Abstract
Excitotoxic insults induce c-Jun N-terminal kinase (JNK) activation, which leads to neuronal death and contributes to many neurological conditions such as cerebral ischemia and neurodegenerative disorders. The action of JNK can be inhibited by the D-retro-inverso form of JNK inhibitor peptide (D-JNKI1), which totally prevents death induced by N-methyl-D-aspartate (NMDA) in vitro and strongly protects against different in vivo paradigms of excitotoxicity. To obtain optimal neuroprotection, it is imperative to elucidate the prosurvival action of D-JNKI1 and the death pathways that it inhibits. In cortical neuronal cultures, we first investigate the pathways by which NMDA induces JNK activation and show a rapid and selective phosphorylation of mitogen-activated protein kinase kinase 7 (MKK7), whereas the only other known JNK activator, mitogen-activated protein kinase kinase 4 (MKK4), was unaffected. We then analyze the action of D-JNKI1 on four JNK targets containing a JNK-binding domain: MAPK-activating death domain-containing protein/differentially expressed in normal and neoplastic cells (MADD/DENN), MKK7, MKK4 and JNK-interacting protein-1 (IB1/JIP-1).
Similar content being viewed by others
Main
In many nervous system disorders, including cerebral ischemia, traumatic brain injury and neurodegenerative diseases, overactivation of N-methyl-d-aspartate (NMDA) receptors leads to neuronal damage, resulting in neuronal loss and consequent severe neurological impairment. This cascade of neuronal injury, referred to as ‘excitotoxicity’, is still only partly understood. Dying neurons activate complex signal transduction events to trigger their death program, and the c-Jun N-terminal kinase (JNK) pathway plays an important role in this process.1
D-retro-inverso form of JNK inhibitor (D-JNKI1) is an extremely potent neuroprotectant against excitotoxicity of cortical neurons and against different in vivo paradigms of neurodegeneration.2, 3, 4 The active part of this peptide contains a retro–inverso form of a 20-amino-acid sequence (JBD20) from the JNK-binding domain (JBD) of the scaffold protein JNK-interacting protein-1 (IB1/JIP-1), and it blocks the access of JNK to many of its targets.5, 6, 7 Recently, Negri et al.8, 9 performed a detailed re-examination of the JBD-containing proteins and identified 19 different substrates of JNK. They then proved in a cell-free assay that the JBD20 sequence prevented interactions and phosphorylations by JNK of nine of these targets. Among these nine substrates, we have studied the following four that might participate in regulating the death of cortical neurons: (1) MADD/DENN (MAPK-activating death domain-containing protein/differentially expressed in normal and neoplastic cells); (2–3) mitogen-activated protein kinase kinase 4 (MKK4) and mitogen-activated protein kinase kinase 7 (MKK7), the two direct upstream activators of JNK and (4) the scaffold protein IB1/JIP-1.
In particularly, MADD/DENN was identified as a substrate for JNK3,10 the isoform most clearly involved in excitotoxicity11, 12 and mostly expressed in the brain.10, 11, 12 Increasing evidence supports a strong correlation between low MADD/DENN expression and neuronal loss.13, 14
MKK4 and MKK7 are the only known JNK activators. In some cell types, MKK4 activates JNK primarily by stress stimuli and MKK7 by inflammatory cytokines15, 16, 17 but in neuronal excitotoxicity their contributions are still not known.15 Both MKK48, 9, 18 and MKK78, 9 have a JBD indicating that these activators of JNK are also targets and potentially sensitive to D-JNKI1.
Finally, the scaffold protein IB1/JIP-1 itself presents a JBD20 domain: it plays an important role in organizing the JNK cascade and shows a tight relationship with cell death. IB1/JIP-1 can influence JNK signalling both positively and negatively, depending on the amounts of IB1/JIP-1 and its interacting kinases.19, 20, 21
Our aim is to identify the critical features of the powerful protective action of the D-JNKI1 peptide and to examine JNK targets in excitotoxic neuronal death pathways. This can offer important clues to produce new inhibitors of excitotoxicity.
Results
Phase contrast microscopy indicated that exposure of neurons to 100 μM NMDA induced immediate (within 1 h) marked swelling of neuronal cell bodies, followed by total neuronal degeneration over the next 48 h (97% cell death at 48 h, as reported previously by Borsello et al.6).
Dying neurons were visualized with propidium iodide (PI, 2 μg/ml, 5 min). Morphological observation of control cultures indicated the presence of very few apoptotic neurons (cellular shrinkage and punctate staining, indicating clumping of chromatin in a condensed nucleus, see Figure 1a). D-JNKI1-protected neurons presented hardly any apoptotic bodies such as control (Figure 1b) and this clearly contrasted with NMDA-treated cultures where most neurons were necrotic (nuclei stained uniformly with PI, Figure 1c). In our model, all neurons received the excitotoxic stimulus at the same time but death was not tightly synchronized: some neurons died early, others late – a fact that has been analyzed in some details in the context of excitotoxicity.22
Morphological observation of neuronal death (a–c). Phase contrast microscopy of cortical neuronal morphology, scale bar, 60 μm. In control cultures, very few neurons presented apoptotic nuclei visualized by PI staining as shown at higher magnification in the inset scale Bar, 5 μm (a). The NMDA-treated cultures presented strong sign of necrosis: arrows indicated swollen neurons, containing enlarged nuclei. The necrotic nuclei are stained uniformly with PI (c). NMDA-treated cultures protected by D-JNKI1 presented control morphology (b) without any necrotic features and as in the control condition PI hardly labeled only a very few cells with apoptotic features. Effect of D-JNKI1 on NMDA-induced changes in Ca2+ (d–m). Control neurons visualized by interference contrast microscopy. Scale bar, 20 μm (d). Image of Fura-2-loaded neurons recorded under basal conditions (e). Ca2+ response to NMDA stimulation after stimulation with 10 and 100 μM NMDA. Intracellular Ca2+ responses range from blue to red for elevated values (f). Cortical neurons were pretreated with 2 μM D-JNKI1 and visualized by interference contrast microscopy (h). The same Fura-2-loaded neurons are shown in the basal condition (i) and after exposure to 10 and 100 μM NMDA (l). Intracellular Ca2+ response shows normal behavior in the presence of D-JNKI1. The plots present Ca2+ traces from selected cells in (g) the control group of neurons (n=41 cells from two experiments) and (m) the D-JNKI1-treated group (n=44 cells from three experiments)
NMDA-induced changes in Ca2+i in control neurons compared to D-JNKI1-treated neurons
We first analyzed the molecular events underlying excitotoxic signalling induced by NMDA, and the protective effects of D-JNKI1, from the cell surface through the cytoplasm to the nucleus.
We started by checking whether D-JNKI1 affected the normal functioning of the NMDA receptor, monitoring the Ca2+i response to NMDA stimulation of cortical neurons by fluorescence microscopy using the Fura-2 probe. Exposure of cortical neurons to 10 or 100 μM NMDA markedly raised the intracellular Ca2+ concentration in control neurons (Figure 1e–g). In D-JNKI1 pretreated neurons, this stimulation resulted in similar Ca2+i responses to those in control neurons (Figure 1i–m). In four independent experiments, the proportion of neurons responding to the stimulus was the same in the control group (20 responders/41 total) and in the D-JNKI1-treated group (21 responders/44 total). In non-treated neurons responders, the amplitude of the Ca2+i response was respectively 536±127 and 376±130 nM after application of 10 and 100 μM NMDA. The amplitude of the Ca2+i responses of D-JNKI1-treated neurons responders was not significantly different from the non-treated cells and averaged 357±57 nM (P=0.199) and 443±82 nM (P=0.662). Therefore, D-JNKI1 does not significantly affect the function of the NMDA receptor or its associated Ca2+i response.
