Abstract
Tumour necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL) exhibits potent antitumour activity upon systemic administration in mice without showing the deleterious side effects observed with other apoptosis-inducing members of the TNF family such as TNF and CD95L. TRAIL may, thus, have great potential in the treatment of human cancer. However, about 60% of tumour cell lines are not sensitive to TRAIL. To evaluate the mechanisms of tumour resistance to TRAIL, we investigated hepatocellular carcinoma (HCC) cell lines that exhibit differential sensitivity to TRAIL. Pretreatment with chemotherapeutic drugs, for example, 5-fluorouracil (5-FU), rendered the TRAIL-resistant HCC cell lines sensitive to TRAIL-induced apoptosis. Analysis of the TRAIL death-inducing signalling complex (DISC) revealed upregulation of TRAIL-R2. Caspase-8 recruitment to and its activation at the DISC were substantially increased after 5-FU sensitisation, while FADD recruitment remained essentially unchanged. 5-FU pretreatment downregulated cellular FLICE-inhibitory protein (cFLIP) and specific cFLIP downregulation by small interfering RNA was sufficient to sensitise TRAIL-resistant HCC cell lines for TRAIL-induced apoptosis. Thus, a potential mechanism for TRAIL sensitisation by 5-FU is the increased effectiveness of caspase-8 recruitment to and activation at the DISC facilitated by the downregulation of cFLIP and the consequent shift in the ratio of caspase-8 to cFLIP at the DISC.
Similar content being viewed by others
Introduction
Apoptosis is essential to maintain tissue homeostasis. Impaired apoptosis has been implicated in many human diseases, including tumours. TNF-related apoptosis-inducing ligand (TRAIL), also called Apo-2 ligand (Apo-2L), has been shown to kill various tumour cell lines in vitro. Interestingly, TRAIL induced apoptosis in about 40% of tumour cell lines, while the remaining tumour cell lines and most normal cells were resistant.1, 2, 3 Systemic administration of TRAIL suppressed human tumour growth in SCID mice and non-human primates without being toxic to normal tissue.2, 4, 5 Conflicting results have been published with regard to the sensitivity of normal human hepatocytes to TRAIL exposure.6 However, recent studies suggested that liver toxicity in humans is unlikely.7, 8 Thus, TRAIL may serve as a novel therapeutic drug for the treatment of cancer. Therefore, it is important to understand the mechanisms involved in resistance and sensitisation to TRAIL-induced apoptosis.
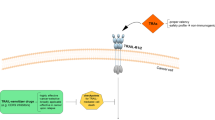
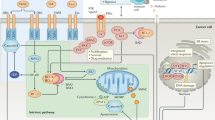
TRAIL interacts with five distinct receptors. TRAIL induces apoptosis upon binding to TRAIL-R1 (DR4)9, 10 and TRAIL-R2 (DR5/TRICK2/Killer).11, 12, 13, 14, 15 These receptors transmit a caspase-activating death signal due to the presence of a cytoplasmic death domain (DD).16, 17 In contrast, neither TRAIL-R3 (DcR1/TRID/LIT)12, 15, 18, 19, 20 nor TRAIL-R4 (DcR2/TRUNDD)21 can mediate apoptosis due to complete or partial absence of an intracellular DD, respectively. Osteoprotegerin (OPG) is a soluble receptor reported to bind OPG ligand (OPGL/RANKL/TRANCE/ODF) and TRAIL.22 The molecular events leading to apoptosis induction via TRAIL have recently been analysed. Crosslinking of the TRAIL receptors leads to the formation of a death-inducing signalling complex (DISC).23 The death adaptor protein FADD (MORT1) and the proteolytic enzymes caspase-8 and -10 are recruited to the TRAIL DISC.24, 25, 26, 27, 28 In a homotypic interaction, the DD of FADD binds to the DD of the TRAIL receptors. The death effector domain (DED) of FADD in turn interacts with the DED of procaspase-8, which is thereby recruited to the TRAIL DISC. Procaspase-8 is proteolytically cleaved and activated at the DISC. Activated caspase-8 then initiates the apoptosis executing caspase cascade.23
For successful cancer treatment, it is important to understand the biochemical mechanisms that distinguish TRAIL-resistant from TRAIL-sensitive cells. Hepatocellular carcinoma (HCC) is one of the most common carcinomas. Therapeutic options are very limited because of chemotherapy resistance of this tumour type. A number of studies have shown that cotreatment with chemotherapeutic agents and irradiation results in sensitisation of TRAIL-resistant tumour cell lines.29, 30, 31, 32, 33 However, the mechanisms leading to TRAIL sensitivity are controversial.5, 34, 35, 36, 37, 38, 39, 40, 41 Upregulation of the apoptosis-inducing TRAIL receptors after treatment with 5-fluorouracil (5-FU) has been implicated in sensitising human leukaemic and glioma cells.31, 35 Bernard et al.42 presented data supporting the role of TRAIL-R3 and TRAIL-R4 as putative decoy receptors, preventing apoptosis. Overexpression of cFLIPL and cFLIPS inhibited TRAIL-induced apoptosis.43 In other studies, no correlation with cFLIP expression and sensitivity versus resistance in TRAIL-induced apoptosis was found.44 Thus, the role of the TRAIL DISC and cFLIP in sensitisation for TRAIL-induced apoptosis is still unclear.45
To dissect the molecular mechanisms of TRAIL-induced apoptosis, we analysed the different biochemical levels at which TRAIL sensitivity versus resistance of HCC cell lines could be regulated. We show that constitutive resistance of HCC cell lines is not mediated by TRAIL-R3 or TRAIL-R4. Our data indicate that in hepatocytes sensitisation with 5-FU is mediated through enhanced caspase-8 recruitment and activation at the DISC.
Both cFLIPL and cFLIPS are downregulated by 5-FU treatment and specific downregulation of cFLIP by small interfering RNA (siRNA) in Hep3b cells sensitises for TRAIL-induced apoptosis. Downregulation of cFLIP results in an increase of the ratio of caspase-8 to cFLIP within the DISC. The observed changes synergise in 5-FU-mediated sensitisation to TRAIL-induced apoptosis.
Results
TRAIL induces apoptosis in the human HCC cell line HepG2 but not in HepG2-TR or Hep3b
Although TRAIL has been reported to induce apoptosis in a variety of tumour cell types including HCC cell lines,46 the molecular mechanisms that regulate sensitivity versus resistance of tumour cells to TRAIL-induced apoptosis remain poorly defined. We identified two cell lines that exhibit a pronounced difference in TRAIL sensitivity, TRAIL-resistant Hep3b cells and TRAIL-sensitive HepG2 cells. In addition, we established a TRAIL-resistant variant of HepG2 cells, HepG2-TR cells, obtained by in vitro selection for TRAIL resistance. HepG2 cells undergo TRAIL-induced apoptosis in a dose-dependent manner (Figure 1). In contrast, treatment of HepG2-TR and Hep3b cells with TRAIL did not result in significant apoptosis induction. At a concentration of 1?μg/ml TRAIL less than 15% of the cells underwent apoptosis, while more than 90% of HepG2 cells were killed with 100?ng/ml TRAIL. We have, thus, identified a model system that serves to study the mechanisms of TRAIL resistance in HCC cells.
Dose dependency of TRAIL-induced apoptosis in various HCC cell lines. Different HCC cell lines HepG2 (○), HepG2-TR (Δ) and Hep3b (□) were treated with 12–1000?ng/ml TRAIL and analysed after 24?h for subdiploid DNA content.72 Data are shown as the percent apoptosis with S.D. (three wells each condition). Three independent experiments with similar results have been performed
5-FU and other chemotherapeutic drugs sensitise Hep3b and HepG2-TR cells for TRAIL-induced apoptosis
We were interested in studying mechanisms that result in TRAIL resistance and how TRAIL resistance can be broken. We tested whether chemotherapeutic drugs influence TRAIL sensitivity of HCC cell lines. We incubated Hep3b cells with different chemotherapeutic drugs before the addition of TRAIL (Figure 2a). We had determined subtoxic concentrations of the different chemotherapeutic drugs on the resistant cell lines (data not shown). The most striking synergistic effect was observed when Hep3b or HepG2-TR cells were cotreated with 5-FU and TRAIL, while the effect was less pronounced in the cases of pretreatment with the other chemotherapeutic drugs tested (Figure 2a). In approximately 50% of Hep3b cells, apoptosis was induced after 24?h, while both reagents alone induced apoptosis in less than 5% of cells. In addition, we found that cotreatment with 5-FU could further sensitise the constitutively sensitive HepG2 cells for TRAIL-induced apoptosis (Figure 2b). This cellular system at hand, we determined the molecular level at which 5-FU-mediated sensitisation to TRAIL-induced apoptosis occurred.
