Abstract
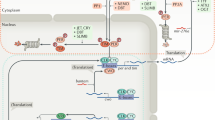
In this era of jet travel, our body 'remembers' the previous time zone, such that when we travel, our sleep–wake pattern, mental alertness, eating habits and many other physiological processes temporarily suffer the consequences of time displacement until we adjust to the new time zone. Although the existence of a circadian clock in humans had been postulated for decades, an understanding of the molecular mechanisms has required the full complement of research tools. To gain the initial insights into circadian mechanisms, researchers turned to genetically tractable model organisms such as Drosophila.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Pittendrigh, C. S. Circadian systems. I. The driving oscillation and its assay in Drosophila pseudoobscura. Proc. Natl Acad. Sci. USA 58, 1762–1767 (1967).
Shaw, P. J., Cirelli, C., Greenspan, R. J. & Tononi, G. Correlates of sleep and waking in Drosophila melanogaster. Science 287, 1834–1837 (2000).
Hendricks, J. C. et al. Rest in Drosophila is a sleep-like state. Neuron 25, 129–138 (2000).
Konopka, R. J. & Benzer, S. Clock mutants of Drosophila melanogaster. Proc. Natl Acad. Sci. USA 68, 2112–2116 (1971).
Handler, A. M. & Konopka, R. J. Transplantation of a circadian pacemaker in Drosophila. Nature 279, 236–238 (1979).
Bargiello, T. A., Jackson, F. R. & Young, M. W. Restoration of circadian behavioural rhythms by gene transfer in Drosophila. Nature 312, 752–754 (1984).
Reddy, P. et al. Molecular analysis of the period locus in Drosophila melanogaster and identification of a transcript involved in biological rhythms. Cell 38, 701–710 (1984).
Baylies, M. K., Bargiello, T. A., Jackson, F. R. & Young, M. W. Changes in abundance or structure of the per gene product can alter periodicity of the Drosophila clock. Nature 326, 390–392 (1987).
Ewer, J., Frisch, B., Hamblen-Coyle, M. J., Rosbash, M. & Hall, J. C. Expression of the period clock gene within different cell types in the brain of Drosophila adults and mosaic analysis of these cells' influence on circadian behavioral rhythms. J. Neurosci. 12, 3321–3349 (1992).
Vosshall, L. B. & Young, M. W. Circadian rhythms in Drosophila can be driven by period expression in a restricted group of central brain cells. Neuron 15, 345–360 (1995).
Colot, H. V., Hall, J. C. & Rosbash, M. Interspecific comparison of the period gene of Drosophila reveals large blocks of non-conserved coding DNA. EMBO J. 7, 3929–3937 (1988).
Reppert, S. M., Tsai, T., Roca, A. L. & Sauman, I. Cloning of a structural and functional homolog of the circadian clock gene period from the giant silkmoth Antheraea pernyi. Neuron 13, 1167–1176 (1994).
Sauman, I. & Reppert, S. M. Circadian clock neurons in the silkmoth Antheraea pernyi: novel mechanisms of Period protein regulation. Neuron 17, 889–900 (1996).
Regier, J. C. et al. Evolution and phylogenetic utility of the period gene in Lepidoptera. Mol. Biol. Evol. 15, 1172–1182 (1998).
Toma, D. P., Bloch, G., Moore, D. & Robinson, G. E. Changes in period mRNA levels in the brain and division of labor in honey bee colonies. Proc. Natl Acad. Sci. USA 97, 6914–6919 (2000).
Levine, J. D., Sauman, I., Imbalzano, M., Reppert, S. M. & Jackson, F. R. Period protein from the giant silkmoth Antheraea pernyi functions as a circadian clock element in Drosophila melanogaster. Neuron 15, 147–157 (1995).
Siwicki, K. K., Strack, S., Rosbash, M., Hall, J. C. & Jacklet, J. W. An antibody to the Drosophila period protein recognizes circadian pacemaker neurons in Aplysia and Bulla. Neuron 3, 51–58 (1989).
Siwicki, K. K., Schwartz, W. J. & Hall, J. C. An antibody to the Drosophila period protein labels antigens in the suprachiasmatic nucleus of the rat. J. Neurogenet. 8, 33–42 (1992).
Sehgal, A., Price, J. L., Man, B. & Young, M. W. Loss of circadian behavioral rhythms and per RNA oscillations in the Drosophila mutant timeless. Science 263, 1603–1606 (1994).