Excitotoxic signalling pathway in cortical neurons
To better characterize NMDA-induced neuronal death of cortical neurons, we tested whether different inhibitor combinations would prevent it.
The addition of 1–3 μg/ml of cycloheximide to neuronal cultures efficiently inhibited protein synthesis more than 90% (data not shown), but adding it before the NMDA gave no protective effect on neuronal loss (Figure 2a). Neuronal death was assessed after 5 h by lactate dehydrogenase (LDH) release into the bathing medium as described previously.6 We used a combination of cycloheximide and D-JNKI1 given before NMDA stress, and the peptide still provided total protection after 5 h (Figure 2a). We then examined whether the broad-spectrum caspase inhibitor z-Val-Ala-Asp fluoromethylketone (z-VAD.fmk) partially reduced NMDA-induced death (Figure 2b). Addition of 150 or 300 μM of z-VAD.fmk to neuronal cultures before NMDA treatment efficiently inhibited caspase-3 activation (Figure 2c) but did not block death measured after 5 h by LDH (Figure 2b). Although the caspase inhibitor alone did not prevent induced neuronal death, D-JNKI1 did prevent caspase-3 activation (Figure 2c).
Inefficacy of caspase inhibitor cycloheximide against NMDA-induced neuronal death. Cycloheximide is ineffective against NMDA-induced excitotoxicity, but either alone or with cycloheximide D-JNKI1 protects cortical neurons against NMDA at 5 h (a). Effect of z-VAD.fmk and 2 μM D-JNKI1 on neuronal death as assayed by LDH release after 5 h NMDA exposure, z-VAD50: 50 μM z-VAD.fmk; z-VAD150: 150 μM z-VAD.fmk; ZVAD300: 300 μM z-VAD.fmk (b). Effect of z-VAD.fmk and D-JNKI1 on pro-caspase-3 cleavage: in Western blot anti-caspase-3 antibodies reveal both pro-caspase-3 and the cleaved p12 product (c). Protective effect of the combined treatment with z-VAD.fmk and cycloheximide. The joint treatment provides 60% neuroprotection against NMDA. Neuronal death was measured with LDH release at 24 h (d). Quantifications were performed in 10 independent experiments (±S.E.M.)
As NMDA-induced death persisted with the protein synthesis inhibitor and with the caspase inhibitor alone, blocking only one pathway is not enough to prevent neuronal loss. However, D-JNKI1 conferred total protection even in the presence of the protein synthesis inhibitor. We tried a combined treatment with z-VAD.fmk and cycloheximide to prevent NMDA-induced death, and it did provide 60% neuroprotection (Figure 2d). It thus appears that NMDA activates at least two different death pathways, one independent of protein synthesis and the other dependent on it. In the first, post-translational modifications of pre-existing proteins are taking part in the death process and are associated with the JNK action. The second pathway depends on protein synthesis, owing to the well-known activation of transcription factors such as c-jun,11, 13, 23 ATF2, Elk-1 and c-fos induction.6, 8, 9, 24
To identify the nature of the death pathway not blocked by z-VAD.fmk and cycloheximide but inhibited by D-JNKI1, we studied the JBD–containing non-transcription factor targets whose post-translational modifications are related to JNK phosphorylation and inhibited by JBD20 in cell-free competitive assay.8, 25
MADD/DENN role in excitotoxicity and the effect of D-JNKI1
To study MADD/JNK interaction, we first performed in vitro kinase assays with MADD substrate produced as glutathione S-transferase (GST) fusion proteins. Recombinant JNK2 and JNK3 protein kinases were incubated with [γ-33P]ATP, the constructed GST-MADD fusion proteins and either with JBD20 or JBDmut for 30 min.
As shown in Figure 3g, JNK2 and JNK3 specifically associate10 with and phosphorylate MADD/DENN. No phosphorylation was detected for both isoforms in the presence of the retro–inverso JBD20 peptide, whereas the control mutated peptide JBDmut2, 8 had no effect.
MADD/DENN in excitotoxicity. Immunofluorescence study (a–f): neurons are labeled with anti-MADD/DENN antibody (a, b and c) and nuclei are counterstained with DAPI (d, e and f). In control condition, the staining of MADD/DENN is cytoplasmic (a) but after NMDA treatment MADD is also present in the nuclear compartment and colocalizes with DAPI (b, arrow and e). NMDA-treated neurons show a more strongly labeled nucleus with DAPI than control ones, and dying neurons are easily identifiable (e). D-JNKI1 pretreatment prevents MADD translocation within nuclei (c) and DAPI reveals very few intense and uniformly stained nuclei (f). Scale bar, 10 μm. In vitro JNK competitive assay showing specificity of the JNK inhibitor peptide (g): kinase assays with recombinant activated JNK2 and -3, using GST-MADD as substrate as described previously.5, 6 The JBD20 peptide inhibits MADD phosphorylation whereas the control-mutated peptide JBDmut has no effect. Western blot study (h) of MADD/DENN after an NMDA time course on total extracts of cortical neurons, with the corresponding quantification. The MADD/DENN-reactive band at ≅200 kDa decreases immediately after 5 min of treatment and at 15 min of NMDA exposure presents a reduction of 90%. MADD degradation pathways following NMDA (i–k). In (i), cortical neurons were pre-treated with 5 and 10 μM calpastatin peptide for 1 h and incubated with NMDA for 5 h. Calpain inhibitor has no effect on MADD degradation, whereas D-JNKI1 strongly prevents it. The Western blot densitometry clearly proves that calpains are not involved in MADD degradation. Proteasome inhibitors (j–k): the lactacystin proteasome inhibitor (against the 20S complex) was tested and compared with the D-JNKI1 effect on MADD degradation (j). The lactacystin prevents MADD degradation induced by NMDA in a dose-dependent way, D-JNKI1 protects as well. MG132, the second proteasome inhibitor (of the 26S complex), was tested in parallel with the two neuroprotective treatments (k): D-JNKI1 (2 μM) and z-VAD.fmk/cycloheximide (150–300 μM, see Figure 2). Western blot analysis and quantification demonstrates that MG132 powerfully prevented MADD degradation. Both MG132 and D-JNKI1 preserve a level of MADD comparable to the control condition, whereas the z–Vad.fmk/cycloheximide has no effect. Quantifications were performed in five independent experiments (±S.E.M.)
We then analyzed the expression of MADD in response to NMDA and the effect of D-JNKI1 by immunofluorescence and Western blots in cortical neurons. In control conditions, MADD staining was sparse and diffusely distributed in the cytoplasm of neurons (Figure 3a,d), whereas after 1 h of NMDA treatment MADD was strikingly relocalized into the nuclei and nucleoli of dying neurons (Figure 3b, e: MADD colocalizes with DAPI, N.B.: here DAPI is more intensely and uniformly labeled). The nuclear translocation of MADD was mostly prevented by D-JNKI1 pretreatment (Figure 3c–f).
