Abstract
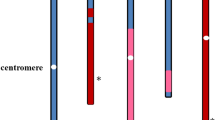
The higher plant Arabidopsis thaliana (Arabidopsis) is an important model for identifying plant genes and determining their function. To assist biological investigations and to define chromosome structure, a coordinated effort to sequence the Arabidopsis genome was initiated in late 1996. Here we report one of the first milestones of this project, the sequence of chromosome 4. Analysis of 17.38 megabases of unique sequence, representing about 17% of the genome, reveals 3,744 protein coding genes, 81 transfer RNAs and numerous repeat elements. Heterochromatic regions surrounding the putative centromere, which has not yet been completely sequenced, are characterized by an increased frequency of a variety of repeats, new repeats, reduced recombination, lowered gene density and lowered gene expression. Roughly 60% of the predicted protein-coding genes have been functionally characterized on the basis of their homology to known genes. Many genes encode predicted proteins that are homologous to human and Caenorhabditis elegans proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Accessions
EMBL/GenBank/DDBJ
Data deposits
The sequence and preliminary analysis of clones and PCR products were made available immediately after completion through the MATDB database12. The results of computational analyses, including the functional and structural characterization of the protein sequences involved, are available at the PEDANT-pro genome analysis server (http://pedant.mips.biochem.mpg.de). Underlying recombinant clones can be obtained from the NASC (http://www.nasc.ac.uk/). The accession numbers for chromosome 4 are: short arm, AJ270058; long arm, AJ270060.
References
Meinke,D. W., Cherry,J. M., Dean,C. D., Rounsley,S. & Koornneef,M. Arabidopsis thaliana: a model plant for genome analysis. Science 282, 662–682 (1998).
Copenhaver,G. C. & Pikaard,C. S. Two dimensional RFLP analyses reveal megabase-sized clusters of rRNA gene variants in Arabidopsis thaliana, suggesting local spreading of variants as the mode for gene homogenization during concerted evolution. Plant J. 9, 273–282 (1996).
Lister,C. & Dean,C. Recombinant inbred lines for mapping RFLP and phenotypic markers in Arabidopsis thaliana. Plant J. 4, 745–750 (1993).
Bevan,M. et al. Analysis of 1.9 Mb of contiguous sequence from chromosome 4 of Arabidopsis thaliana. Nature 391, 485–488 (1998).
Kotani,H., Nakamura,Y., Sato,S., Kaneko,T., Asamizu,E. et al. Structural analysis of Arabidopsis thaliana chromosome 5. II. Sequence features of 1,044,062 bp covered by thirteen physically-assigned P1 clones. DNA Res. 4, 291–300 (1997).
The C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1999).
Gardner,M. J. et al. Chromosome 2 sequence of the human malarial parasite Plasmodium falciparum. Science 282, 1126–1132 (1998).
Bowman,S. et al. The complete nucleotide sequence of chromosome 3 of Plasmodium falciparum. Nature 400, 532–538 (1999).
Richards,E. J. & Ausubel,F. M. Isolation of a higher eukaryotic telomere from Arabidopsis thaliana. Cell 53, 127–136 (1988).
Copenhaver,G. C. & Pikaard,C. S. RFLP and physical mapping with an rDNA-specific endonuclease reveals that nucleolus organiser regions of Arabidopsis thaliana adjoin the telomeres on chromosomes 2 and 4. Plant J. 9, 259–272 (1996).
Richards,E. J., Goodman,H. M. & Ausubel,F. M. The centromeric region of Arabidopsis thaliana chromosome 1 contains telomere-similar sequences. Nucleic Acids Res. 19, 3351–3357 (1991).
Mewes,H. W. et al. MIPS: a database for genomes and protein sequences. Nucleic Acids Res. 27, 44–48 (1999).
Hubbard,T. J. P., Ailey,B., Brenner,S. E., Murzin,A. G. & Chothia,C. SCOP: a structural classification database of proteins. Nucleic Acids Res. 27, 254–256 (1999).
Gerstein,M. A structural census of genomes: comparing bacterial, eukaryotic and archeal genomes in terms of protein structure. J. Mol. Biol. 274, 562–576 (1997).