Sensitisation of Hep3b cells for TRAIL-induced apoptosis with chemotherapeutic drugs. (a) Hep3b cells (black bars) and HepG2-TR cells (grey bars) were treated with different chemotherapeutic drugs at the indicated concentrations for 24?h in the presence or absence of 1?μg/ml TRAIL. Apoptosis was determined by subdiploid DNA content. (b) HepG2 cells were treated with different concentrations of TRAIL in the presence (Δ) or absence (○) of 5-FU (25?μg/ml). After 24?h, cell survival was analysed by MTT assay as described in Materials and methods. Data are shown as the mean±S.D. of three independent experiments
Drug-induced upregulation of TRAIL death receptors is p53 independent
Upregulation of TRAIL death receptors, downregulation of nonapoptosis-inducing TRAIL receptors or a combination of both could result in enhanced sensitivity for TRAIL-induced apoptosis. We, therefore, examined surface expression of the different TRAIL receptors, and CD95 as a control, on all three cell lines before and after treatment with 5-FU (Figure 3). In accordance with our previous studies,47 we found that p53-deficient Hep3b cells did not express CD95 and that treatment with 5-FU did not result in its upregulation, while CD95 upregulation was readily observed in p53-wildtype HepG2 and HepG2-TR cells. In TRAIL-resistant Hep3b and HepG2-TR cells, TRAIL-R1 was expressed at lower levels than in the TRAIL-sensitive HepG2, and only in HepG2-TR cells TRAIL-R1 was upregulated by 5-FU treatment. Thus, TRAIL-R1 upregulation potentially contributes to increased sensitivity of HepG2-TR cells to TRAIL-induced apoptosis upon 5-FU treatment. TRAIL-R2 was expressed at comparable levels on all three cell lines. Upon treatment with 5-FU, TRAIL-R2 was not only upregulated in HepG2 and HepG2-TR cells but also in the p53-deficient Hep3b cells. TRAIL-R3 was only marginally detected on Hep3b, HepG2-TR and HepG2 cells, while TRAIL-R4 was apparent and increased upon 5-FU treatment. HepG2-TR and HepG2 cells also expressed TRAIL-R3 after treatment with 5-FU. Taken together, the chemotherapeutic drug 5-FU led to the upregulation of TRAIL-R2 in all cell lines and, to a lesser degree, of TRAIL-R1 in the HepG2-TR and HepG2 cells. Thus, increased surface expression of apoptosis-inducing TRAIL receptors could contribute to the increased sensitivity of the examined cell lines to TRAIL-induced apoptosis.
Comparison of TRAIL receptor surface expression of resistant and sensitive cell lines with and without 5-FU pretreatment. FACS analysis of surface expression of TRAIL-R1 to TRAIL-R4 and CD95 with (solid bold line) and without (filled line) 5-FU (100?μg/ml) treatment for 16?h as compared to an isotype-matched control mIgG1 mAb (dashed line). To exclude unspecific staining of dead cells, 1 × 104 7-AAD-negative cells were counted. Staining of isotype-matched control mIgG1 mAb was comparable with or without incubation with 5-FU (data not shown). The data represent the results of an experimental setup, which was repeated at least three times
TRAIL-R1- and TRAIL-R2-triggered pathways contribute to drug-induced sensitisation of HCC cell lines to TRAIL-induced apoptosis
Next, we tested whether resistance to TRAIL was mediated by the putative decoy function of the nonapoptosis-inducing receptors TRAIL-R3 and TRAIL-R4 or whether it was due to the blockage of apoptotic signalling of the receptors TRAIL-R1 and TRAIL-R2. Therefore, we specifically triggered TRAIL-R1 and TRAIL-R2, either alone or in combination, in the absence or presence of 5-FU (Figure 4). This allowed us to investigate whether putative decoy receptors were responsible for the resistance of Hep3b cells, since the antibodies would only stimulate specific receptors irrespective of the presence of TRAIL-R3 and TRAIL-R4. In TRAIL-resistant Hep3b cells, significant apoptosis induction was neither possible through triggering of TRAIL-R1 nor TRAIL-R2 alone, yet concomitant triggering of TRAIL-R1 and TRAIL-R2 resulted in about 10% specific apoptosis induction comparable to what was seen with TRAIL at high doses. Upon incubation with 5-FU, however, cell death was readily induced either by triggering TRAIL-R1, TRAIL-R2 or both receptors in combination (Figure 4). Resistance upon triggering TRAIL-R1 and TRAIL-R2 in the absence of 5-FU and sensitisation in its presence demonstrates that the nonapoptosis-inducing TRAIL-R3 and TRAIL-R4 are not necessary for the observed TRAIL resistance of Hep3b cells. Comparable results were obtained for HepG2-TR cells (data not shown). Interestingly, although TRAIL-R1 was not upregulated on Hep3b cells upon 5-FU treatment (Figure 3), the cells became more susceptible to the induction of apoptosis by triggering TRAIL-R1. This indicates that apart from receptor upregulation also intracellular events are involved in 5-FU-mediated sensitisation.
Induction of apoptosis by agonistic TRAIL-R1 and TRAIL-R2 antibodies after sensitisation of TRAIL-resistant Hep3b cells with 5-FU. TRAIL-resistant Hep3b cells were treated with monoclonal agonistic antibodies against TRAIL-R1 and TRAIL-R2 and a combination thereof. After 24?h, cell survival was determined by MTT assay as described in Materials and methods. Data are shown as the percent viability of control with S.D. (three wells each condition). Three independent experiments with similar results have been performed
Upregulation of TRAIL-R1 and TRAIL-R2 is not essential for sensitisation to TRAIL-induced apoptosis
For tumour necrosis factor (TNF), it has been shown that cells need to bind this cytokine directly for only minutes in order to achieve the full cytotoxic effect. Thus far, we had incubated HCC cells in the presence and absence of TRAIL with 5-FU for a 24?h time period. In order to test whether additional TRAIL needs to be present at later time points during sensitisation, or whether already cell-bound TRAIL was sufficient to induce apoptosis, we tested 5-FU sensitisation after removing all noncell-bound TRAIL after an initial incubation time of 25?min. If receptor upregulation alone would be responsible for 5-FU-mediated sensitisation, then cells from which unbound TRAIL had been removed should not undergo apoptosis upon 5-FU treatment. If, however, receptor upregulation was not responsible for sensitisation but rather an intracellular mechanism, then removal of unbound TRAIL should not affect 5-FU sensitisation. We took advantage of this distinction to determine whether increased surface expression of TRAIL-R1 and TRAIL-R2 were, in fact, necessary for the sensitising effect of 5-FU. TRAIL-resistant Hep3b cells were incubated in the presence of 1?μg/ml TRAIL for 25?min. Subsequently, unbound TRAIL was removed by thoroughly washing the cells before new medium with or without 5-FU was added for 24?h (Figure 5). As a control, Hep3b cells were treated with 5-FU in the presence of 1?μg/ml TRAIL throughout the 24?h period. Only small differences in TRAIL-induced apoptosis were observed between 5-FU-sensitised cells when unbound TRAIL was removed and not replaced and 5-FU-sensitised cells that were further incubated in the presence of TRAIL. Comparable results were obtained for HepG2-TR cells (data not shown). Thus, upregulation of the apoptosis-inducing TRAIL receptors by 5-FU treatment might contribute but is not the decisive factor for 5-FU sensitisation. Instead, these findings clearly point towards a mechanism of sensitisation that either takes place intracellularly or at the level of DISC formation.