To further characterize the changes in MADD expression, we studied it during NMDA treatment from 5 min to 5 h in Western blots of total cortical neuronal extracts. The level of MADD fell rapidly after NMDA; there was a clear effect after only 15 min and at 5 h the decrease was 80% (Figure 3h).
We investigated whether calpains or the ubiquitin-proteolytic pathway are implicated in the MADD degradation mediated by NMDA. For this purpose, cortical neurons were pretreated with calpastatin, a specific calpain inhibitor, and with two potent proteasome inhibitors: lactacystin (20S proteasome inhibitor) and MG132 (against the 26S complex). Western blotting showed that MADD degradation was unaffected by calpastatin (Figure 3i) but was strongly inhibited by both proteasome inhibitors, lactacystin and MG132 (Figure 3j,k). These data prove that calpains are not involved in MADD degradation, whereas the proteasome pathway is responsible for it following NMDA treatment. However, inhibition of MADD degradation through the proteasome did not protect neurons from death (data not shown). D-JNKI1 also prevented MADD degradation like lactacystin and MG132 (see Figure 3j,k), whereas with z-VAD.fmk/cycloheximide there is no protection.
Responses of MKK7 and MKK4 to NMDA and the effects of D-JNKI1
We attempted first to document the NMDA-induced changes in expression and phosphorylation of MKK4 and MKK7 and the effects of D-JNKI1. As both MKK4 and MKK7 contain a JBD20 homology domain,8, 18, 25 they are not only JNK activators but also substrates and their phosphorylation by JNK is likely to be sensitive to D-JNKI1.
In the time course (from 10 min to 5 h) of the NMDA effect on total extracts of cortical neurons, the MKK7 level deeply declined (90%) in the first 10 min, whereas the decrease in MKK4 was weaker and reached 90% only after 5 h of NMDA (Figure 4a and Supplementary Figure).
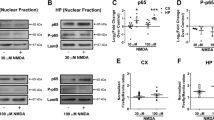
Responses of MKK7 and MKK4 to NMDA. Effect of NMDA on MKK4 and MKK7 levels in total extracts (a). MKK7 decreases after 10 min of treatment, whereas MKK4 is more stable and starts to decrease only at 5 h. Effect of NMDA on fraction extracts (b). In control conditions, there is a pool of MKK7 in both the nuclear and the cytoplasmic fractions. After NMDA treatment, cytoplasmic MKK7 translocates into the nuclear fraction. To locate MKK4, we tested two antibodies. MKK4 is mainly cytoplasmic and does slightly increase in the nuclear fraction in response to NMDA. The first antibody used (1st) presents only a nonspecific (ns) band at the wrong molecular weight in the nuclear fraction. The second one (2nd) shows traces of MKK4 in this fraction. Effect of NMDA on P-MKK7 and P-MKK4 (c, d). MKK7 is activated by NMDA and the activation arrived at the top at 10 min for then rapidly decreasing at the control level. MKK4 is not activated by NMDA and the excitotoxic stimulation induced a vigorous decrease of P-MKK4 level. MKK7 phosphorylation assessed by combined immunoprecipitation (anti-P-MKK7-antibody) and Western blotting (anti-total MKK7 antibody) (e). Immunoprecipitation results were obtained using pools of 10 different dishes and were performed in three independent experiments to quantify the MKK7 activation with densitometry analysis: there is strong MKK7 activation at 10 min in the nuclear fraction (e, g). MKK4 phosphorylation monitored by combined immunoprecipitation (anti-P-MKK4-antibody) and Western blotting (anti-total MKK4 antibody) (f). Results are quantified (in h) and MKK4 is not activated at 10 min in the nuclear or the cytoplasmic fraction. Effects of D-JNKI1 on P-MKK7 and P-MKK4 (i, j). P-MKK7 is activated by NMDA (i) compared to control and the D-JNKI1 treatment (2 μM) does not affect the P-MKK7 level. P-MKK4 level was analyzed in control condition and D-JNKI1-treated neurons (24 h) do not present any changes in P-MKK4 level (j). Quantifications were performed in eight independent experiments (±S.E.M.)
To assess the changes in their subcellular localization, we performed an immunofluorescence analysis and a fraction separation. Nuclear and cytoplasmic fractions of control and NMDA-treated neurons were charged and blotted for MKK7 and MKK4 (Figure 4b). In the control condition, MKK7 was distributed in both the cytoplasm and nucleus, but exposure to NMDA caused a massive translocation of MKK7 into neuronal nuclei (Figures 4b and 5). In contrast, MKK4 was mainly cytoplasmic, and NMDA caused no changes in its distribution in the first hours of treatment (Figures 4b and 6). Only at 5 h of NMDA exposure it was possible to detect scant traces of MKK4 in the nuclear fraction using the anti-MKK4 antibody from Upstate (no. 07–194; see Materials and Methods). We repeated this experiment using a second antibody (from Santa-Cruz, see Coffey et al.26), but were still not able to detect an increase of MKK4 in the nuclei after NMDA stimulation.
MKK7 in excitotoxicity. Immunofluorescence study (a–j): after 1 h of 100 μm of NMDA cortical neurons are stained with MKK7 antibody (a, b, c, h and i) and DAPI (d, e, f, g and j). In the control condition, the labelling is punctate in the cell body (a), and sometimes nuclear (b, arrow). After 1 h of NMDA treatment, the staining becomes almost exclusively nuclear (c, arrow). D-JNKI1 mostly prevents nuclear translocation (h, i) although if it is still possible to find neurons with nuclear labelling (i, arrow). Scale bar, 10 μm. Western blotting (k, l) of MKK7 following a NMDA time course on nuclear and cytoplasmic extracts of cortical neurons, in the presence or absence of D-JNKI1. In control conditions, there is a pool of MKK7 in the cytoplasmic fractions and hardly any MKK7 is detectable in the nuclear fraction. After only 5 min NMDA treatment, cytoplasmic MKK7 translocates into the nuclear fraction. D-JNKI1 prevents nuclear translocation and retains MKK7 in the cytoplasm (k, l). Results were similar in nine independent experiments (±S.E.M.)
MKK4 immunofluorescence (using 2nd antibody). After 1 h of 100 μm NMDA, cortical neurons are labeled with this MKK4 antibody (a, c and e) and stained with DAPI (b, d and f). In the control condition (a), like after 1 h NMDA stimulation (c), the labelling is equally distributed in both nuclei and cytoplasm. D-JNKI1 pretreatment has no effect on MKK4 staining. Scale bar, 10 μm. Western blotting (i) of MKK4 following an NMDA time course on cytoplasmic extracts of cortical neurons in the presence or absence of D-JNKI1. NMDA reduced the MKK4 level, whereas D-JNKI1 strongly preserved it (i, j). Results were similar in six independent experiments (±S.E.M.)
We then evaluated the effect of NMDA on the degradation and translocation of MKK7 by immunofluorescence. In control neurons, MKK7 staining was typically punctate in the perikaryal cytoplasm (Figure 5a, d) and sometimes a nuclear pool of MKK7 was also detectable (Figure 5b, e). After 1 h of NMDA stimulation, the staining was localized almost exclusively in the nuclei of the stressed neurons (Figure 5c, f). D-JNKI1 largely prevented this nuclear translocation (as shown in Figure 5g, h i and j). Using fraction separation Western blotting this effect is more appreciable and measurable (Figure 5k, l). The peptide entirely prevented the decrease in cytoplasmic MKK7 from 5 min to 1 h of NMDA treatment and even at 5 h D-JNKI1 still prevented 50% of the decrease (Figure 5k, l).