Mizutani,M., Ward,E. & Ohta,D. Cytochrome P-450 superfamily in Arabidopsis thaliana: isolation of cDNAS, differential expression, and RFLP mapping of multiple cytochromes P-450. Plant Mol. Biology 37, 39–52 (1998).
Joazeiro,C. A. P. et al. The tyrosine kinase negative regulator c-Cbl as a RING-type, E2-dependent ubiquitin-protein ligase. Science 286, 309–312 (1999).
Jensen,R. B., Jensen,K. L., Jespersen,H. M. & Skriver,K. Widespread occurrence of a highly conserved RING-H2 zinc finger motif in the model plant Arabidopsis thaliana. FEBS Lett. 436, 283–287 (1998).
Moncrief,N. D., Kretsinger,R. H. & Goodman,M. Evolution of EF-hand calcium-modulated proteins 2. Domains of several sub-families have diverse evolutionary history. J. Mol. Evol. 30, 522–562 (1991).
McAinsh,M. R. & Hetherington,A. M. Encoding specificity in Ca2+ signaling systems. Trends Plant Sci. 3, 32–36 (1998).
Douglas,S. E. Plastid evolution: origins, diversity, trends. Curr. Opin. Genet. Dev. 8, 655–661 (1998).
Emanuelsson,O., Nielsen,H. & von Heijne,G. ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci. 8, 978–984 (1999).
Gamas,P., de Carvalho Niebel,F., Lescure,N. & Cullimore,J. V. Use of a subtractive hybridisation approach to identify new Medicago truncatula genes induced during nodule development. Mol. Plant Microbe Interaction 9, 233–242 (1996).
Parniske,M. et al. Novel disease resistance specificities result from sequence exchange between tandemly repeated genes at the Cf-4/9 locus in tomato. Cell 9, 821–832 (1997).
Mewes,H.-W. et al. Nature 387 (Suppl.) 7–8 (1997).
Round,E., Flowers,S. K. & Richards,E. J. Arabidopsis thaliana centromere regions: genetic map positions and repetitive DNA structure. Genome Res. 7, 1045–1054 (1997).
Fransz,P. F. et al. Cytogenetics for the model system Arabidopsis thaliana. Plant J. 13, 867–876 (1998).
Richards,E. J. & Dawe,R. K. Plant centromeres: structure and control. Curr. Opin. Plant Biol. 1, 130–135 (1998).
Murphy,T. D. & Karpen,G. H. Localization of centromere function in a Drosophila minichromosome. Cell 82, 599–609 (1995).
Henning,K. A. et al. Human artificial chromosomes generated by modification of a yeast artificial chromosome containing both human alpha satellite and single-copy sequences. Proc. Natl Acad. Sci. USA 96, 592–597 (1999).
Copenhaver,G. P., Browne,W. E. & Preuss,D. Assaying genome-wide recombination and centromere functions with Arabidopsis tetrads. Proc. Natl Acad. Sci. USA 95, 247–252 (1998).
Grewal,S. I. S. & Klar,A. J. S. A recombinationally repressed region between mat2 and mat3 loci shares homology to centromeric repeats and regulates directionality of mating type switching in fission yeast. Genetics 146, 1221–1238 (1997).
Allshire,R. C., Nimmo,E. R., Ekwall,K., Javerzat,J.-P. & Cranston,G. Mutations derepressing silent centromeric domains in fission yeast disrupt chromosome segregation. Genes Dev. 9, 218–233 (1995).
Xu,X. J., Hsai,A.-P., Zhang,L., Nikolau,B. J. & Schnable,P. S. Meiotic recombination breakpoints resolve at high rates at the 5′ end of a maize coding sequence. Plant Cell 7, 2151–2161 (1995).
Lin,X. et al. Sequence and analysis of chromosome 2 of the plant Arabidopsis thaliana. Nature 402, 761–768 (1999).
Choi,S. D., Creelman,R., Mullet,J. & Wing,R. A. Construction and characterisation of a bacterial artificial chromosome library from Arabidopsis thaliana. Weeds World 2, 17–20 (1995).
