Abstract
Hearing impairment is the most frequent sensory defect in children, with a genetic basis in about 50% of cases. Several point mutations and deletions in mitochondrial DNA (mtDNA) have been identified in non-syndromic sensorineural hearing loss (NSSNHL). Beside the frequent A1555G mutation, a number of mutations in tRNAs have been reported recently, but their incidence remains unknown. We identified the T7511C mutation in the tRNASer(UCN) gene in two French families with isolated deafness. Maternal transmission was obvious in both. The 15 patients with hearing impairment exhibited a variable disease phenotype in terms of onset, severity, and progression. T7511C was present in all the patients screened. Homoplasmic and heteroplasmic levels were observed and did not correlate with the severity of the disease. T7511C was also present in 12 hearing offspring of the oldest deaf mothers, confirming the existence of modulatory factors. Our data suggest that this mtDNA mutation should be screened for in all cases of familial NSSNHL compatible with maternal transmission.
Similar content being viewed by others
Introduction
Deafness is a relatively common disorder, with 1 in 1000 children presenting a severe or profound sensorineural deafness at birth. A genetic defect is thought to account for up to 60% of cases of congenital deafness in developed countries. Remarkable progress in identifying genes involved in deafness has been made over the past few years. So far, 60 loci involved in NSSNHL have been reported and 27 genes cloned.1 Furthermore, heteroplasmic mtDNA mutations have been found in syndromic hearing loss associated with systemic neuromuscular syndromes,2 diabetes3 or skin diseases.4
In 1992, Jaber first described an Arab-Israeli family affected with a NSSNHL which demonstrated mitochondrial inheritance.5 In 1993, a A1555G mutation in the mitochondrial 12S ribosomal RNA gene was isolated in this pedigree.6 This mutation is thought to predispose individuals to aminoglycoside-induced hearing loss. Until now, more than 120 families carrying this mutation have been described throughout the world.7,8 In these families, the penetrance and the expression of the hearing loss varied considerably among individuals, suggesting that other factors are involved. Other mtDNA mutations located in the tRNASer (UCN) gene were demonstrated to be responsible for NSSHI.9 The T7511C point mutation was identified in a large African American family.10 The occurrence of mtDNA mutations prompted us to perform screening for the known mtDNA mutations in French families with maternally inherited NSSNHL. We report here two separate families found to have a T7511C hetero or homoplasmic mtDNA mutation. We discuss the variability of the phenotype and recommend systematic screening of mtDNA mutations in maternally inherited NSSNHL or in cases due to aminoglycoside administration.
Materials and methods
Mutation analysis
DNA was extracted from white blood cells by standard techniques. DNA was also extracted from urine, hair root cells and mouthwash samples when these samples were available.
Denaturing gradient gel electrophoresis (DGGE)
MtDNA mutation screening was performed by denaturing gradient gel electrophoresis (DGGE) of tRNA Ser(UCN) and tRNA Leu(UUR), following previously described techniques.11 In addition, 12S rRNA surrounding A1555G was analysed by DGGE. MtDNA was amplified by PCR with primers corresponding to nt positions 1440–1465 (forward) with an additional GC tail at the 5′ end (backward) and 1592–1617 (reverse). PCR consisted of 35 step cycles of 94°C for 30 s, 55°C for 30 s and 72°C for 30 s.
Electrophoresis was performed in a 6% polyacrylamide gel with a 10–60% gradient (60°C, 145 V, 4 h). Each PCR product was analysed ‘pure’ and mixed with PCR-amplified DNA of a healthy control individual who had the reference sequence in order to detect homoplasmic mutations.
Direct sequencing of PCR amplified fragments
PCR products generated for DGGE analysis were subjected to direct sequencing in an ABI Prism 310 Genetic Analyser (Perkin-Elmer Applied Biosystems, Foster City, CA, USA) using the Big Dye Terminator Cycle sequencing reaction kit (Perkin-Elmer Applied Biosystems). We used the updated Cambridge human mitochondrial DNA light strand sequence as the reference sequence.
PCR/restriction fragment length polymorphism analysis
MtDNA surrounding tRNA Ser(UCN) was amplified by PCR with the same primers and in the same conditions as these used for DGGE analysis. PCR product was digested with MboII. The mutant mtDNA creates a novel MboII restriction site. The percentage of mutant mtDNA was studied using the method described by Sue.10 Briefly, after 30 cycles of PCR, 3 μCu of α-32P was added to each reaction tube. The PCR products were digested by MboII and analysed by acrylamide electrophoresis. Quantification of the mutant mtDNA was performed in a phosphoimager (Bio-Rad).