For MKK4 immunofluorescence, only the Santa-Cruz antibody was utilizable: the labelling was equally distributed in nuclei and cytoplasm in control conditions as well as after NMDA stimulation, without any changes in localization throughout the 5 h examined (Figure 6a–f). In Western blot studies, D-JNKI1 blocked the NMDA-induced decrease in MKK4 level (Figure 6i, j).
Activation of MKK7 and MKK4 after NMDA stimulation
To determine to what extent MKK4 and MKK7 contribute to JNK activation after NMDA stress, we analyzed their phosphorylation in the first hours of NMDA treatment. In this model, maximal JNK activation occurs in the first hour of NMDA treatment, followed by a decrease6 and it is reasonable to expect a prior phosphorylation of the two direct upstream JNK activators.
We looked at P-MKK4 and P-MKK7 forms from 5 min to 5 h after NMDA exposure (Figure 4c,d).
As clearly shown in Figure 4c and d, NMDA caused a striking increase in MKK7 phosphorylation in the first 30 min of stimulation, whereas no increase in phosphorylation of MKK4 was detectable, and the level of P-MKK4 in fact decreased progressively to virtually zero (Figure 4d).
To examine and quantify more exactly the activation of these two proteins, we did additional studies involving immunoprecipitation with the P-MKK7 and P-MKK4 antibodies. P-MKK7 and P-MKK4 were immunoprecipitated at 10, 20, 30 min and at 5 h of NMDA stimulation, from nuclear and cytoplasmic fractions.
The results confirmed that MKK7 was powerfully activated by NMDA in the nuclear fraction, with maximal activation after 10 min (Figure 4e, g), as revealed previously by Western blot (Figure 4c, d). MKK7 activation by NMDA returned to basal levels by 40 min (mirroring the JNK response reported in this model).
Immunoprecipitated P-MKK7 was quantified with densitometry analysis and the maximal activation was four-fold higher than the basal (control) level (Figure 4g). The immunoprecipitation experiments also confirmed that MKK4 activation was unaffected by NMDA at all time points and that it remained almost exclusively cytoplasmic (Figure 4f and h). Thus, NMDA selectively activated MKK7, whereas MKK4 was unaltered.
Effect of D-JNKI1 on P-MKK7 and P-MKK4 activation
We tested whether D-JNKI1 (2 μM as in all the presented experiments) affects NMDA-induced activation/phosphorylation of the two kinases. We examined MKK7 activation after 10 min NMDA stimulation as at this time point MKK7 phosphorylation was maximal. Pretreatment with D-JNKI1 did not affect the level of P-MKK7 (Figure 4i). We also analyzed the effect of D-JNKI1 effect on P-MKK4: as shown before MKK4 presented its maximal activation in control condition. As for MKK7, D-JNKI1 did not affect the phosphorylation of MKK4 (Figure 4j).
Role of IB1/JIP-1 in excitotoxicity and the effect of D-JNKI1
We investigated IB1/JIP-1 modifications in NMDA-induced death and the inhibitory action on it of D-JNKI1.
Immunofluorescence showed that in the control condition, IB1/JIP-1 was almost exclusively in axons and as weak spots in the neuronal soma (including some nuclei) (Figure 7a), whereas after NMDA treatment it became strongly nuclear in the first hour of treatment (Figure 7c). D-JNKI1 pretreatment prevented the nuclear translocation but did not preserve the axonal staining characteristic of control neurons (Figure 7e).
IB1/JIP-1 in excitotoxicity. IB1/JIP-1 immunofluorescence study (a–f): after 1 h of 100 μm NMDA cortical neurons are stained with IB1/JIP-1 antibody (a, c, e) and DAPI (b, d, f). In the control condition, IB1/JIP-1 is localized in dendrites and axons (arrowhead) and in the soma of cortical neurons, including some nuclei (a). After NMDA, the labelling becomes strongly nuclear (c, arrow) and the staining of neurites is limited (arrowhead). D-JNKI1 prevents nuclear translocation but does not preserve the labelling of dendrites and axons (e). Scale bar, 10 μm. Western blot study (g–h) of IB1/JIP-1 after a NMDA time course on total extracts (g) and fraction extracts (h) of cortical neurons. IB1/JIP-1 level is stable from 5 min to 1 h of NMDA, but decreases clearly at 5 h, when two bands are visible (g). Fraction separation analysis confirms the immunofluorescence data (h): in the control condition IB1/JIP-1 is distributed in both nuclear and cytoplasmic compartments, whereas after NMDA stimulation it increases in the nuclear fraction. In vitro JNK-competitive assay with recombinant activated JNK3 (i), using GST-IB1/JIP-1 as substrate as described previously.5, 6 The JBD20 peptide inhibits IB1/JIP-1 phosphorylation, whereas the control-mutated peptide JBDmut has no effect. Western blot study of D-JNKI1 effect on IB1/JIP-1 following NMDA: NMDA strongly reduces the IB1/JIP-1 level and the D-JNKI1 strongly prevents it (j). Similar results were obtained in seven independent experiments (±S.E.M.)
In Western blot analyses of total extracts, the IB1/JIP-1 level had already decreased by 30% after 10 min of NMDA treatment, and at 5 h it was very weak. Moreover, the single band seen after short times of treatment had segregated into two separate bands27, 28 (Figure 7g, h and j).
Using fraction separation, the resolution of IB1/JIP-1 was improved and it was confirmed that in the control condition it distributed in both nuclear and cytoplasmic fractions, whereas after excitotoxic (NMDA) stimulation IB1/JIP-1 increased in the nuclear fraction during the first 20 min before subsequently decreasing (Figure 7a–f, h).
In order to clarify IB1/JIP1 interaction with JNK, we performed in vitro (cell-free) kinase assays with GST-IB1/JIP1 fusion proteins as a substrate. Activated JNK3 protein kinase was incubated with [γ-33P]ATP, the GST-IB1/JIP1 fusion proteins and either with JBD20 or JBDmut for 30 min. As shown in Figure 7i, JNK3 phosphorylated IB1/JIP-1. The JBD20 peptide powerfully prevented this interaction, whereas the JBDmut28 peptide had no effect. D-JNKI1 is known to block JNK-mediated phosphorylation of IB1/JIP-1.8, 9, 28
We then investigated the neuroprotective effect of D-JNKI1 on IB1/JIP-1: it prevented 70% of its NMDA-induced decrease and diminished the segregation into upper and lower bands27, 28 (Figure 7j).
Until now 35 sites of phosphorylation are known on IB1/JIP1,29 but there is no specific phospho-antibody to study JNK-mediated JIP-1 phosphorylation, which occurs at Thr-103. To better assess changes in IB1/JIP-1 at 5 h of NMDA and to verify the protective effect of D-JNKI1, we did a proteomic analysis.