Upregulation of the death-inducing TRAIL receptors is not essential for sensitisation of resistant cell lines. Hep3b cells were incubated with TRAIL at 1?μg/ml for 25?min and then washed five times with the medium. Cells were then cultured either in the medium containing 100?μg/ml 5-FU in the absence (open bars) or in the presence of TRAIL at 1?μg/ml (black bars). Control cells were washed five times without any addition of TRAIL (grey bars). After 24?h, apoptosis was analysed by subdiploid DNA content.72 Data are shown as the percent apoptosis with S.D. (three wells each condition). One of three experiments with comparable results is shown
Decrease of cFLIPL and cFLIPS expression and increased caspase-8 activation after 5-FU and TRAIL cotreatment
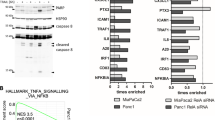
To test changes at the intracellular level, we stimulated Hep3b cells in the presence and absence of 5-FU with TRAIL for different time periods and subsequently analysed cell lysates for caspase-8 cleavage (Figure 6a). In Hep3b cells cotreated with 5-FU and TRAIL, caspase-8 cleavage was detected already after 1?h peaking after 2?h with the active p18 subunit detected. After 4?h of apoptosis induction by TRAIL, detection of the p18 and p43/p41 subunits of caspase-8 was reduced most likely due to rapid degradation. Remarkably, cleavage of procaspase-8 was nearly complete after 8?h. In marked contrast, only minimal caspase-8 cleavage and no active p18 subunit could be detected in Hep3b cells that were treated with TRAIL in the absence of 5-FU (Figure 6a). Caspase-8 activation was shown to be inhibited by cFLIPL and/or cFLIPS.48 In addition, cFLIPL overexpression has been shown to block TRAIL-induced apoptosis in transformed keratinocytes.43, 49 To test whether cFLIP downregulation by 5-FU treatment may contribute to the intracellular event that is responsible for the observed TRAIL sensitisation, we determined expression levels of cFLIP depending on treatment with 5-FU. We found that 5-FU treatment resulted in a marked reduction of cFLIPL and cFLIPS in Hep3b cells (Figure 6b). Already 2?h after onset of 5-FU treatment, both cFLIP isoforms were greatly reduced in the cell lysates, while the other DISC components were expressed at constant levels or, as in the case of FADD, only at slightly reduced levels (Figure 6c). This correlation between TRAIL sensitisation of Hep3b cells and downregulation of both cFLIP isoforms suggested the involvement of cFLIP downregulation in 5-FU sensitisation to TRAIL-induced apoptosis.
Rapid caspase-8 cleavage induced by TRAIL after 5-FU pretreatment of TRAIL-resistant cells, which markedly decreased protein levels of cFLIPL and cFLIPS. Hep3b cells were left untreated or were pretreated with 100?μg/ml 5-FU for 12?h before TRAIL was added for the indicated time points. Subsequently, cells were lysed and 20?μg of cell lysates were analysed by 12% SDS-PAGE and Western blot for caspase-8 (a) and cFLIP (b). The same blots were incubated with anti-ERK1 as a protein loading control. (c) To compare protein expression of the three cell lines in the absence and presence of 100?μg/ml, 5-FU cells were lysed after 16?h 5-FU treatment and 20?μg of cell lysates were analysed by 12% SDS-PAGE and Western blot for caspase-8, FADD and cFLIP. The same blots were incubated with anti-ERK1 as a protein loading control. These blots illustrate one representative Western blot of at least three
Enhanced DISC formation in TRAIL-sensitive and -sensitised cell lines after treatment with 5-FU
To compare directly the TRAIL-sensitive HepG2 cells with TRAIL-resistant Hep3b cells, we immunoprecipitated TRAIL DISCs from identical numbers of untreated cells next to each other. Although TRAIL-R1 and TRAIL-R2 are expressed at similar levels on the cell surface, caspase-8, FADD and also cFLIPS and cFLIPL recruitment to the TRAIL DISC is much more pronounced in the TRAIL-sensitive cell line than in the TRAIL-resistant cell line (Figure 7a).
Enhanced caspase-8 and constant levels of cFLIP recruitment revealed by native TRAIL DISC analysis after sensitisation with 5-FU. (a) HepG2 and Hep3b cells were then either stimulated with 1?μg/ml biotinylated TRAIL for 15?min (+) or left unstimulated (−) before cell lysis. TRAIL was added to the lysates in the unstimulated control. Resulting protein complexes bound to biotinylated TRAIL were analysed for the presence of TRAIL-R1, TRAIL-R2, caspase-8, FADD/MORT1 and cFLIP by SDS-PAGE and Western blot following affinity precipitation with Neutravidin beads. (b) HepG2, HepG2-TR and Hep3b cells were left untreated or were preincubated for 12?h with 5-FU (100?μg/ml), then DISC analysis was performed as described above. Three independent experiments with similar results have been performed
Enhanced caspase-8 activation and downregulation of cFLIP in cell lysates after treatment with 5-FU suggested an important function of the DISC. We therefore investigated changes induced by 5-FU in the formation and/or composition of the DISC formed upon TRAIL stimulation. We analysed the native TRAIL DISC of all three cell lines with and without pretreatment with 5-FU for 12?h (Figure 7b). This analysis showed that 5-FU treatment resulted in markedly enhanced DISC formation in the primarily resistant Hep3b and HepG2-TR cells and, to a lesser degree, also in sensitive HepG2 cells. In accordance with the changes in receptor surface expression, 5-FU treatment enhanced the recruitment of TRAIL-R2 to the DISC. The increased amount of TRAIL death receptor correlated with increased procaspase-8 recruitment, while the p43/41 levels were only slightly elevated. Interestingly, the increasing procaspase-8 recruitment was not paralleled to the same extent by an increase in FADD recruitment. The most surprising finding was that, although cFLIPS and cFLIPL was significantly reduced in the lysates after 16?h of 5-FU treatment (Figure 6c), cFLIPL was recruited to the TRAIL DISC in similar amounts in all three cell lines whether preincubated with 5-FU or not, while cFLIPS was only detected in the TRAIL DISC of the untreated sensitive HepG2 cells (Figure 7b). Interestingly, the DISC of the TRAIL-resistant HepG2-TR cells contained cFLIP at a level comparable to the one in the TRAIL DISC of the TRAIL-sensitive HepG2 cells. These results suggest that DISC formation is enhanced after sensitisation with 5-FU and that the effectiveness of DISC formation and caspase-8 recruitment is the determining difference between resistance and sensitivity in these cell lines.
Specific downregulation of cFLIP by siRNA is sufficient to sensitise Hep3b cells for TRAIL-induced apoptosis
Upregulation of TRAIL death receptors by 5-FU was not necessary for TRAIL sensitisation of hepatoma cell lines. Thus an intracellular event must be crucial. 5-FU treatment resulted in rapid downregulation of cFLIPL and cFLIPS. To test whether cFLIP downregulation alone might be sufficient to mediate TRAIL sensitisation, we transfected Hep3b cells with cFLIP-targeting siRNA vectors.50 Our results show that transfection of Hep3b cells with the cFLIP-targeting siRNA vector results in the downregulation of cFLIP (Figure 8a) and that these cells are sensitised for TRAIL-induced apoptosis (Figure 8b and c). Thus downregulation of cFLIP is sufficient to sensitise resistant tumour cells for TRAIL-induced apoptosis.