Vosshall, L. B., Price, J. L., Sehgal, A., Saez, L. & Young, M. W. Block in nuclear localization of period protein by a second clock mutation, timeless. Science 263, 1606–1609 (1994).
Sehgal, A. et al. Rhythmic expression of timeless: a basis for promoting circadian cycles in period gene autoregulation. Science 270, 808–810 (1995).
Gekakis, N. et al. Isolation of timeless by PER protein interaction: defective interaction between timeless protein and long-period mutant PERL. Science 270, 811–815 (1995).
Hoffman, E. C. et al. Cloning of a factor required for activity of the Ah (dioxin) receptor. Science 252, 954–958 (1991).
Crews, S. T., Thomas, J. B. & Goodman, C. S. The Drosophila single-minded gene encodes a nuclear protein with sequence similarity to the per gene product. Cell 52, 143–151 (1988).
Huang, Z. J., Edery, I. & Rosbash, M. PAS is a dimerization domain common to Drosophila Period and several transcription factors. Nature 364, 259–262 (1993).
Hao, H., Allen, D. L. & Hardin, P. E. A circadian enhancer mediates PER-dependent mRNA cycling in Drosophila melanogaster. Mol. Cell. Biol. 17, 3687–3693 (1997).
Darlington, T. K. et al. Closing the circadian loop: CLOCK-induced transcription of its own inhibitors per and tim. Science 280, 1599–1603 (1998).
Rutila, J. E. et al. CYCLE is a second bHLH-PAS clock protein essential for circadian rhythmicity and transcription of Drosophila period and timeless. Cell 93, 805–814 (1998).
Allada, R., White, N. E., So, W. V., Hall, J. C. & Rosbash, M. A mutant Drosophila homolog of mammalian Clock disrupts circadian rhythms and transcription of period and timeless. Cell 93, 791–804 (1998).
Hogenesch, J. B., Gu, Y. Z., Jain, S. & Bradfield, C. A. The basic-helix-loop-helix-PAS orphan MOP3 forms transcriptionally active complexes with circadian and hypoxia factors. Proc. Natl Acad. Sci. USA 95, 5474–5479 (1998).
Gekakis, N. et al. Role of the CLOCK protein in the mammalian circadian mechanism. Science 280, 1564–1569 (1998).
Price, J. L. et al. double-time is a novel Drosophila clock gene that regulates PERIOD protein accumulation. Cell 94, 83–95 (1998).
Blau, J. & Young, M. W. Cycling vrille expression is required for a functional Drosophila clock. Cell 99, 661–671 (1999).
Martinek, S., Inonog, S., Manoukian, A. S. & Young, M. W. A role for the segment polarity gene shaggy/GSK-3 in the Drosophila circadian clock. Cell 105, 769–779 (2001).
Kloss, B., Rothenfluh, A., Young, M. W. & Saez, L. Phosphorylation of PERIOD is influenced by cycling physical associations of DOUBLE-TIME, PERIOD, and TIMELESS in the Drosophila clock. Neuron 30, 699–706 (2001).
Rothenfluh, A., Young, M. W. & Saez, L. A TIMELESS-independent function for PERIOD proteins in the Drosophila clock. Neuron 26, 505–514 (2000).
Bao, S., Rihel, J., Bjes, E., Fan, J. Y. & Price, J. L. The Drosophila double-timeS mutation delays the nuclear accumulation of period protein and affects the feedback regulation of period mRNA. J. Neurosci. 21, 7117–7126 (2001).
Bae, K., Lee, C., Sidote, D., Chuang, K. Y. & Edery, I. Circadian regulation of a Drosophila homolog of the mammalian Clock gene: PER and TIM function as positive regulators. Mol. Cell. Biol. 18, 6142–6151 (1998).
Lee, C., Bae, K. & Edery, I. The Drosophila CLOCK protein undergoes daily rhythms in abundance, phosphorylation, and interactions with the PER-TIM complex. Neuron 21, 857–867 (1998).
Glossop, N. R., Lyons, L. C. & Hardin, P. E. Interlocked feedback loops within the Drosophila circadian oscillator. Science 286, 766–768 (1999).
Ralph, M. R. & Menaker, M. A mutation of the circadian system in golden hamsters. Science 241, 1225–1227 (1988).
Moore, R. Y. & Eichler, V. B. Loss of a circadian adrenal corticosterone rhythm following suprachiasmatic lesions in the rat. Brain Res. 42, 201–206 (1972).