IB1/JIP-1 by proteomic analysis
In silver-stained two-dimensional (2D) gels IB1/JIP-1 was barely detectable (Figure 8a, see ring), but on 2D Western blots there was a complex IB1/JIP-1 staining pattern (Figure 8b–d). Blots from untreated neurons revealed up to six spots, showing that IB1/JIP-1 exists in a variety of isoforms and undergoes several post-translational modifications. The NMDA stimulus resulted in a decrease of IB1/JIP-1 immunoreactivity and the disappearance of the most acidic forms. A shift was observed towards more basic proteins and also towards forms of smaller Mw (suggesting a decrease in phosphorylation). Pretreatment with D-JNKI1 reduced these changes indicating that D-JNKI1 partially counteracts the effects of the NMDA, probably by blocking the action of JNK kinase on IB1/JIP-1.28, 30 The counteracting effect of D-JNKI1 was confirmed when silver-stained 2D gels of control, NMDA-stimulated and NMDA-stimulated D-JNKI1-treated neurons were compared with the computer software Melanie, using a heuristic clustering algorithm to classify similar gels in the same group by comparing the similarity of protein spots. The control gel and the NMDA plus D-JNKI1 inhibition gel were placed in the same group, confirming that D-JNKI1 must counteract the effects of NMDA.
2D gels for IB1/JIP-1. Different samples are separated on 2D gels and either stained by an ammoniacal silver method or immunostained after electric transfer to nitrocellulose. Panel a is an example of a silver-stained gel of a control samples. IB1/JIP-1 is barely visible (see the circle) in the 100 kD Mw region and between pH 3 and 5. Only the specific areas of Western blots are shown (six spots) from a control sample (b), after NMDA stimulation (c), or NMDA and D-JNKI1 exposure (d). The lower panels show the entire silver-stained gels for b, c and d. The pH range of isoelectrophocusing is shown on top of the gels, and the molecular weight range is indicated to the right. 2D gel results were obtained using pools of three different dishes and in three independent experiments
Discussion
In multiple nervous system acute pathological conditions, including stroke and traumatic brain injury, overactivation of NMDA receptors leads to neuronal injury, resulting in neurological impairments. A treatment for these conditions would have a massive impact in modern society and D-JNKI1 represents a new neuroprotective candidate for this aim. In fact, the D-JNKI1 peptide is able to completely prevent neuronal loss after NMDA treatment and to give powerful protection against cerebral ischemia.6 However, the molecular mechanisms that modulate this powerful neuroprotection remain partially unclear.
Excitotoxic neuronal death can show features of apoptosis or necrosis (or a combination of the two,31 depending on the duration and intensity of the insult and on the model used2, 32, 33). In the present model involving cortical neurons exposed to a high concentration of NMDA (100 μM), the neuronal death has generally been described as necrotic rather than apoptotic,31 and our present, albeit limited, observations support this view. The neurons initially swelled and presented strong staining with PI, with no clumped chromatin. The fact that caspase-3 was activated shows that apoptotic pathways were also activated, but the experiments with z-VAD.fmk and cycloheximide indicate that apoptotic signalling was by no means the only contributor to cell death.
We analyzed NMDA-induced neuronal death from the initial calcium entry to the effects on the JNK signalling pathway. We examined the Ca++ entry through NMDA receptor channels, using imaging experiments with Fura-2, to see whether the D-JNKI1 peptide interferes at the receptor level or further downstream in the death cascade. The peptide did not change Ca++ influx, implying that D-JNKI1 did not protect at this level, but more downstream.
In characterizing the NMDA-induced death pathway, we found that z-VAD.fmk and cycloheximide alone were ineffective against NMDA toxicity, but combined they conferred 60% protection on NMDA-treated cortical neurons at 24 h. This suggests that NMDA activated at least two neuronal death pathways: one is caspase-dependent and -independent of protein synthesis and the other dependent on protein synthesis. Interestingly, D-JNKI1 was neuroprotective even in the presence of cycloheximide, implying that the neuroprotection by D-JNKI1 affected both pathways. The action of D-JNKI1 on c-jun and related transcription factors is well known.6
Here, we focus our attention on the JNK targets which contain a JBD homology sequence and whose phosphorylation by JNK is blocked by JBD20 at the concentration of 40 μM in cell-free experiments.8, 9 Among the JBD20-dependent targets, four substrates (MADD/DENN, MKK4, MKK7 and IB1/JIP-1) may be potentially implicated in the NMDA stress response of cortical neurons.
These targets act at different levels in the JNK cascade. MKK4 and MKK7 are upstream of JNK and are both activators and substrates. In contrast, MADD and IB1/JIP1 are at a lower level, downstream of JNK.
MADD/DENN
Our data suggest that post-translational modification of MADD may be involved in the excitotoxic process. NMDA stress caused a strong degradation of MADD/DENN, and a nuclear translocation. This degradation appears to be mediated by the ubiquitin proteolytic pathway, as the two specific proteasome inhibitors lactacystin and MG132 prevented it, whereas the calpain inhibitor had no effect. D-JNKI1 prevented both MADD degradation, as did the proteasome inhibitors, and translocation inside the nucleus. Moreover, the JNK competitive assay on MADD demonstrated that D-JNKI1 prevented the JNK-mediated phosphorylation of MADD. Taken together, our data strongly suggest that inhibition of JNK-mediated phosphorylation of MADD prevents its entry into the proteasome pathway.
Thus, MADD/DENN post-translational modification may represent an important component in excitotoxicity, but further studies are needed to determine its role in the death process and to determine the meaning of its phosphorylation and translocation.
The two direct activators of JNK: MKK4 and MKK7
We studied the localization and the contribution of these two JNK activators in the response of JNK to excitotoxic stress. We showed that already in the control condition the two kinases present a different compartmentation in cortical neurons. MKK7 is present in both nuclear and cytoplasmic compartments, whereas MKK4 is confined to the cytoplasm. Following NMDA stimulation, MKK7 strongly translocated into the nuclei of dying neurons, whereas MKK4 did not change its localization. In order to better clarify the contribution of these two kinases to JNK activation in excitotoxicity, we also analyzed MKK7 and MKK4 activation/phosphorylation in basal and stress conditions.
MKK7 showed a minor activation/phosphorylation in the control condition, whereas after NMDA stimulation it was quickly and powerfully activated. In fact, after 10 min of NMDA stimulation MKK7 reached maximal activation, then dropped back to the basal level in 1 h, as did JNK.
In contrast, the basal activity of MKK4 was elevated in the control condition and did not respond with an increase in phosphorylation after excitotoxic stress.
As D-JNKI1 selectively prevents inter-molecular interactions between JNK and its JBD-containing targets, this can result in an inhibition of MKK4 and MKK7 activity. At the concentration (2 μM) used to prevent neuronal loss, we did not detect any effects of D-JNKI1 on the activities of MKK4 and MKK7, which was expected as we previously found no effect on the activity of JNK.6 This suggests that the protective effects of D-JNKI1 were due to its prevention of the activation of its downstream targets (such as MADD) rather than to inhibition of the feedback activation of MKK4 and MKK7. The lack of effect of D-JNKI1 on the activities of MKK4 and MKK7 may be due to the fact that the concentration used in our experiments (2 μM) was too low compared to the concentration (40 μM) used in cell-free competitive assays where MKK7 and MKK4 phosphorylation is inhibited by JBD20.8, 9 The significance of MKK4 and MKK7 phosphorylation by JNK is not known, but it clearly constitutes a positive feedback loop in the JNK cascade. Such feedback has been predicted to explain the occurrence of bistability and ultrasensitivity in the activation of JNK.34, 35
The scaffold protein IB1/JIP-1
IB1/JIP-1 under excitotoxic stimuli altered its subcellular localization and accumulated in the nuclei. A similar behavior was observed in hippocampal neurons.20
We established that JNK participates actively in the phosphorylation of IB1/JIP-1 and this consequently regulated its stability. Exposure to NMDA stress induced a large degradation of IB1/JIP-1. D-JNKI1 protected against NMDA-induced death by preventing IB1/JIP-1 phosphorylation and degradation. In fact, having determined the effect of the peptide in cell-free assays, we assume that D-JNKI1 blocks JNK-mediated phosphorylation of IB1/JIP-18, 9, 28 and prevents its degradation. We suggest here two possible roles for the IB1/JIP-1 scaffold, in addition to its widely accepted linking of the upstream signalling elements to JNK. First, it may play a major role in directing the activated cascade into a particular cellular compartment. Second, it may serve to ‘select’ some of the downstream substrates of JNK. By blocking the action of JNK on JBD20-dependent substrates, IB1/JIP-1 may be promoting its access to the JBD20-independent ones.