Specific downregulation of cFLIP by siRNA is sufficient to sensitise Hep3b cells for TRAIL-induced apoptosis. (a) Hep3b cells were transfected with a vector encoding for GFP and cFLIP siRNA or control siRNA. After 48?h, cells were sorted and GFP-positive cells were lysed and examined for cFLIP expression by Western blot analysis. (b) Cells were transfected with the respective vectors encoding for GFP only, control siRNA and cFLIP siRNA. After 48?h, cells were incubated with LZ-TRAIL (1?μg/ml) and GFP-positive cells were analysed by FACS at the indicated time points. Three independent experiments with similar results have been performed. (c) At 2?h after incubation with (1?μg/ml) LZ-TRAIL, same cell populations as described in (b) were stained for active caspase-3 (PE) and DAPI, and examined under the fluorescence microscope
Discussion
In the present study, we investigated the mechanisms of 5-FU sensitisation for TRAIL-induced apoptosis of HCC cells. Two TRAIL-resistant cell lines and a TRAIL-sensitive cell line showed similar surface expression of the TRAIL receptors. Interestingly, surface expression of TRAIL-R4 could be detected in all three cell lines. TRAIL-R3 and TRAIL-R4 have been suggested to function as apoptosis-inhibitory receptors, as ectopic overexpression has been shown to inhibit TRAIL-induced apoptosis.18, 19 However, in more recent publications no correlation between the expression of the putative TRAIL decoy receptors and TRAIL sensitivity was found in various cellular systems.34, 35 To test whether the sensitising effect of 5-FU was independent of TRAIL-R3 and TRAIL-R4 expression, we specifically stimulated TRAIL-R1 and TRAIL-R2 with agonistic antibodies. The direct stimulation of TRAIL-R1 or TRAIL-R2 alone and in combination failed to induce apoptosis in TRAIL-resistant cells. However, after sensitisation with 5-FU direct stimulation of TRAIL-R1 or TRAIL-R2 resulted in apoptosis. Therefore, TRAIL-R1 or TRAIL-R2 triggering alone or in combination is insufficient to induce apoptosis in TRAIL-resistant cells demonstrating that constitutive resistance of HCC cells is independent of TRAIL-R3 and TRAIL-R4 expression. Furthermore, sensitisation to TRAIL-induced apoptosis after 5-FU treatment was observed for both death-inducing TRAIL receptors, although TRAIL-R1 upregulation was marginal after 5-FU treatment in Hep3B cells.
Analysis of TRAIL receptor surface expression revealed that, concomitantly with sensitisation after treatment with 5-FU, TRAIL-R2 was upregulated on both resistant cell lines.51 Since Hep3b cells lack functional p53, our data confirmed that TRAIL-induced apoptosis is independent of the p53 status.52, 53, 54, 55, 56 In addition, the upregulation of TRAIL-R2 by 5-FU in the p53-negative Hep3b cells shows that TRAIL-R2 can be efficiently upregulated (Figure 3) and expressed (Figure 4) in the absence of functional p53.54
Recently, it has been shown that in a number of cell types TRAIL-induced apoptosis requires Bax-dependent release of Smac/DIABLO to achieve inactivation of the inhibitor of apoptosis proteins (IAPs).57, 58, 59, 60 To test whether sensitisation in HCC cells to TRAIL-induced apoptosis is a receptor-proximal event or whether it is regulated further downstream in the apoptotic signalling cascade, we examined the kinetics of cleavage of the initiator caspase-8. In contrast to resistant cells, we found massive and early caspase-8 cleavage in the sensitised cells with a maximum detected in the lysates after 1–2?h. Next, we tested whether caspase-8 activation was directly activated at the DISC to discriminate caspase-8 activation secondary to a mitochondrial amplification loop.30, 61 Postmitochondrial changes as described by Leverkus et al.58 were not observed in these cell lines, as we found no change in the expression of IAPs after incubation with 5-FU (data not shown). However, the HCC cells sensitised with 5-FU recruited massive caspase-8 at the native DISC already after 15?min. Therefore, increased caspase-8 activation at the DISC is a decisive initiator for the greatly increased apoptosis of these sensitised cells.
These results suggest that sensitisation by 5-FU for TRAIL-induced apoptosis is a receptor-proximal event. Upregulation of the death-inducing TRAIL receptors after treatment with 5-FU implicated an important role in the process of sensitisation of hepatoma cells. Interestingly, we found a comparable degree of apoptosis independent of whether unbound TRAIL was removed from the supernatant of resistant cells before sensitisation with 5-FU or was carried out in the continuous presence of TRAIL. Thus, additional binding of TRAIL to the upregulated TRAIL receptors did not result in significant additional apoptosis. Hence, the main mechanism for sensitisation occurs independent of the modulation of TRAIL receptor surface expression. Interestingly, FADD levels at the DISC remain relatively unchanged, while there is an upregulation of TRAIL-R2 together with increased caspase-8 recruitment.
The protein cFLIP has been shown to act as an inhibitor of TRAIL-induced apoptosis in other cellular systems.43, 62 Overexpression of cFLIPL or cFLIPS has resulted in the inhibition of the CD95 and TRAIL DISC.62, 63, 64 While cFLIPS blocks cleavage of procaspase-8, cFLIPL inhibits further processing of the intermediate cleavage products (p43/41) into the catalytically active caspase-8 (p18/p10).48, 62, 65 Recently, it has been suggested that cFLIPL at physiological levels enhances CD95-induced apoptosis and only overexpression of cFLIPL blocks CD95-induced apoptosis,45 while this has not been shown for TRAIL. In contrast to other known components of the DISC, for example, caspase-8, the expression levels of cFLIPL and cFLIPS in the cytosol were markedly reduced after 5-FU treatment. We precipitated the native TRAIL DISC in sensitive and resistant HCC cells before and after sensitisation with 5-FU to investigate whether the changes in the lysates were reflected by less efficient or absent cFLIP recruitment. DISC analysis demonstrated that both isoforms of cFLIP bind to the TRAIL DISC. Surprisingly, at the DISC level only a minimal decrease of cFLIPL could be observed, while cFLIPS was only efficiently precipitated in the TRAIL-sensitive cell line and could not be detected after treatment with 5-FU. However, procaspase-8 recruitment was enhanced after sensitisation to TRAIL-induced apoptosis in all cell lines, which was paralleled by massive caspase-8 cleavage in the cytosol. At the same time, processed p43/p41 caspase-8 were only slightly increased in the DISC, suggesting a high turnover of caspase-8 at the TRAIL DISC. In contrast, cFLIPL-mediated inhibition of caspase-8 activation at the DISC can be recognised by an accumulation of p43/p41.48 Thus, the appearance of less p43/p41 in comparison to p55/p53 upon treatment with 5-FU supports a function for cFLIPL downregulation in the observed sensitisation of TRAIL-resistant cell lines. The downregulation of cFLIP with siRNA and the consequent sensitisation of Hep3b cells for TRAIL-induced apoptosis (Figure 8) showed that cFLIP downregulation is sufficient for TRAIL sensitisation. Together with the increased recruitment of caspase-8 to the TRAIL-DISC following 5-FU treatment, the downregulation of cFLIP in the lysates of 5-FU-treated cells results in an increase in the ratio of caspase-8 to cFLIP within the DISC (Figure 7). This suggests that non-pretreated, resistant cells mainly contain inhibited DISCs, whereas 5-FU-pretreated cells contain active DISCs that catalyse the activation of caspase-8 at a rate that allows for subsequent induction of apoptosis.66
Taken together, our findings allow for the following conclusions: first, TRAIL-R3 and TRAIL-R4 do not function as decoy receptors in Hep3b and HepG2-TR HCC cell lines. Second, 5-FU treatment upregulates the TRAIL death receptors, but this upregulation is insufficient to explain 5-FU-mediated sensitisation, indicating that TRAIL sensitivity is also regulated at the level of caspase-8 recruitment and activation.67, 68, 69 Third, we show that both cFLIPL and cFLIPS are recruited to the native TRAIL DISC, that they are downregulated by 5-FU treatment and that specific downregulation of cFLIP by siRNA in Hep3b cells sensitises these cells for TRAIL-induced apoptosis. Therefore, enhanced activation of caspase-8 can be explained by the increased ratio of caspase-8 to cFLIP within the TRAIL DISC of tumour cells.70 Efficient recruitment of caspase-8 to the DISC and its activation at this complex is decisive for TRAIL sensitivity. Notably, the described sensitisation mechanism for tumour cells does not apply for primary human hepatocytes. These cells remained resistant when cotreated with 5-FU and TRAIL (own unpublished data). This difference between sensitisation of HCC cells, on the one hand, and normal hepatocytes, on the other, may open up a new therapeutic window for TRAIL in combination with chemotherapeutics.
Materials and Methods
Cell lines
The human HCC cell lines HepG2, HepG2-TR and Hep3b were maintained in DMEM (Gibco-BRL, Karlsruhe, Germany) containing 10% foetal calf serum (Gibco-BRL). The HepG2-TR cells are a TRAIL-resistant variant of HepG2 cells that were obtained by in vitro selection for TRAIL resistance. They were periodically treated with increasing concentrations of TRAIL and selected for resistance to TRAIL-induced apoptosis.
