Abstract
Atopic diseases, including asthma, allergic rhinitis and atopic dermatitis, are caused by both environmental and genetic factors. Here we show that infection by hepatitis A virus (HAV) may protect individuals from atopy if they carry a particular variant of the gene that encodes TIM-1 (also known as HAVcr-1) — the cell-surface receptor used by HAV to infect human cells1. Exposure to HAV is associated with poor hygiene, large family size and attendance at day-care centres, all factors that are also inversely associated with atopy2,3,4,5,6. Our discovery indicates that interaction between HAV and TIM-1 genotype may contribute to the aetiology of atopic diseases, and provides a mechanism to account for the hygiene hypothesis.
Similar content being viewed by others
Main
Using a congenic positional cloning strategy, we identified TIM-1 as a candidate gene for atopy and asthma in a region of mouse chromosome 11, which is homologous to a segment of human chromosome 5q31–33 that has been linked to atopy7,8. TIM-1 is expressed by activated CD4+ T cells during the development of helper-T-cell (Th2) responses and regulates cytokine production7. We therefore investigated whether the interaction between HAV and TIM-1 on lymphocytes can modify T cells in a way that protects against atopy, and whether polymorphisms in TIM-1 can alter susceptibility to atopy7.
By sequencing complementary DNA from human lymphocytes, we identified a six-amino-acid insertion (ins) at residue 157, termed 157insMTTTVP (one-letter amino-acid notation), as well as two single-amino-acid changes, 195delT (where 'del' signifies a deletion) and A206T. The insertion 157insMTTTVP is located at the centre of an extracellular mucin-like region that is required for efficient HAV uncoating9 and, because 157insMTTTVP lengthens this critical region by 12–14%, this variation may affect the efficiency of viral entry (Fig. 1).
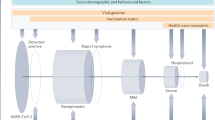
a, A variant of TIM-1 ('hook') that carries a 6-amino-acid insert (red), 157insMTTTVP, may increase binding of HAV (large pentagons) to the receptor, thereby enhancing HAV viral uncoating (small pentagons) and infection of TIM-1-expressing T cells. This could lead to deletion of certain lymphocyte subsets, such as Th2 cells, or reduce Th2-cell differentiation, causing a reduction in atopy and asthma. b, Alternatively, HAV may bind less efficiently to the form of TIM-1 without the insertion, resulting in less HAV infection and hence more Th2-cell development and more atopy. The mechanism that underpins this interaction between TIM-1, its 157insMTTTVP region and HAV could relate to viral uncoating9, the extent and duration of HAV viraemia10, or to a direct effect on Th1/Th2-subset differentiation.
To determine the effect of the insertion 157insMTTTVP on the occurrence of atopy, we carried out a cross-sectional study of 375 individuals who were evaluated by history and tested serologically for atopy and prior HAV infection. To correct for potentially confounding effects of population admixture, we used stratified Mantel–Haenszel χ2 tests to quantify the association between atopy and 157insMTTTVP in the total sample. We found that HAV seropositivity protects against atopy, but only in individuals with the 157insMTTTVP variant of TIM-1 (P = 0.0005; Table 1).
The protective effects of HAV therefore depend upon a common TIM-1 allele that is carried by 63% of Caucasians, 46% of Asians and 64% of African Americans in this population (see supplementary information). As allelic variation in TIM-1 does not affect HAV-infection rates in our population (χ2 = 1.567, P = 0.211), we conclude that the interaction of HAV with TIM-1 genotype seen here is not due to variation in the rate of seroconversion following HAV exposure.
Before 1970, the seroprevalence of antibodies against HAV approached 100% in Western countries4, and infection with HAV may have protected many individuals against atopy3. However, modernization has led to a reduction in average family size and significant improvements in public health, causing anti-HAV seroprevalence to fall to 25–30%, while the prevalence of atopic disease has doubled4. Our finding that TIM-1 is associated with atopy in HAV-seropositive individuals indicates that exposure to a specific pathogen may influence the expression of atopy — so a declining prevalence of HAV infection could contribute to an increase in atopy by association with TIM-1. It will be necessary to determine whether HAV exposure must occur during childhood to have a protective effect, whether HAV can mitigate the severity of existing atopic disease, and whether HAV vaccination can reproduce the effects of natural HAV infection.
References
Feigelstock, D., Thompson, P., Mattoo, P., Zhang, Y. & Kaplan, G. G. J. Virol. 72, 6621–6628 (1998).
Matricardi, P. M. et al. Br. Med. J. 314, 999–1003 (1997).
Matricardi, P. M., Rosmini, F., Panetta, V., Ferrigno, L. & Bonini, S. J. Allerg. Clin. Immunol. 110, 381–387 (2002).
Bach, J. F. N. Engl. J. Med. 347, 911–920 (2002).
Strachan, D. P. Br. Med. J. 299, 1259–1260 (1989).
Kramer, U., Heinrich, J., Wjst, M. & Wichmann, H. E. Lancet 353, 450–454 (1999).
McIntire, J. J. et al. Nature Immunol. 2, 1109–1116 (2001).
Marsh, D. G. et al. Science 264, 1152–1156 (1994).
Silberstein, E. et al. J. Virol. 77, 8765–8774 (2003).
Bower, W. A., Nainan, O. V., Han, X. & Margolis, H. S. J. Infect. Dis. 182, 12–17 (2000).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
McIntire, J., Umetsu, S., Macaubas, C. et al. Hepatitis A virus link to atopic disease. Nature 425, 576 (2003). https://doi.org/10.1038/425576a
Issue Date:
DOI: https://doi.org/10.1038/425576a
This article is cited by
-
Genomic heterozygosity is associated with parasite abundance, but the effects are not mediated by host condition
Evolutionary Ecology (2023)
-
Tumor-derived exosomal HMGB1 fosters hepatocellular carcinoma immune evasion by promoting TIM-1+ regulatory B cell expansion
Journal for ImmunoTherapy of Cancer (2018)
-
The phospholipid code: a key component of dying cell recognition, tumor progression and host–microbe interactions
Cell Death & Differentiation (2015)
-
Protective Association of TIM1−1454G>A Polymorphism with Asthma in a North Indian Population
Lung (2015)
-
The relationship between polymorphisms in the promoter region of Tim-3 and unexplained recurrent spontaneous abortion in Han Chinese women
Reproductive Biology and Endocrinology (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.