MKK7 and MKK4 as potential targets for regulating the JNK signalling pathway
It will be important to devise tools (e.g. siMKK7 and siMKK4 or new specific cell-permeable MKK7 and MKK4 inhibitor peptides) for preventing selectively MKK7 and MKK4 activation with the aim of determining the direct causality of JNK activation induced by different stress stimuli.
Materials and Methods
Cortical neuronal culture
Small pieces of cortex were dissected from the brains of 2-day-old rat pups, incubated with 200 U of papain for 30 min at 34°C and after trypsin inhibitor treatment (10 μg) were mechanically dissociated. Neurons were then plated at densities of approximately 1 × 106 cells/plate on dishes precoated with 25 μg/ml poly-D-lysine. For immunocytochemistry, neurons were plated at densities of 1 × 105 cells/coverslip on coverslips precoated with 50 μg/ml poly-D-lysine and 20 μg/ml laminin. The plating medium consisted of B27/Neurobasal (Life Technologies, Gaithersburg, MD, USA) supplemented with 0.5 mM glutamine, 100 U/ml penicillin and 100 μg/ml streptomycin. PI was used to identify dying neurons at a concentration of 2 μg/ml, for 5 min immediately before fixation. Experiments were performed after 11–13 days in culture, at which time the neurons had elaborate axonal and dendritic arbors and had formed many synapses. D-JNKI1 2 μM was added to the dishes 1 hour before NMDA treatment (100 μM). Cycloheximide (1 or 3 μg/ml, fluka) and z-VAD.fmk (50, 150 or 300 μM, Sigma-Aldrich, St Louis, MO, USA) were added to the cultures 1 hour before NMDA. For the degradation studies, the calpain inhibitor calpastatin was added at concentrations of 5 and 10 μM 60 min before the addition of NMDA. Different concentrations of the proteasome inhibitors lactacystin and MG132 (10, 20 μM) were also added to the dishes 1 hour before NMDA treatment.
LDH cytotoxicity assay
Neuroprotection was evaluated by an LDH assay. LDH released into the bathing medium 5–48 h after NMDA was measured using the Cytotox 96 non-radioactive cytotoxicity assay kit (Promega, WI, USA).
Cytoplasmic and nuclear fractionation and total protein extracts
After stimulation, cells were washed twice in ice-cold PBS +1 mM MgCl2, and lysed for 10 min in lysis buffer5 at 4°C. Samples were then centrifuged at 900 × g for 10 min at 4°C to separate the supernatants (cytoplasmic fractions) from the pellets (nuclear fractions).26 Nuclear pellets were washed once in the lysis buffer and centrifuged at 900 × g for 5 min at 4°C, after which the pellets were reconstituted in lysis buffer and sonicated. Samples were analyzed for protein determination and equal proportions of nuclear and cytoplasmic fractions were analyzed by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Total protein extracts were obtained by scraping cells in lysis buffer.6
Western blot analysis
Proteins were separated by 10–14% SDS-PAGE and transferred to a PVDF membrane. Incubation with primary antibodies was overnight at 4°C using: 1 : 10 000 anti-MKK7 (#M-86920, BD Transduction Laboratories, San Jose, CA, USA), 1 : 1000 anti-P-MKK7 (a generous gift from Andreas H Nelsbach, Cell Signaling Technology, Beverly, MA, USA), 1 : 5000 or 1 : 10 000 anti-MKK4 (#SC-964, Santa-Cruz Biotechnology, CA, USA and # 07-194 Upstate, Charlottesville, VA, USA), 1 : 2000 anti-P-MKK4 (no. 9151 Cell Signaling Technology, Beverly, MA, USA), 1 : 50 000 anti–IB1/JIP-1 (polyclonal IB1 antiserum was prepared against recombinant IB1 (aa 1280) and the antiserum was subsequently affinity purified36), 1 : 5000 anti-MADD (a generous gift from Dr. Ulrich Blank, INSERM U 699, Paris Cedex 18, France), 1 : 750 anti-caspase-3 antibody (#SC-1225 Santa-Cruz Biotechnology, CA, USA; which reacts with both pro-caspase-3 and the cleaved p12 product). Both P-antibodies, P-MKK7 and P-MKK4, are specific and recognize only the phosphorylated form of these proteins (they do not recognize cortical neuronal extracts dephosphorylated with alkaline phosphatase overnight). In a single experiment, a very large number of neurons were analyzed, providing very consistent results, and the blots were all normalized with respect to tubulin (#SC-8035 Santa-Cruz Biotechnology, CA, USA).
Immunoprecipitation
All steps were performed on ice. About 1 mg of cell extract was incubated with an equivalent of 180 μg Ig anti-P-MKK7 (see WB) or P-MKK4 (#9156, Cell Signaling Technology, Beverly, MA, USA) for 30 min at 4°C under agitation. Protein A beads were added in the same tubes and samples were incubated overnight at 4°C. The immunoprecipitate was washed four times with complete cold lysis buffer. The final pellet was dissolved in 40 μl 2 × electrophoresis buffer, loaded into electrophoresis gels and transferred to a PVDF membrane. To assess the activation of MKK4 and MKK7 compared to control conditions, membranes were incubated overnight with the anti-non-phosphorylated specific antibodies (MKK7: #559676, BD Pharmingen, San Jose, CA, USA; MKK4: #07–194, Upstate, Charlottesville, VA, USA).
Quantification
The quantification of Western blots and immunoprecipitations was performed using ImageQuant TL software (Amersham Biosciences) and was based on at least five independent experiments.
Immunocytochemistry
All immunocytochemical reactions were performed on cortical neurons plated on coverslips as described above. Cells were fixed in 4% paraformaldehyde in PBS for 30 min, and then incubated for 1 h with 10% serum in PBS-Triton 0.3%. Primary antibodies were diluted 1 : 200 (MKK7), 1 : 100 (MKK4, #SC-964 Santa-Cruz Biotechnology, CA, USA), 1 : 500 (MADD), 1 : 1000 (IB/JIP-1) in PBS-Triton 0.1% and incubated overnight at 4°C. After washing in PBS, cells were incubated with secondary fluorescein isothiocyanate (FITC)-anti mouse and FITC-anti rabbit antibodies (FI-2000 and FI-1000, Vector Laboratories Inc., Burlingame, CA, USA), 1 : 200 in PBS for 1.5 h. Finally, coverslips were mounted in Vectashield mounting medium for fluorescence with DAPI (H-1200, Vector Laboratories Inc., Burlingame, CA, USA).