Clinical and audiological evaluation
We recruited patients from the genetic counselling clinic for deaf people at The Armand-Trousseau Children's Hospital, Paris, France. The protocol was accepted by the Committee for the Protection of Individuals in Biochemical Research as required by French legislation and informed consent was obtained from all patients.
The proband of both families was clinically examined by a clinical geneticist to rule out a syndromic form of deafness. An interview excluded a personal or family history of diabetes, muscle disease, visual or neurological disorders. One or more members of each family had an ophthalmological review (including fundoscopy), an electrocardiogram and tests for proteinuria and haematuria.
Audiological data were collected. Pure-tone-air-conduction thresholds in the conversational frequencies (0.5, 1, 2 kHz) were calculated for each ear and were used to define the severity of deafness: mild (20 db
Results
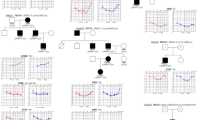
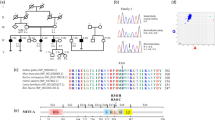
The pedigree of families no. 1 and 2 are reported in Figures 1 and 2.
Pedigree of family 1. Hearing-impaired individuals with audiogram are indicated by filled symbols, patients known to be deaf but without an audiogram are symbolised with hatched symbols. The question marks symbolise patients with apparently normal hearing for whom we did not obtain an audiogram. The probands are indicated by arrows. Asterisks indicate the individuals from whom DNA samples were obtained and tested. Generations are indicated on the left in Roman numerals, and the numbers under the individuals represent identification numbers for each generation.
Pedigree of family 2. For comments see Figure 1.
DNA analysis
In families 1 and 2, the variation T7511C was identified by direct sequencing and confirmed by PCR and restriction fragment length polymorphism (RFLP) analysis (Figures 3,4,5).
PCR-RFLP detection of T7511C in blood samples of family 1. Ethidium bromide-stained gel showing the diagnostic PCR-RFLP analysis used for rapid detection of the T7511C mutation. Patients are numbered as in the family pedigree. MW, 50-bp DNA molecular marker. PCR, undigested PCR fragment. The 23-bp is not visible on the gel.
PCR-RFLP detection of T7511C in blood samples urine, hair root cells and mouthwash samples of family 2 and quantification of mutant mtDNA. The PCR fragments were labelled, MboII digested and and analysed by electrophoresis on a 12% acrylamide gel. Patients are numbered as in the family pedigree. C: PCR fragment control. B: buccal cells from mouthwash samples, U: urine, H: hair root cells.
In the two index cases a homoplasmic variation was observed in PCR amplified fragments of tRNA Ser(UCN). No variation was detected in the tRNA Leu(UUR) or in the 12S rRNA. PCR and RFLP analysis were performed in 25 relatives of the two index cases. In family 1, T7511C was present in all the eight maternal relatives tested and absent in the three paternal relatives (Figure 4). The mutation was homoplasmic in the eight maternal relatives tested. In family 2, T7511C was present in all the 14 maternal relatives tested. Homoplasmic levels were observed in five relatives, whereas heteroplasmic levels were obtained in nine (Figure 5). In the heteroplasmic patients, the level of mutant mtDNA in blood, urine, hair root cells and mouthwash samples was highly variable (51–85%).
Clinical findings
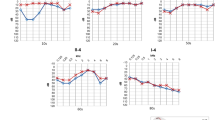
The propositus of family no. 1 (V1) (Figure 1) was born at term after a normal pregnancy and delivery. She began walking at a normal age. The first audiogram was performed at the age of 3 years because of a delay in language development and revealed a moderate bilateral sensorineural hearing loss. We saw her in the genetic counselling clinic at the age of 23 years. Her hearing loss was severe. She had no significant medical history except for recurrent venous thrombosis due to a mutation in the gene coding for the proaccelerin factor, which was transmitted by her mother. Karyotype, fundoscopy and a CT scan of the temporal bones were normal. She had no haematuria or proteinuria. The molecular analysis of CX26 (which is implicated in the most frequent form of NSSNHL) did not reveal any mutation. Her familial details revealed a genetic history with a pattern compatible with a mitochondrial inheritance. Indeed, seven out of 19 offspring of the deaf mothers were hearing impaired, whereas none of the three offspring of deaf fathers had any hearing impairment (see pedigree in Figure 1). The audiograms of eight patients carrying the mutation were analysed. Five of them had hearing impairment. The hearing loss was mild to severe with an average loss of 51 db (Table 1). The diagnosis of hearing impairment was established between 3 and 33 years (median: 21 years). One of the patients (IV6) presented with tinnitus, whereas her audiogram was normal. The hearing loss was progressive in the propositus but in none of the other deaf members of the family. In general, U-shaped curves were observed.