Welsh, D. K., Logothetis, D. E., Meister, M. & Reppert, S. M. Individual neurons dissociated from rat suprachiasmatic nucleus express independently phased circadian firing rhythms. Neuron 14, 697–706 (1995).
Jin, X. et al. A molecular mechanism regulating rhythmic output from the suprachiasmatic circadian clock. Cell 96, 57–68 (1999).
Bunger, M. K. et al. Mop3 is an essential component of the master circadian pacemaker in mammals. Cell 103, 1009–1017 (2000).
Bae, K. et al. Differential functions of mPer1, mPer2, and mPer3 in the SCN circadian clock. Neuron 30, 525–536 (2001).
Zheng, B. et al. Nonredundant roles of the mPer1 and mPer2 genes in the mammalian circadian clock. Cell 105, 683–694 (2001).
Gotter, A. L. et al. A time-less function for mouse Timeless. Nature Neurosci. 3, 755–756 (2000).
Lee, C., Parikh, V., Itsukaichi, T., Bae, K. & Edery, I. Resetting the Drosophila clock by photic regulation of PER and a PER-TIM complex. Science 271, 1740–1744 (1996).
Myers, M. P., Wager-Smith, K., Rothenfluh-Hilfiker, A. & Young, M. W. Light-induced degradation of TIMELESS and entrainment of the Drosophila circadian clock. Science 271, 1736–1740 (1996).
Hunter-Ensor, M., Ousley, A. & Sehgal, A. Regulation of the Drosophila protein timeless suggests a mechanism for resetting the circadian clock by light. Cell 84, 677–685 (1996).
Zimmerman, W. F. & Goldsmith, T. H. Photosensitivity of the circadian rhythm and of visual receptors in carotenoid-depleted Drosophila. Science 171, 1167–1169 (1971).
Wheeler, D. A., Hamblen-Coyle, M. J., Dushay, M. S. & Hall, J. C. Behavior in light-dark cycles of Drosophila mutants that are arrhythmic, blind, or both. J. Biol. Rhythms 8, 67–94 (1993).
Thompson, C. L. et al. Preservation of light signaling to the suprachiasmatic nucleus in vitamin A-deficient mice. Proc. Natl Acad. Sci. USA 98, 11708–11713 (2001).
Czeisler, C. A. et al. Suppression of melatonin secretion in some blind patients by exposure to bright light. N. Engl. J. Med. 332, 6–11 (1995).
Stanewsky, R. et al. The cryb mutation identifies cryptochrome as a circadian photoreceptor in Drosophila. Cell 95, 681–692 (1998).
Emery, P., So, W. V., Kaneko, M., Hall, J. C. & Rosbash, M. CRY, a Drosophila clock and light-regulated cryptochrome, is a major contributor to circadian rhythm resetting and photosensitivity. Cell 95, 669–679 (1998).
Ceriani, M. F. et al. Light-dependent sequestration of TIMELESS by CRYPTOCHROME. Science 285, 553–556 (1999).
Naidoo, N., Song, W., Hunter-Ensor, M. & Sehgal, A. A role for the proteasome in the light response of the timeless clock protein. Science 285, 1737–1741 (1999).
Lin, F. J., Song, W., Meyer-Bernstein, E., Naidoo, N. & Sehgal, A. Photic signaling by cryptochrome in the Drosophila circadian system. Mol. Cell. Biol. 21, 7287–7294 (2001).
Krishnan, B. et al. A new role for cryptochrome in a Drosophila circadian oscillator. Nature 411, 313–317 (2001).
Kume, K. et al. mCRY1 and mCRY2 are essential components of the negative limb of the circadian clock feedback loop. Cell 98, 193–205 (1999).
Lee, C., Etchegaray, J. P., Cagampang, F. R., Loudon, A. S. & Reppert, S. M. Posttranslational mechanisms regulate the mammalian circadian clock. Cell 107, 855–867 (2001).
van der Horst, G. T. et al. Mammalian Cry1 and Cry2 are essential for maintenance of circadian rhythms. Nature 398, 627–630 (1999).
Takahashi, J. S., DeCoursey, P. J., Bauman, L. & Menaker, M. Spectral sensitivity of a novel photoreceptive system mediating entrainment of mammalian circadian rhythms. Nature 308, 186–188 (1984).
Lucas, R. J. et al. Identifying the photoreceptive inputs to the mammalian circadian system using transgenic and retinally degenerate mice. Behav. Brain Res. 125, 97–102 (2001).