Kinase assays
Before initiating the kinase reactions, 2 μCi [γ-33P]ATP and 40 μM of peptides (JBD20 or JBDmut) were added to the activated recombinant JNK2 or JNK3 (0.5 μg, Upstate Biotechnology) for 15 min. Kinase reactions (Bonny et al.,5) were performed for 30 min at 30°C using 1 μg of the GST-MADD or GST-IB1/JIP-1 fusion proteins. Samples were then analyzed by SDS-PAGE and autoradiography.
2D electrophoresis
Tissue was homogenized with 1:10 w/v IEF sample buffer (5 M urea+2 M thiourea 35 mM Tris+4% CHAPS+1% DTT, with a protease inhibitor cocktail (Sigma p2714)) followed by a protein determination. First, Zoom Strips pH 3–10, 8 cm long (ZOOM IPGRunner System, Invitrogen, Switzerland) were rehydrated with 8 M urea, 2% CHAPS, 0.5% ampholytes pH 3–10, 0.02% bromphenolblue and 20 mM DTT was added just before the use. For the second dimension, strips were incubated in equilibration buffer (6 M urea+1.5 M Tris-HCl pH 8.8+2% SDS+20% Glycerol v/v) with 200 mg DTT per 10 ml buffer (buffer A) and 250 mg iodoacetamide per 10 ml buffer (buffer B). Each channel was filled with buffer A for 15 min, and then buffer B for another 15 min. The second dimension was run on NuPage 4–12% Bis-Tris Zoom gel (Invitrogen). Proteins were visualized by silver staining37 or transferred to nitrocellulose. Gels were scanned with an Amersham Molecular Scanner and results were analyzed with Melanie 4.0 software (Genebio, Geneva, Switzerland).
Microscopy for Ca2+i measurements
Experiments were carried out on the stage of an inverted epifluorescence microscope (Carl Zeiss, Jena) and observed through a 40 × 1.3 NA oil-immersion Neofluar objective lens (Zeiss). Fluorescence excitation wavelengths were selected using a holographic monochromator (Polychrome II, Till Photonics, Planegg, Germany) and fluorescence was detected using a 12-bit cooled CCD camera (Micromax, Princeton Instruments, Trenton, NJ, USA). Acquisition and digitization of images, and time series were computer controlled using the software Metafluor (Universal Imaging, West Chester, PA, USA) running on a Pentium computer. The acquisition rate of ratio images was varied between 0.5 and 0.1 Hz. Once loaded with dye, cells were placed in a perfusion chamber designed for rapid exchange of perfusion solutions.38 Up to ∼25 individual neurons were simultaneously analyzed in the selected field of view. Ca2+i was measured using Fura-2 (Molecular Probes, Eugene, OR, USA) loaded into cells by incubation with 5 μM Fura-2 AM for 30 min at 37°C. Experiments were run in CO2/bicarbonate-buffered solutions (NaCl 135, KCl 5.4, CaCl2 1.3, MgSO4 0.8, NaH2PO4 0.78, NaHCO3 25, glucose 5, glycine 3 μM, bubbled with 5% CO2/95% air). Calibration of cytoplasmic signal was accomplished in situ at the end of some experiments as described previously.38 Fluorescence was sequentially excited at 340 and 380 nm and detected at >515 nm. Data are presented as means±S.E.M. and Student's t-test was used to assess statistical significance.
Abbreviations
- D-JNKI1:
-
D-retro-inverso form of c-Jun N-terminal kinase-inhibitor
- FITC:
-
fluorescein isothiocyanate
- GST:
-
glutathione S-transferase
- IB1/JIP-1:
-
JNK-interacting protein-1
- JBD:
-
JNK-binding domain
- JNK:
-
c-Jun N-terminal kinase
- LDH:
-
lactate dehydrogenase
- MADD/DENN:
-
MAPK-activating death domain-containing protein/differentially expressed in normal and neoplastic cells
- MAPK:
-
mitogen-activated protein kinase
- MKK4:
-
MAP kinase kinase 4
- MKK7:
-
MAP kinase kinase 7
- NMDA:
-
N-methyl-D-aspartate
- PI:
-
propidium iodide
- SDS-PAGE:
-
sodium dodecyl sulfate polyacrylamide gel electrophoresis
- z-VAD.fmk:
-
z-Val-Ala-Asp Fluoromethylketone
References
Brecht S, Kirchhof R, Chromik A, Willesen M, Nicolaus T et al. Specific pathophysiological functions of JNK isoforms in the brain. Eur J Neurosci 2005; 21: 363–377.
Borsello T, Croquelois K, Hornung JP, Clarke PGH . N-methyl-d-aspartate-triggered neuronal death in organotypic hippocampal cultures is endocytic, autophagic and mediated by the c-Jun N-terminal kinase pathway. Eur J Neurosci 2003; 18: 473–485.
Wang LH, Besirli CG, Johnson Jr EM . Mixed-lineage kinases: a target for the prevention of neurodegeneration. Annu Rev Pharmacol Toxicol 2004; 44: 451–474.
Tezel G, Chauhan BC, LeBlanc RP, Wax MB . Immunohistochemical assessment of the glial mitogen-activated protein kinase activation in glaucoma. Invest Ophthalmol Vis Sci 2003; 44: 3025–3033.
Bonny C, Oberson A, Negri S, Sauser C, Schorderet DF . Cell-permeable peptide inhibitors of JNK: novel blockers of beta-cell death. Diabetes 2001; 50: 77–82.
Borsello T, Clarke PGH, Hirt L, Vercelli A, Repici M et al. A peptide inhibitor of c-Jun N-terminal kinase protects against excitotoxicity and cerebral ischemia. Nat Med 2003; 9: 1180–1186.
Borsello T, Bonny C . Use of cell-permeable peptides to prevent neuronal degeneration. Trends Mol Med 2004; 10: 239–244.
Negri S . Role of the Scaffold Protein IB1 and IB2 in JNK Mediated Apoptosis of Insulin Secreting Cells PhD thesis, UNIL, Lausanne, 2002.
Negri S, Guenat S, Oberson A, Allaman-Pillet N, Schorderet DF, Bonny C . Jip-1/IB1 interference on JNK-targets. Swiss Apoptosis Meeting B20 2002.
Zhang Y, Zhou L, Miller CA . A splicing variant of a death domain protein that is regulated by a mitogen-activated kinase is a substrate for c-Jun N-terminal kinase in the human central nervous system. Proc Natl Acad Sci USA 1998; 95 (5): 2586–2591.
Kuan CY, Whitmarsh AJ, Yang DD, Liao G, Schloemer AJ et al. A critical role of neural-specific JNK3 for ischemic apoptosis. Proc Natl Acad Sci USA 2003; 100: 15184–15189.
Yang DD, Kuan CY, Whitmarsh AJ, Rincon M, Zheng TS et al. Absence of excitotoxicity-induced apoptosis in the hippocampus of mice lacking the Jnk3 gene. Nature 1997; 389: 865–870.
Del Villar K, Miller CA . Down-regulation of DENN/MADD, a TNF receptor binding protein, correlates with neuronal cell death in Alzheimer's disease brain and hippocampal neurons. Proc Natl Acad Sci USA 2004; 101: 4210–4215.