Mozo,T. et al. A complete BAC-based physical map of the Arabidopsis thaliana genome. Nature Genet. 22, 271–275 (1999).
Marra,M. et al. A map or sequence analysis of the Arabidopsis thaliana genome. Nature Genet. 22, 265–270 (1999).
Bent,E., Johnson,S. & Bancroft,I. BAC representation of two low-copy regions of the genome of Arabidopsis thaliana. Plant J. 13, 849–855 (1998).
Vos,P. et al. AFLP, a new technique for DNA fingerprinting. Nucleic Acids Res. 23, 4407–4414 (1995).
Borodovsky,M. & Peresetsky,A. Deriving non-homogeneous DNA Markov chain models by cluster analysis algorithm minimizing multiple alignment entropy. Comput. Chem. 18, 259–267 (1994).
Uberbacher,E. C. & Mural,R. J. Locating protein coding regions in human DNA sequences by a multiple sensor-neural network approach. Proc. Natl Acad. Sci. 88, 1261–1265 (1991).
Burge,C. & Karlin,S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94 (1997).
Hebsgaard,S. M. et al. Splice site prediction in Arabidopsis thaliana pre-mRNA by combining local and global sequence information. Nucleic Acids Res. 24, 3439–3452 (1996).
Altschul,S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Fichant,G. A. & Burks,C. Identifying potential tRNA genes in genomic DNA sequences. J. Mol. Biol. 220, 659–671 (1991).
Wootton,J. C. & Federhen,S. Statistics of local complexity in amino acid sequences and sequence databases. Comput. Chem. 17, 149–163 (1993).
Lupas,A. N., van Dyke,M. & Stock,J. Predicting coiled coils from protein sequences. Science 252, 1162–1164 (1991).
Klein,P., Kanehisa,M. & DeLisi,C. The detection and classification of membrane-spanning proteins. Biochim. Biophys. Acta 815, 468–476 (1985).
Schäffer,A. A. et al. IMPALA: Software to match a protein sequence against a collection of PSI-BLAST-constructed position–specific score matrices. Bioinformatics, in the press.
Pearson,W. R. & Lipman,D. J. Improved tools for biological sequence comparison. Proc. Natl Acad. Sci. 85, 2444–2448 (1988).
Acknowledgements
We wish to thank A. Schäffer for invaluable assistance with the IMPALA software and S. Brenner for providing an up-to-date version of the SCOP database. We are grateful to S. Choi for a copy of his large-insert BAC library. Scientists at the John Innes Centre are acknowledged for their help in interpreting gene function. This work was funded in part by Contracts from the European Commission, by the National Science Foundation (NSF) Cooperative Agreement (funded by the NSF, US Department of Agriculture and the US Department of Energy), and by a grant from the USDA NRI Plant Genome Program. Additional support from the Biotechnology and Biological Sciences Research Council, Bundesministerium f. Bildung, Forschung und Technologie, Groupe de Recherche et d'etude des Genomes, Plan Nacional de Investigacion Cientifica y Technica, Westvaco Corporation and D. L. Luke III is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mayer, K., Schüller, C., Wambutt, R. et al. Sequence and analysis of chromosome 4 of the plant Arabidopsis thaliana. Nature 402, 769–777 (1999). https://doi.org/10.1038/47134
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/47134
This article is cited by
-
Transcriptome mapping related genes encoding PR1 protein involved in necrotic symptoms to soybean mosaic virus infection
Molecular Breeding (2023)
-
Comparative transcriptome analysis of MeJA-responsive AP2/ERF transcription factors involved in notoginsenosides biosynthesis
3 Biotech (2020)
-
A novel single-base mutation in CaBRI1 confers dwarf phenotype and brassinosteroid accumulation in pepper
Molecular Genetics and Genomics (2020)
-
Genome-Wide Identification and Expression Profiling of Ascorbate Peroxidase (APX) and Glutathione Peroxidase (GPX) Genes Under Drought Stress in Sorghum (Sorghum bicolor L.)
Journal of Plant Growth Regulation (2018)
-
Expression of four phosphate transporter genes from Finger millet (Eleusine coracana L.) in response to mycorrhizal colonization and Pi stress
3 Biotech (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.