Antibodies and reagents
Monoclonal antibodies (mAb) against FADD/MORT1 were purchased from Transduction Laboratories (San Diego, CA, USA). The mAb anti-FLICE C15 recognises the p18 subunit of caspase-871 and the mAb anti-cFLIP NF6 recognises the N-terminal portion of cFLIP;64 both antibodies are also available from Alexis (San Diego, CA, USA). Leucine Zipper (LZ)-TRAIL is a stable trimer of TRAIL and induces apoptosis upon binding to TRAIL-sensitive cells.11 LZ-TRAIL was purified from the supernatant of COS-7 cells transfected with LZ-TRAIL-pcDNA3.1. The purification was carried out as described.11 The antibodies specific for the different TRAIL receptors were anti-TRAIL-R1 HS101, TRAIL-R2 HS201, TRAIL-R3 HS301 and TRAIL-R4 HS402 for FACS analysis; for agonistic antibody-mediated killing, we used TRAIL-R1 HS101 and TRAIL-R2 HS201. Anti-TRAIL-R1 polyclonal rabbit Ab 210-730-C100 and anti-TRAIL-R2 polyclonal goat Ab 210-743-R100 were used for Western blot detection of TRAIL-R1 and TRAIL-R2, respectively. All TRAIL receptor antibodies are available from Alexis (San Diego, CA, USA). Horseradish peroxidase (HRPO)-conjugated goat anti-mouse IgG1, IgG2a and IgG2b and anti-rabbit and anti-goat polyclonal antibodies (pAb) were obtained from Southern Biotechnology Associates (Birmingham, AL, USA). All other chemicals used were of analytical grade and purchased from Merck (Darmstadt, Germany) or Sigma Chemical Co. (St. Louis, MO, USA).
MTT assay
A total of 5 × 104 cells were seeded in 96-well plates (Costar, Cambridge, MA, USA) with or without apoptotic stimuli in 150?μl medium at 37°C. After 20?h, 30?μl of an MTT solution (5?mg/ml MTT, Sigma Chemical Co., St. Louis, MO, USA, in phosphate-buffered saline (PBS)) was added for additional 4?h, the medium was then removed and cells lysed with 95% isopropanol/5% acetic acid (v/v). Absorption was determined at 560?nm. The percentage viable cells were calculated as follows: 100 × (absorption measured−absorption lysed cells)/(absorption medium−absorption lysed cells).
FACS analysis
Subconfluent HCC cell lines were detached from the plates by EDTA (2?mM) treatment. Cells were incubated with mAbs of the same isotype (mIgG1) against the four surface-expressed TRAIL receptors (TRAIL-R1 HS101, TRAIL-R2 HS201, TRAIL-R3 HS301 and TRAIL-R4 HS402) (Alexis, San Diego, CA, USA) or control mIgG1 followed by biotinylated secondary goat anti-mouse antibodies (Southern Biotechnology Associates, CA, USA) and streptavidin–PE (Pharmingen, Hamburg, Germany). Surface staining was determined with a FACScan cytometer (Becton Dickinson, Heidelberg, Germany). Specificity in FACS staining of the respective anti-TRAIL-R mAbs used here was determined by staining of TRAIL-R1 to TRAIL-R4 on CV1/EBNA cells transfected with pcDNA3.1-based expression vectors coding for the individual surface-bound TRAIL receptors (data not shown).
Quantification of apoptotic cell death
As a direct measurement of apoptotic cell death, DNA fragmentation was quantified as described.72 Briefly, 1.5 × 105 cells were incubated in 12-well plates (Costar, Cambridge, MA, USA) with or without apoptotic stimuli in 1?ml medium at 37°C. Cells were trypsinised and then collected by centrifugation at 600 × g for 10?min at 4°C, washed twice with PBS and then resuspended in 100?μl lyses solution containing 0.1% (v/v) Triton X-100, 0.1% (w/v) sodium citrate and 50?μg/ml propidium iodide (PI). Apoptosis was quantitatively determined by flow cytometry after incubation at 4°C in the dark for at least 24?h as cells containing nuclei with subdiploid DNA content.
Preparation of cell lysates
Subconfluent HCC cell lines were detached from the plates by EDTA (2?mM) treatment. Cells were harvested by centrifugation at 300 × g for 10?min at 4°C, washed twice with PBS and lysates were prepared by resuspending the resulting cell pellets in 100?μl lysis buffer per 1 × 107 cells (30?mM Tris-HCl pH 7.5, 150?mM NaCl, 10% glycerol 1% Triton X-100) supplemented with Complete™ protease inhibitors (Roche Diagnostics, Mannheim, Germany) according to the manufacturer's instructions. After 30?min incubation on ice, the lysates were centrifuged once at 15?000 × g at 4°C to remove nuclei. In the case of lysate preparation for ligand affinity precipitation, an intermediate centrifugation step (600 × g for 15?min at 4°C) was added after lysis in order to remove cellular debris.
Western blot analysis
For Western blot analysis, the resulting postnuclear supernatants or ligand affinity precipitates were supplemented with two-fold concentrated standard reducing sample buffer (2 × RSB). Subsequently, lysate containing 20?μg of protein as determined by the BCA method (Pierce, Rockford, IL, USA) or proteins eluted from beads after ligand affinity immunoprecipitation were separated on 4–12% NuPage Bis-Tris gradient gels (Novex, San Diego, CA, USA) in MOPS buffer according to the manufacturer's instructions. After protein transfer onto nitrocellulose membranes (Amersham Pharmacia Biotech, Freiburg, Germany) by electroblotting, membranes were blocked with 5% nonfat dry milk (NFDM) in PBS/Tween (PBS containing 0.05% Tween-20) for at least 2?h, washed with PBS/Tween, and incubated in PBS/Tween containing 3% NFDM and primary antibodies against FADD/MORT1, TRAIL-R1, TRAIL-R2 (all at 1?μg/ml), cFLIP (1?:?10 diluted NF6 hybridoma supernatant), caspase-8 (1?:?10 diluted C15 hybridoma supernatant)71 or ERK1 from Transduction Laboratories (San Diego, CA, USA). Specificity of the respective anti-TRAIL-R mAbs used here was determined by Western blot analysis of TRAIL-R1 to TRAIL-R4 in lysates prepared from CV1/EBNA cells transfected with expression vectors coding for the individual TRAIL receptors (data not shown). After six washes for 5?min each in PBS/Tween, the blots were incubated with HRPO-conjugated isotype-specific secondary antibody diluted 1?:?20?000 in PBS/Tween for 1?h. After washing six times for 5?min with PBS/Tween, the blots were developed by enhanced chemoluminescence following the manufacturer's protocol (Amersham Pharmacia Biotech, Uppsala, Sweden). For stripping, blots were either incubated for 30?min in a buffer containing 62.5?mM Tris-HCl, pH 6.8, 2% SDS and 100?mM mercaptoethanol at 60°C or in 50?mM glycine HCl pH 2.3 for 20?min at room temperature when only secondary antibodies needed to be removed. Subsequently, blots were washed six times for 10?min in PBS/Tween and blocked again.