Miyoshi J, Takai Y . Dual role of DENN/MADD (Rab3GEP) in neurotransmission and neuroprotection. Trends Mol Med 2004; 10: 476–480.
Harper SJ, LoGrasso P . Signalling for survival and death in neurones: the role of stress-activated kinases, JNK and p38. Cell Signal 2001; 13: 299–310.
Papa S, Zazzeroni F, Bubici C, Jayawardena S, Alvarez K et al. Gadd45 beta mediates the NF-kappa B suppression of JNK signalling by targeting MKK7/JNKK2. Nat Cell Biol 2004; 6: 146–153.
Zhou G, Golden T, Aragon IV, Honkanen RE . Ser/thr protein phosphatase 5 (PP5) inactivates hypoxia-induced activation of an ASK-1/MKK-4/JNK-signaling cascade. J Biol Chem 2004; 279 (45): 46595–46605.
Ho DT, Bardwell AJ, Abdollahi M, Bardwell LA . Docking site in MKK4 mediates high affinity binding to JNK MAPKs and competes with similar docking sites in JNK substrates. J Biol Chem 2003; 278: 32662–32672.
Dickens M, Rogers JS, Cavanagh J, Raitano A, Xia Z et al. A cytoplasmic inhibitor of the JNK signal transduction pathway. Science 1997; 277: 693–696.
Whitmarsh AJ, Kuan CY, Kennedy NJ, Kelkar N, Haydar TF et al. Requirement of the JIP1 scaffold protein for stress-induced JNK activation. Genes Dev 2001; 15: 2421–2432.
Magara F, Haefliger JA, Thompson N, Riederer B, Welker E et al. Increased vulnerability to kainic acid-induced epileptic seizures in mice underexpressing the scaffold protein Islet-Brain 1/JIP-1. Eur J Neurosci 2003; 17: 2602–2610.
Dubinsky JM, Kristal BS, Elizondo-Fournier M . On the probabilistic nature of excitotoxic neuronal death in hippocampal neurons. Neuropharmacology 1995; 34: 701–711.
Behrens A, Sibilia M, Wagner EF . Amino-terminal phosphorylation of c-Jun regulates stress-induced apoptosis and cellular proliferation. Nat Genet 1999; 21: 326–329.
Gupta S, Campbell D, Derijard B, Davis RJ . Transcription factor ATF2 regulation by the JNK signal transduction pathway. Science 1995; 267: 389–393.
Negri S, Guenat S, Oberson A, Allaman-Pillet N, Schorderet DF, Bonny C . Jip-1/IB1 interference on JNK-targets. Swiss Apoptosis Meeting B20 2002.
Coffey ET, Hongisto V, Dickens M, Davis RJ, Courtney MJ . Dual roles for c-Jun N-terminal kinase in developmental and stress responses in cerebellar granule neurons. J Neurosci 2000; 20: 7602–7613.
Kim IJ, Lee KW, Park BY, Lee JK, Park J et al. Molecular cloning of multiple splicing variants of JIP-1 preferentially expressed in brain. J Neurochem 1999; 72: 1335–1343.
Allaman-Pillet N, Storling J, Oberson A, Roduit R, Negri S et al. Calcium- and proteasome-dependent degradation of the JNK scaffold protein islet-brain 1. J Biol Chem 2003; 278: 48720–48726.
D'Ambrosio C, Arena S, Fulcoli G, Scheinfeld MH, Zhou D et al. Hyperphosphorylation of JNK-interacting Protein 1, a protein associated with Alzheimer disease. Mol Cell Proteomics 2006; 5: 97–113.
Nihalani D, Wong HN, Holzman LB . Recruitment of JNK to JIP1 and JNK-dependent JIP1 phosphorylation regulates JNK module dynamics and activation. J Biol Chem 2003; 278: 28694–28702.
Portera-Cailliau C, Price DL, Martin LJ . Non-NMDA and NMDA receptor-mediated excitotoxic neuronal deaths in adult brain are morphologically distinct: further evidence for an apoptosis-necrosis continuum. J Comp Neurol 1997; 378: 88–104.
Ankarcrona M, Dypbukt JM, Bonfoco E, Zhivotovsky B, Orrenius S et al. Glutamate-induced neuronal death: a succession of necrosis or apoptosis depending on mitochondrial function. Neuron 1995; 15: 961–973.
Kajta M, Lason W, Kupiec T . Effects of estrone on N-methyl-D-aspartic acid- and staurosporine-induced changes in caspase-3-like protease activity and lactate dehydrogenase-release: time- and tissue-dependent effects in neuronal primary cultures. Neuroscience 2004; 123: 515–526.
Bagowski CP, Ferrell Jr JE . Bistability in the JNK cascade. Curr Biol 2001; 11: 1176–1182.
Kanda H, Miura M . Regulatory roles of JNK in programmed cell death. J Biochem (Tokyo) 2004; 136: 1–6.
Bonny C, Nicod P, Waeber G . IB1, a JIP-1-related nuclear protein present in insulin-secreting cells. J Biol Chem 1998; 273: 1843–1846.
Hochstrasser DF, Merril CR . ‘Catalysts’ for polyacrylamide gel polymerization and detection of proteins by silver staining. Appl Theor Electrophor 1988; 1: 35–40.
Chatton JY, Idle JR, Vogbo CB, Magistretti PJ . Insights into the mechanisms of ifosfamide encephalopathy: drug metabolites have agonistic effects on α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA)/Kainate receptors and induce cellular acidification in mouse cortical neurons. J Pharmacol Exp Ther 2001; 299 (3): 1161–1168.
Acknowledgements
Tiziana Borsello dedicates this work to Martina Giordano. This work was supported by grants from the Botnar Foundation and the Swiss National Science Foundation (FNRS) Grants 310000-107888, 3100A0-101696 and from the European Community's Sixth Framework Programme to the Specific Targeted Research Project stressprotect (LHSM-CT-2004-005310). Dr. Repici was supported by a Roche grant. We thank Dr. S Negri and S Guenat, E Bernardi, Vincent Mottier and Irene Riederer, for assistance, Dr. Andreas H Nelsbach, Cell Signaling Technology for kindly providing us the P-MKK7 antibody and Dr. Ulrich Blank for providing us the MADD/DENN antibody. We also thank Dr. Amar Abderrahmani and Dr. G De Simoni for critical comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by SA Lipton
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Centeno, C., Repici, M., Chatton, JY. et al. Role of the JNK pathway in NMDA-mediated excitotoxicity of cortical neurons. Cell Death Differ 14, 240–253 (2007). https://doi.org/10.1038/sj.cdd.4401988
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401988
Keywords
This article is cited by
-
Identification of novel proteins for lacunar stroke by integrating genome-wide association data and human brain proteomes
BMC Medicine (2022)
-
The selective disruption of presynaptic JNK2/STX1a interaction reduces NMDA receptor-dependent glutamate release
Scientific Reports (2019)
-
Quantitative proteomic analyses of dynamic signalling events in cortical neurons undergoing excitotoxic cell death
Cell Death & Disease (2019)
-
The Parkinson’s Disease-Linked Protein DJ-1 Associates with Cytoplasmic mRNP Granules During Stress and Neurodegeneration
Molecular Neurobiology (2019)
-
Attenuation of Glutamate-Induced Excitotoxicity by Withanolide-A in Neuron-Like Cells: Role for PI3K/Akt/MAPK Signaling Pathway
Molecular Neurobiology (2018)