Antoch, M. P. et al. Functional identification of the mouse circadian Clock gene by transgenic BAC rescue. Cell 89, 655–667 (1997).
Newby, L. M. & Jackson, F. R. A new biological rhythm mutant of Drosophila melanogaster that identifies a gene with an essential embryonic function. Genetics 135, 1077–1090 (1993).
Renn, S. C., Park, J. H., Rosbash, M., Hall, J. C. & Taghert, P. H. A pdf neuropeptide gene mutation and ablation of PDF neurons each cause severe abnormalities of behavioral circadian rhythms in Drosophila. Cell 99, 791–802 (1999).
Sarov-Blat, L., So, W. V., Liu, L. & Rosbash, M. The Drosophila takeout gene is a novel molecular link between circadian rhythms and feeding behavior. Cell 101, 647–656 (2000).
Harmer, S. L. et al. Orchestrated transcription of key pathways in Arabidopsis by the circadian clock. Science 290, 2110–2113 (2000).
Grundschober, C. et al. Circadian regulation of diverse gene products revealed by mRNA expression profiling of synchronized fibroblasts. J. Biol. Chem. 276, 46751–46758 (2001).
McDonald, M. J. & Rosbash, M. Microarray analysis and organization of circadian gene expression in Drosophila. Cell 107, 567–578 (2001).
Claridge-Chang, A. et al. Circadian regulation of gene expression systems in the Drosophila head. Neuron 32, 657–671 (2001).
Ceriani, M. F. et al. Genome-wide expression analysis in Drosophila predicts genes controlling circadian behavior. (Submitted.)
Panda, S. et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell (in the press).
Alabadi, D. et al. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293, 880–883 (2001).
Doi, M., Nakajima, Y., Okano, T. & Fukada, Y. Light-induced phase-delay of the chicken pineal circadian clock is associated with the induction of cE4bp4, a potential transcriptional repressor of cPer2 gene. Proc. Natl Acad. Sci. USA 98, 8089–8094 (2001).
Mitsui, S., Yamaguchi, S., Matsuo, T., Ishida, Y. & Okamura, H. Antagonistic role of E4BP4 and PAR proteins in the circadian oscillatory mechanism. Genes Dev. 15, 995–1006 (2001).
Kaneko, M. & Hall, J. C. Neuroanatomy of cells expressing clock genes in Drosophila: transgenic manipulation of the period and timeless genes to mark the perikarya of circadian pacemaker neurons and their projections. J. Comp. Neurol. 422, 66–94 (2000).
van Esseveldt, K. E., Lehman, M. N. & Boer, G. J. The suprachiasmatic nucleus and the circadian time-keeping system revisited. Brain Res. Brain Res. Rev. 33, 34–77 (2000).
Weiner, J. Time, Love, Memory (Vintage Books, New York, 1999).
Lowrey, P. L. et al. Positional syntenic cloning and functional characterization of the mammalian circadian mutation tau. Science 288, 483–492 (2000).
Keesler, G. A. et al. Phosphorylation and destabilization of human period 1 clock protein by human casein kinase Iε. NeuroReport 11, 951–955 (2000).
Vielhaber, E., Eide, E., Rivers, A., Gao, Z. H. & Virshup, D. M. Nuclear entry of the circadian regulator mPER1 is controlled by mammalian casein kinase Iε. Mol. Cell. Biol. 20, 4888–4899 (2000).
Toh, K. L. et al. An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291, 1040–1043 (2001).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Panda, S., Hogenesch, J. & Kay, S. Circadian rhythms from flies to human. Nature 417, 329–335 (2002). https://doi.org/10.1038/417329a
Issue Date:
DOI: https://doi.org/10.1038/417329a
This article is cited by
-
DVT: a high-throughput analysis pipeline for locomotion and social behavior in adult Drosophila melanogaster
Cell & Bioscience (2023)
-
Muscle quality index is associated with trouble sleeping: a cross-sectional population based study
BMC Public Health (2023)
-
The circadian rhythm gene Bmal1 ameliorates acute deoxynivalenol-induced liver damage
Archives of Toxicology (2023)
-
In Search of Regained Time? Autism and Organizational [A]temporality in the Light of Humanistic Management
Journal of Business Ethics (2023)
-
Conservation of dynamic characteristics of transcriptional regulatory elements in periodic biological processes
BMC Bioinformatics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.