Ligand affinity precipitation
We performed ligand affinity precipitation by using biotinylated LZ-TRAIL (Bio-LZ-TRAIL) in combination with Neutravidin beads (Pierce). Bio-LZ-TRAIL was prepared by incubation of purified LZ-TRAIL at 1?mg/ml with Sulfo-NHS-LC-Biotin at 1?mg/ml (Pierce) for 1?h on ice before the reaction was stopped by adding 1/10 volume of 1?M Tris-HCl at pH 7.5. Unincorporated biotin was removed from Bio-LZ-TRAIL preparations by buffer exchange into 150?mM NaCl, 30?mM HEPES pH 7.5 on PD-10 columns (Amersham Pharmacia Biotech). Protein preparations were checked for purity and incorporation of biotin by SDS-PAGE. The biological activity of Bio-LZ-TRAIL was determined by its apoptosis-inducing capacity and found to be comparable to nonbiotinylated LZ-TRAIL. For ligand affinity precipitation, 4 × 107 cells were used per sample. Cells were washed twice with 50?ml RPMI medium at 37°C and subsequently incubated for the indicated time periods at 37°C in the presence of 1?μg/ml Bio-LZ-TRAIL or, for the unstimulated control, in the absence of Bio-LZ-TRAIL. DISC formation was stopped by the addition of at least 15 volumes of ice-cold PBS. Cells were then washed with 50?ml ice-cold PBS before cell lysates were prepared. A total of 4 × 107 cells were lysed by the addition of 2?ml lysis buffer. The resulting protein complexes were precipitated from the lysates by coincubation with 20?μl streptavidin beads (Pierce) for 24?h on an end-over-end shaker at 4°C. For the precipitation of the nonstimulated receptors, Bio-LZ-TRAIL was added to the lysates prepared from nonstimulated cells at 1?μg/ml to control for protein association to nonstimulated receptors. Ligand affinity precipitates were washed four times with lysis buffer before the protein complexes were eluted from the beads by the addition of 15?μl 2 × standard RSB. Subsequently, proteins were separated on SDS-PAGE before the presence of antigens was determined in the different precipitates by Western blot analysis.
RNA interference
Oligonucleotides encoding short hairpin RNA targeting c-FLIP were selected as described by Brummelkamp et al.73 The specific target sequence for cFLIP (Acc. no.: U97074) was: nucleotides 909–929; and for the control we used a nonfunctional siRNA: nucleotides 651–671. Oligonucleotides were mixed in equal amounts of sense and antisense strands in annealing buffer (100?mM potassium acetate, 30?mM HEPES pH 7.4, 2?mM Mg acetate), incubated for 4?min at 95°C, for 10?min at 70°C and slowly cooled down to room temperature, phosphorylated and inserted into the BglII and HindIII sites of pSUPER.gfp/neo (Oligoengine). Sequences of the resulting vectors were verified by sequencing.
Cells were then transfected with the according vectors with Fugene 6 according to the manufacturer's instructions. At 48?h after transfection, cells were incubated with LZ-TRAIL. Kill of GFP-positive cells was determined by PI uptake and analysed by FACScan cytometer (Becton Dickinson, Heidelberg, Germany). Alternatively transfected Hep3b cells were cultured on chamber slides and stimulated with LZ-TRAIL. The cells were washed twice with PBS and fixed for 15?min on ice with 4% formaldehyde in PBS. The cells were washed with PBS and fixed 10?min in acetone. To inhibit unspecific binding of the antibody, the cells were incubated with blocking solution (BSA 20?mg/ml, human IgG 1?mg/ml in PBS). After 15?min, the blocking solution was replaced with rabbit anti-active caspase-3 (BD, 559565) and the cells were incubated for 1?h at room temperature. After washing in PBS, the cells were incubated with 20% normal goat serum in PBS for 15?min and then with biotinylated goat anti-rabbit for 30?min. The cells were again washed twice with PBS, incubated with streptavidin–PE (Pharmingen, 534061), washed twice again and then they were counterstained with DAPI (2?μg/ml). For the evaluation of immunofluorescence, a Zeiss Axioskop 40 was used.
Abbreviations
- 5-FU:
-
5-fluorouracil
- cFLIPL/S:
-
cellular FLICE-inhibitory proteinlong/short
- cIAP:
-
cellular inhibitor of apoptosis protein
- DD:
-
death domain
- DED:
-
death effector domain
- DISC:
-
death-inducing signalling complex
- HCC:
-
hepatocellular carcinoma
- OPG:
-
osteoprotegerin
- siRNA:
-
small interfering RNA
- TNF:
-
tumour necrosis factor
- TRAIL:
-
TNF-related apoptosis-inducing ligand
References
Wiley SR, Schooley K, Smolak PJ, Din WS, Huang CP, Nicholl JK, Sutherland GR, Smith TD, Rauch C and Smith CA (1995) Identification and characterization of a new member of the TNF family that induces apoptosis. Immunity 3: 673–682
Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M, Chin W, Jones J, Woodward A, Le T, Smith C, Smolak P, Goodwin RG, Rauch CT, Schuh JC and Lynch DH (1999) Tumoricidal activity of tumor necrosis factor-related apoptosis-inducing ligand in vivo. Nat. Med. 5: 157–163
Gores GJ and Kaufmann SH (2001) Is TRAIL hepatotoxic? Hepatology 34: 3–6
Ashkenazi A, Pai RC, Fong S, Leung S, Lawrence DA, Marsters SA, Blackie C, Chang L, McMurtrey AE, Hebert A, DeForge L, Koumenis IL, Lewis D, Harris L, Bussiere J, Koeppen H, Shahrokh Z and Schwall RH (1999) Safety and antitumor activity of recombinant soluble Apo2 ligand. J. Clin. Invest. 104: 155–162
Gliniak B and Le T (1999) Tumor necrosis factor-related apoptosis-inducing ligand's antitumor activity in vivo is enhanced by the chemotherapeutic agent CPT-11. Cancer Res. 59: 6153–6158
Jo M, Kim TH, Seol DW, Esplen JE, Dorko K, Billiar TR and Strom SC (2000) Apoptosis induced in normal human hepatocytes by tumor necrosis factor-related apoptosis-inducing ligand. Nat. Med. 6: 564–567
Ichikawa K, Liu W, Zhao L, Wang Z, Liu D, Ohtsuka T, Zhang H, Mountz JD, Koopman WJ, Kimberly RP and Zhou T (2001) Tumoricidal activity of a novel anti-human DR5 monoclonal antibody without hepatocyte cytotoxicity. Nat. Med. 7: 954–960
Lawrence D, Shahrokh Z, Marsters S, Achilles K, Shih D, Mounho B, Hillan K, Totpal K, DeForge L, Schow P, Hooley J, Sherwood S, Pai R, Leung S, Khan L, Gliniak B, Bussiere J, Smith CA, Strom SS, Kelley S, Fox JA, Thomas D and Ashkenazi A (2001) Differential hepatocyte toxicity of recombinant Apo2L/TRAIL versions. Nat. Med. 7: 383–385
Pan G, O'Rourke K, Chinnaiyan AM, Gentz R, Ebner R, Ni J and Dixit VM (1997) The receptor for the cytotoxic ligand TRAIL. Science 276: 111–113
Schneider P, Bodmer JL, Thome M, Hofmann K, Holler N and Tschopp J (1997) Characterization of two receptors for TRAIL. FEBS Lett. 416: 329–334
Walczak H, Degli-Esposti MA, Johnson RS, Smolak PJ, Waugh JY, Boiani N, Timour MS, Gerhart MJ, Schooley KA, Smith CA, Goodwin RG and Rauch CT (1997) TRAIL-R2: a novel apoptosis-mediating receptor for TRAIL. EMBO J. 16: 5386–5397
Sheridan JP, Marsters SA, Pitti RM, Gurney A, Skubatch M, Baldwin D, Ramakrishnan L, Gray CL, Baker K, Wood WI, Goddard AD, Godowski P and Ashkenazi A (1997) Control of TRAIL-induced apoptosis by a family of signaling and decoy receptors. Science 277: 818–821
Wu GS, Burns TF, McDonald III ER, Jiang W, Meng R, Krantz ID, Kao G, Gan DD, Zhou JY, Muschel R, Hamilton SR, Spinner NB, Markowitz S, Wu G and el-Deiry WS (1997) KILLER/DR5 is a DNA damage-inducible p53-regulated death receptor gene. Nat. Genet. 17: 141–143
