie a dominant form of deafness.6 No mutation was detected in the coding regions of these genes. Identification of additional families with a recessive hearing impairment mapping to the DFNB40 interval will be useful to refine the critical region. Finally, it is noteworthy that the Bronx waltzer (bv) mouse mutant, characterized by waltzing behavior, deafness, and degeneration of the cochlear inner hair cells, has been mapped in the region syntenic to the DFNB40 interval, on murine chromosome 5 (∼63 cM from the centromere).7
Segregation analysis with chromosome 22q11–q12 polymorphic microsatellite markers. Deaf individuals are indicated by filled symbols and unaffected individuals by open symbols. The ‘at risk’ haplotypes inherited from generation III are boxed in all generation IV individuals. The smallest region of homozygosity associated with the mutated DFNB40 allele is boxed on the left side of the figure. For each polymorphic marker, the physical distance to the centromere is indicated on the left, and the calculated lod score is indicated on the right. The DFNB40 critical interval, between D22S427 and D22S1144, spans 8.8 Mb.
References
Thompson D, McPhillips H, Davis R, Lieu T, Homer C, Helfand M : Universal newborn hearing screening: summary of evidence. JAMA 2001; 286: 2000–2010.
Petit C, Levilliers J, Hardelin J-P : Molecular genetics of hearing loss. Annu Rev Genet 2001; 35: 589–646.
Kruglyak L, Daly MJ, Lander ES : Rapid multipoint linkage analysis of recessive traits in nuclear families, including homozygosity mapping. Am J Hum Genet 1995; 56: 519–527.
Merscher S, Funke B, Epstein J et al: TBX1 is responsible for cardiovascular defects in velo-cardio-facial/DiGeorge syndrome. Cell 2001; 104: 619–629.
Funke B, Epstein J, Kochilas L et al: Mice overexpressing genes from the 22q11 region deleted in velo-cardio-facial syndrome/DiGeorge syndrome have middle and inner ear defects. Hum Mol Genet 2001; 10: 2549–2556.
Abe S, Katagiri T, Saito-Hisaminato A et al: Identification of CRYM as a candidate responsible for nonsyndromic deafness, through cDNA microarray analysis of human cochlear and vestibular tissues. Am J Hum Genet 2003; 72: 73–82.
Bussoli TJ, Kelly A, Steel KP : Localization of the bronx waltzer (bv) deafness gene to mouse chromosome 5. Mamm Genome 1997; 8: 714–717.
Acknowledgements
The authors thank the family members for their participation in the study, Mehdi Ashtiani for his support, Francoise Denoyelle, Jacqueline Levilliers and Jean-Pierre Hardelin for their help in the preparation of the manuscript, and Sebastien Chardenoux for genetic linkage calculations. This work was supported by grants from Fondation pour la Recherche Medicale (ARS 2000) and European Community (QLG2-CT-1999-00988). SD is a recipient from the Pasteur Institute (Paris, Teheran).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Delmaghani, S., Aghaie, A., Compain-Nouaille, S. et al. DFNB40, a recessive form of sensorineural hearing loss, maps to chromosome 22q11.21–12.1. Eur J Hum Genet 11, 816–818 (2003). https://doi.org/10.1038/sj.ejhg.5201045
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201045