The propositus of family no. 2 (II.3) (Figure 2) came to the genetic counselling clinic at the age of 37 years. He presented with a prelingual (which begins before 2 years of age) bilateral severe sensorineural deafness. His hearing loss had been stable since diagnosis, even after exposure to streptomycin during childhood. He began walking at a normal age and he had no vestibular symptoms. Bilateral and permanent tinnitus in both ears began when he was 20 years old. His mother and four of his siblings presented with hearing impairment (see Figure 2). The age of diagnosis of their defect was very variable and the severity of the hearing impairment ranged from a mild unilateral to a severe bilateral deafness (Table 1). The propositus and all his deaf relatives had the T7511C mutation. The level of the mutated mtDNA was variable, patients II3 and III1 were homoplasmic and patients I2, II7, III12 were heteroplasmic. Moreover, four of his brothers and sisters and 11 of the children of his sisters had no known hearing problem. Among them, eight have been analysed and found to be heteroplasmic or homoplasmic for the T7511C mutation.
Discussion
The prevalence of mitochondrial anomalies in hearing loss is currently unknown however the large number of mtDNA mutations implicated in maternally inherited NSSHL suggests an important role for mitochondria in the function of the inner ear.
The age of onset and the severity may vary widely, even within a family. Some individuals are deaf from birth, some have a slow progressive hearing loss in adulthood, whereas others are audiological normal at the age of 50 years. Such variation suggests that environmental or genetic factors are involved, which influence the age of onset and progression of the hearing impairment. For the A1555G mutation, one of these factors is the administration of aminoglycosides. In our two families, only one patient received streptomycin, however, his hearing loss was discovered before, and it did not worsen following, exposure to this drug. If the level of mtDNA mutation in the blood was an accurate reflection of the level of the mutation in the cochlear hair cells, we would expect to find a correlation between the mutational load in blood and the severity of the hearing phenotype. However, this was not the case in our two families nor in the family described by Sue et al.10 Also, no correlation between severity of the hearing loss and the mutational load in urine hair root cells and mouthwash samples was observed. Recently, some data has been reported suggesting involvement of a nuclear factor in the variability of the phenotype associated with the mitochondrial 12S RNA mutation. Further analysis revealed that the maternal inheritance was only relevant in mice homozygous for a nuclear allele.12 Differences in some biochemical functions were demonstrated between lymphoblastoid cell lines derived from symptomatic and asymptomatic patients of a family carrying a homoplasmic form of the 12S RNA mutation.13 In addition, this variability among cells from symptomatic and asymptomatic individuals was reduced in transformants carryng the same nuclear background.14 Even though no modifier gene has been cloned in NSSNHL, Bykhovskaya et al15 have proposed a candidate locus for a modifier gene for the A1555G mutation. It would be interesting to investigate the implication of such a locus in the phenotypic expression in our two families. However, the modifying factor(s) affecting pathogenicity might differ between the various mutations. Therefore, a genome-wide search will be required in our two families to try and identify a new nuclear modifier gene.
All the subjects clinically observed in this study presented with an isolated deafness, even the patients carrying a homoplasmic mutation. The reason why a homoplasmic mutation can specifically affect only the ear, despite being present in all organs, is unknown. This tissue specificity could result from environmental factors, such as the concentration of aminoglycosides in the cochlea. Such an environmental factor has not been isolated in our study. A tissue specific pattern of gene expression could also explain the tissue-specific effects of the mtDNA mutations. But this hypothesis will not be tested since it is not possible to study the cochlear hair cells molecular function.
Only one family with the T7511C transition has been previously described.10 The origin of our two families is probably different as we did not observe the other two mitochondrial changes (T3308C and T5655C) reported by Sue et al.10 They suggested that the T7511C mutation has a relatively low pathogenic potential because it was present at almost homoplasmic levels in blood. In our study, nine out of the 21 patients carrying the mutation were heteroplasmic in the blood and some deaf subjects had a very low mutational load (Figure 3). Quantitative analysis in the inner ear cells needs to be performed in order to evaluate the pathogenicity of the T7511C mutation. It is noteworthy that Sue et al10 reported gradually progressive and symmetric hearing loss, whereas the patients included in this study presented with congenital, unilateral, or stable deafness. This variability underlines the existence of genetic and/or environmental factors modifying the expression of this mutation.
The identification of a T7511C mtDNA in two French families corroborates the recent findings that mtDNA mutations are a common cause of hearing loss. Screening for the mtDNA mutations, and in particular the T7511C mutation, should be performed in any patient with a family history of hearing loss when mitochondrial inheritance cannot be excluded. The discovery of a mitochondrial mutation in a deaf patient allows appropriate genetic counselling, as well as counselling to warn family members of the danger of exposure to drugs such as aminoglycosides. Understanding the mechanisms leading from a mitochondrial mutation to a hearing defect and the causes of the phenotypic variability should lead to novel targets for specific therapies in the future.