Screaton GR, Mongkolsapaya J, Xu XN, Cowper AE, McMichael AJ and Bell JI (1997) TRICK2, a new alternatively spliced receptor that transduces the cytotoxic signal from TRAIL. Curr. Biol. 7: 693–696
MacFarlane M, Ahmad M, Srinivasula SM, Fernandes-Alnemri T, Cohen GM and Alnemri ES (1997) Identification and molecular cloning of two novel receptors for the cytotoxic ligand TRAIL. J. Biol. Chem. 272: 25417–25420
Krammer PH (2000) CD95's deadly mission in the immune system. Nature 407: 789–795
Peter ME, Scaffidi C, Medema JP, Kischkel F and Krammer PH (1999) The death receptors. Results Probl. Cell Differ. 23: 25–63
Degli-Esposti MA, Smolak PJ, Walczak H, Waugh J, Huang CP, DuBose RF, Goodwin RG and Smith CA (1997) Cloning and characterization of TRAIL-R3, a novel member of the emerging TRAIL receptor family. J. Exp. Med. 186: 1165–1170
Pan G, Ni J, Yu G, Wei YF and Dixit VM (1998) TRUNDD, a new member of the TRAIL receptor family that antagonizes TRAIL signalling. FEBS Lett. 424: 41–45
Mongkolsapaya J, Cowper AE, Xu XN, Morris G, McMichael AJ, Bell JI and Screaton GR (1998) Lymphocyte inhibitor of TRAIL (TNF-related apoptosis-inducing ligand): a new receptor protecting lymphocytes from the death ligand TRAIL. J. Immunol. 160: 3–6
Degli-Esposti MA, Dougall WC, Smolak PJ, Waugh JY, Smith CA and Goodwin RG (1997) The novel receptor TRAIL-R4 induces NF-kappaB and protects against TRAIL-mediated apoptosis, yet retains an incomplete death domain. Immunity 7: 813–820
Emery JG, McDonnell P, Burke MB, Deen KC, Lyn S, Silverman C, Dul E, Appelbaum ER, Eichman C, DiPrinzio R, Dodds RA, James IE, Rosenberg M, Lee JC and Young PR (1998) Osteoprotegerin is a receptor for the cytotoxic ligand TRAIL. J. Biol. Chem. 273: 14363–14367
Walczak H and Sprick MR (2001) Biochemistry and function of the DISC. Trends Biochem. Sci. 26: 452–453
Sprick MR, Weigand MA, Rieser E, Rauch CT, Juo P, Blenis J, Krammer PH and Walczak H (2000) FADD/MORT1 and caspase-8 are recruited to TRAIL receptors 1 and 2 and are essential for apoptosis mediated by TRAIL receptor 2. Immunity 12: 599–609
Kischkel FC, Lawrence DA, Chuntharapai A, Schow P, Kim KJ and Ashkenazi A (2000) Apo2L/TRAIL-dependent recruitment of endogenous FADD and caspase-8 to death receptors 4 and 5. Immunity 12: 611–620
Bodmer JL, Holler N, Reynard S, Vinciguerra P, Schneider P, Juo P, Blenis J and Tschopp J (2000) TRAIL receptor-2 signals apoptosis through FADD and caspase-8. Nat. Cell Biol. 2: 241–243
Sprick MR, Rieser E, Stahl H, Grosse-Wilde A, Weigand MA and Walczak H (2002) Caspase-10 is recruited to and activated at the native TRAIL and CD95 death-inducing signalling complexes in a FADD-dependent manner but cannot functionally substitute caspase-8. EMBO J. 21: 4520–4530
Kischkel FC, Lawrence DA, Tinel A, LeBlanc H, Virmani A, Schow P, Gazdar A, Blenis J, Arnott D and Ashkenazi A (2001) Death receptor recruitment of endogenous caspase-10 and apoptosis initiation in the absence of caspase-8. J. Biol. Chem. 276: 46639–46646
Cuello M, Ettenberg SA, Nau MM and Lipkowitz S (2001) Synergistic induction of apoptosis by the combination of trail and chemotherapy in chemoresistant ovarian cancer cells. Gynecol. Oncol. 81: 380–390
Lacour S, Hammann A, Wotawa A, Corcos L, Solary E and Dimanche-Boitrel MT (2001) Anticancer agents sensitize tumor cells to tumor necrosis factor-related apoptosis-inducing ligand-mediated caspase-8 activation and apoptosis. Cancer Res. 61: 1645–1651
Nagane M, Pan G, Weddle JJ, Dixit VM, Cavenee WK and Huang HJ (2000) Increased death receptor 5 expression by chemotherapeutic agents in human gliomas causes synergistic cytotoxicity with tumor necrosis factor-related apoptosis-inducing ligand in vitro and in vivo. Cancer Res. 60: 847–853
Houghton JA (1999) Apoptosis and drug response. Curr. Opin. Oncol. 11: 475–481
Belka C, Schmid B, Marini P, Durand E, Rudner J, Faltin H, Bamberg M, Schulze-Osthoff K and Budach W (2001) Sensitization of resistant lymphoma cells to irradiation-induced apoptosis by the death ligand TRAIL. Oncogene 20: 2190–2196
Zhang XD, Nguyen T, Thomas WD, Sanders JE and Hersey P (2000) Mechanisms of resistance of normal cells to TRAIL induced apoptosis vary between different cell types. FEBS Lett. 482: 193–199
Wen J, Ramadevi N, Nguyen D, Perkins C, Worthington E and Bhalla K (2000) Antileukemic drugs increase death receptor 5 levels and enhance Apo-2L-induced apoptosis of human acute leukemia cells. Blood 96: 3900–3906
Thomas WD, Zhang XD, Franco AV, Nguyen T and Hersey P (2000) TNF-related apoptosis-inducing ligand-induced apoptosis of melanoma is associated with changes in mitochondrial membrane potential and perinuclear clustering of mitochondria. J. Immunol. 165: 5612–5620
van Noesel MM, van Bezouw S, Salomons GS, Voute PA, Pieters R, Baylin SB, Herman JG and Versteeg R (2002) Tumor-specific down-regulation of the tumor necrosis factor-related apoptosis-inducing ligand decoy receptors DcR1 and DcR2 is associated with dense promoter hypermethylation. Cancer Res. 62: 2157–2161
Spalding AC, Jotte RM, Scheinman RI, Geraci MW, Clarke P, Tyler KL and Johnson GL (2002) TRAIL and inhibitors of apoptosis are opposing determinants for NF-kappaB-dependent, genotoxin-induced apoptosis of cancer cells. Oncogene 21: 260–271
Leaman DW, Chawla-Sarkar M, Vyas K, Reheman M, Tamai K, Toji S and Borden EC (2002) Identification of X-linked inhibitor of apoptosis-associated factor-1 as an interferon-stimulated gene that augments TRAIL Apo2L-induced apoptosis. J. Biol. Chem. 277: 28504–28511
Kim JY, Kim YH, Chang I, Kim S, Pak YK, Oh BH, Yagita H, Jung YK, Oh YJ and Lee MS (2002) Resistance of mitochondrial DNA-deficient cells to TRAIL: role of Bax in TRAIL-induced apoptosis. Oncogene 21: 3139–3148
Walczak H, Bouchon A, Stahl H and Krammer PH (2000) Tumor necrosis factor-related apoptosis-inducing ligand retains its apoptosis-inducing capacity on Bcl-2- or Bcl-xL-overexpressing chemotherapy-resistant tumor cells. Cancer Res. 60: 3051–3057
Bernard D, Quatannens B, Vandenbunder B and Abbadie C (2001) Rel/NF-kappaB transcription factors protect against tumor necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL)-induced apoptosis by up-regulating the TRAIL decoy receptor DcR1. J. Biol. Chem. 276: 27322–27328
Leverkus M, Neumann M, Mengling T, Rauch CT, Brocker EB, Krammer PH and Walczak H (2000) Regulation of tumor necrosis factor-related apoptosis-inducing ligand sensitivity in primary and transformed human keratinocytes. Cancer Res. 60: 553–559
Hao C, Beguinot F, Condorelli G, Trencia A, Van Meir EG, Yong VW, Parney IF, Roa WH and Petruk KC (2001) Induction and intracellular regulation of tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) mediated apoptosis in human malignant glioma cells. Cancer Res. 61: 1162–1170
Chang DW, Xing Z, Pan Y, Algeciras-Schimnich A, Barnhart BC, Yaish-Ohad S, Peter ME and Yang X (2002) c-FLIP(L) is a dual function regulator for caspase-8 activation and CD95-mediated apoptosis. EMBO J. 21: 3704–3714
Yamanaka T, Shiraki K, Sugimoto K, Ito T, Fujikawa K, Ito M, Takase K, Moriyama M, Nakano T and Suzuki A (2000) Chemotherapeutic agents augment TRAIL-induced apoptosis in human hepatocellular carcinoma cell lines. Hepatology 32: 482–490
Muller M, Scaffidi CA, Galle PR, Stremmel W and Krammer PH (1998) The role of p53 and the CD95 (APO-1/Fas) death system in chemotherapy-induced apoptosis. Eur. Cytokine Network 9: 685–686
Krueger A, Schmitz I, Baumann S, Krammer PH and Kirchhoff S (2001) Cellular FLICE-inhibitory protein splice variants inhibit different steps of caspase-8 activation at the cd95 death-inducing signaling complex. J. Biol. Chem. 276: 20633–20640
Irmler M, Thome M, Hahne M, Schneider P, Hofmann K, Steiner V, Bodmer JL, Schroter M, Burns K, Mattmann C, Rimoldi D, French LE and Tschopp J (1997) Inhibition of death receptor signals by cellular FLIP. Nature 388: 190–195
Siegmund D, Hadwiger P, Pfizenmaier K, Vornlocher HP and Wajant H (2002) Selective inhibition of FLICE-like inhibitory protein expression with small interfering RNA oligonucleotides is sufficient to sensitize tumor cells for TRAIL-induced apoptosis. Mol. Med. 8: 725–732
Higuchi H, Bronk SF, Takikawa Y, Werneburg N, Takimoto R, El-Deiry W and Gores GJ (2001) The bile acid glycochenodeoxycholate induces TRAIL-receptor 2/DR5 expression and apoptosis. J. Biol. Chem. 276: 38610–38618
Guan B, Yue P, Clayman GL and Sun SY (2001) Evidence that the death receptor DR4 is a DNA damage-inducible, p53-regulated gene. J. Cell. Physiol. 188: 98–105
Kastan M (1997) On the TRAIL from p53 to apoptosis? Nat. Genet. 17: 130–131
Sheikh MS, Burns TF, Huang Y, Wu GS, Amundson S, Brooks KS, Fornace AJ, Jr and el-Deiry WS (1998) p53-dependent and -independent regulation of the death receptor KILLER/DR5 gene expression in response to genotoxic stress and tumor necrosis factor alpha. Cancer Res. 58: 1593–1598
Sheikh MS, Fornace AJ and Jr (2000) Death and decoy receptors and p53-mediated apoptosis. Leukemia 14: 1509–1513
Wu WG, Soria JC, Wang L, Kemp BL and Mao L (2000) TRAIL-R2 is not correlated with p53 status and is rarely mutated in non-small cell lung cancer. Anticancer Res. 20: 4525–4529
Deng Y, Lin Y and Wu X (2002) TRAIL-induced apoptosis requires Bax-dependent mitochondrial release of Smac/DIABLO. Genes Dev. 16: 33–45
Leverkus M, Sprick MR, Wachter T, Mengling T, Baumann B, Serfling E, Brocker EB, Goebeler M, Neumann M and Walczak H (2003) Proteasome inhibition results in TRAIL sensitization of primary keratinocytes by removing the resistance-mediating block of effector caspase maturation. Mol. Cell. Biol. 23: 777–790
Fulda S, Wick W, Weller M and Debatin KM (2002) Smac agonists sensitize for Apo2L/TRAIL- or anticancer drug-induced apoptosis and induce regression of malignant glioma in vivo. Nat. Med. 8: 808–815
Zhang XD, Zhang XY, Gray CP, Nguyen T and Hersey P (2001) Tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis of human melanoma is regulated by Smac/DIABLO release from mitochondria. Cancer Res. 61: 7339–7348
Fulda S, Meyer E and Debatin KM (2002) Inhibition of TRAIL-induced apoptosis by Bcl-2 overexpression. Oncogene 21: 2283–2294
Bin L, Li X, Xu LG and Shu HB (2002) The short splice form of Casper/c-FLIP is a major cellular inhibitor of TRAIL-induced apoptosis. FEBS Lett. 510: 37–40
Thome M, Schneider P, Hofmann K, Fickenscher H, Meinl E, Neipel F, Mattmann C, Burns K, Bodmer JL, Schroter M, Scaffidi C, Krammer PH, Peter ME and Tschopp J (1997) Viral FLICE-inhibitory proteins (FLIPs) prevent apoptosis induced by death receptors. Nature 386: 517–521
Scaffidi C, Schmitz I, Krammer PH and Peter ME (1999) The role of c-FLIP in modulation of CD95-induced apoptosis. J. Biol. Chem. 274: 1541–1548
Kirchhoff S, Muller WW, Krueger A, Schmitz I and Krammer PH (2000) TCR-mediated up-regulation of c-FLIP short correlates with resistance toward CD95-mediated apoptosis by blocking death-inducing signaling complex activity. J. Immunol. 165: 6293–6300
Harper N, Farrow SN, Kaptein A, Cohen GM and MacFarlane M (2001) Modulation of tumor necrosis factor apoptosis-inducing ligand-induced NF-kappa B activation by inhibition of apical caspases. J. Biol. Chem. 276: 34743–34752
Griffith TS, Chin WA, Jackson GC, Lynch DH and Kubin MZ (1998) Intracellular regulation of TRAIL-induced apoptosis in human melanoma cells. J. Immunol. 161: 2833–2840
Suliman A, Lam A, Datta R and Srivastava RK (2001) Intracellular mechanisms of TRAIL: apoptosis through mitochondrial-dependent and -independent pathways. Oncogene 20: 2122–2133
Srivastava RK (2000) Intracellular mechanisms of TRAIL and its role in cancer therapy. Mol. Cell. Biol. Res. Commun. 4: 67–75
MacFarlane M, Harper N, Snowden RT, Dyer MJ, Barnett GA, Pringle JH and Cohen GM (2002) Mechanisms of resistance to TRAIL-induced apoptosis in primary B cell chronic lymphocytic leukaemia. Oncogene 21: 6809–6818
Scaffidi C, Medema JP, Krammer PH and Peter ME (1997) FLICE is predominantly expressed as two functionally active isoforms, caspase-8/a and caspase-8/b. J. Biol. Chem. 272: 26953–26958
Nicoletti I, Migliorati G, Pagliacci MC, Grignani F and Riccardi C (1991) A rapid and simple method for measuring thymocyte apoptosis by propidium iodide staining and flow cytometry. J. Immunol. Methods 139: 271–279
Zhang L, Fogg DK and Waisman DM (2003) RNA interference-mediated silencing of the S100A10 gene attenuates plasmin generation and invasiveness of Colo 222 colorectal cancer cells. J. Biol. Chem. 279: 2053–2062
Acknowledgements
We thank Michaela Rapp for excellent technical assistance, Ronald Koschny, Sören Eichhorst and Silke Maier for helpful discussions and careful reading of the paper. Henning Walczak is recipient of a BioFuture grant from the Bundesministerium für Bildung und Forschung (BMBF), Tom M Ganten is recipient of a scholarship from the Deutsche Forschungsgemeinschaft (DFG). The work was funded by the Tumorzentrum HD/MA (TM Ganten and H Walczak).
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by GM Cohen
Rights and permissions
About this article
Cite this article
Ganten, T., Haas, T., Sykora, J. et al. Enhanced caspase-8 recruitment to and activation at the DISC is critical for sensitisation of human hepatocellular carcinoma cells to TRAIL-induced apoptosis by chemotherapeutic drugs. Cell Death Differ 11 (Suppl 1), S86–S96 (2004). https://doi.org/10.1038/sj.cdd.4401437
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401437
Keywords
This article is cited by
-
BAX-dependent mitochondrial pathway mediates the crosstalk between ferroptosis and apoptosis
Apoptosis (2020)
-
Defined factors to reactivate cell cycle activity in adult mouse cardiomyocytes
Scientific Reports (2019)
-
A novel TRAIL mutant-TRAIL-Mu3 enhances the antitumor effects by the increased affinity and the up-expression of DR5 in pancreatic cancer
Cancer Chemotherapy and Pharmacology (2018)
-
N-terminal gelsolin fragment potentiates TRAIL mediated death in resistant hepatoma cells
Scientific Reports (2017)
-
Trail (TNF-related apoptosis-inducing ligand) induces an inflammatory response in human adipocytes
Scientific Reports (2017)