References
Petit C, Levilliers J, Hardelin JP . Molecular genetics of hearing loss Annu Rev Genet 2001 35: 589–645
Tamagawa Y, Kitamura K, Hagiwara H et al. Audiologic findings in patients with a point mutation at nucleotide 3243 of mitochondrial DNA Ann Otol Rhinol Laryngol 1997 106: 338–342
Ballinger SW, Shoffner JM, Hedaya EV et al. Maternally transmitted diabetes and deafness associated with a 10.4 kb mitochondrial DNA deletion Nat Genet 1992 1: 11–15
Servior KB, Hatamochi A, Stewart IA et al. Mitochondrial A7445G mutation in two pedigrees with palmoplantar keratoderma and deafness Am J Med Genet 1998 75: 179–185
Jaber L, Shohat M, Bu X et al. Sensorineural deafness inherited as a tissue specific mitochondrial disorder J Med Genet 1992 29: 86–90
Prezant TR, Agapian JV, Bohlman MC et al. Mitochondrial ribosomal RNA mutation associated with both antibiotic-induced and non-syndromic deafness Nat Genet 1993 4: 289–294
Casano RA, Johnson DF, Bykhovskaya Y, Torricelli F, Bigozzi M, Fischel-Ghodsian N . Inherited susceptibility to aminoglycoside ototoxicity: genetic heterogeneity and clinical implications Am J Otolaryngol 1999 20: 151–156
Estivill X, Govea N, Barcelo E et al. Familial progressive sensorineural deafness is mainly due to the mtDNA A1555G mutation and is enhanced by treatment of aminoglycosides Am J Hum Genet 1998 62: 27–35
Hutchin TP, Parker MJ, Young ID et al. A novel mutation in the mitochondrial tRNA(Ser(UCN)) gene in a family with non-syndromic sensorineural hearing impairment J Med Genet 2000 37: 692–694
Sue CM, Tanji K, Hadjigeorgiou G et al. Maternally inherited hearing loss in a large kindred with a novel T7511C mutation in the mitochondrial DNA tRNA(Ser(UCN)) gene Neurology 1999 52: 1905–1908
Sternberg D, Danan C, Lombes A et al. Exhaustive scanning approach to screen all the mitochondrial tRNA genes for mutations and its application to the investigation of 35 independent patients with mitochondrial disorders Hum Mol Genet 1998 7: 33–42
Johnson KR, Zheng QY, Bykhovskaya Y, Spirina O, Fischel-Ghodsian N . A nuclear-mitochondrial DNA interaction affecting hearing impairment in mice Nat Genet 2001 27: 191–194
Guan MX, Fischel-Ghodsian N, Attardi G . Biochemical evidence for nuclear gene involvement in phenotype of non- syndromic deafness associated with mitochondrial 12S rRNA mutation Hum Mol Genet 1996 5: 963–971
Guan MX, Fischel-Ghodsian N, Attardi G . Nuclear background determines biochemical phenotype in the deafness-associated mitochondrial 12S rRNA mutation Hum Mol Genet 2001 10: 573–580
Bykhovskaya Y, Yang H, Taylor K et al. Modifier locus for mitochondrial DNA disease: linkage and linkage disequilibrium mapping of a nuclear modifier gene for maternally inherited deafness Genet Med 2001 3: 177–180
Acknowledgements
This work was supported by the Fondation pour la Recherche Médicale, The Association Française contre les Myopathies, The Institut National de la Santé et de la Recherche Médicale, and The Association pour l'Etude de la Pathologie Pédiatrique.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chapiro, E., Feldmann, D., Denoyelle, F. et al. Two large French pedigrees with non syndromic sensorineural deafness and the mitochondrial DNA T7511C mutation: evidence for a modulatory factor. Eur J Hum Genet 10, 851–856 (2002). https://doi.org/10.1038/sj.ejhg.5200894
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5200894
Keywords
This article is cited by
-
Mitochondrial mutations in maternally inherited hearing loss
BMC Medical Genetics (2017)
-
Genetic etiology study of the non-syndromic deafness in Chinese Hans by targeted next-generation sequencing
Orphanet Journal of Rare Diseases (2013)
-
The Role of Mitochondrial DNA Mutations in Hearing Loss
Biochemical Genetics (2013)
-
The genetic bases for non-syndromic hearing loss among Chinese
Journal of Human Genetics (2009)
-
MRPS18CP2 alleles and DEFA3absence as putative chromosome 8p23.1 modifiers of hearing loss due to mtDNA mutation A1555G in the 12S rRNA gene
BMC Medical Genetics (2007)