Abstract
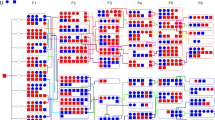
Molecular genetic studies of attention-deficit hyperactivity disorder (ADHD) are a major focus of current research since this syndrome has been shown to be highly heritable.1 Our approach has been to search for quantitative trait loci (QTL) in a genetic animal model of hyperkinesis, the Wistar–Kyoto hyperactive (WKHA) rat, by a whole-genome scan analysis. In a previous article, we reported the detection of a major QTL associated with behavioral activity in an F2 cross between WKHA and Wistar–Kyoto (WKY) rat strains.2 Here, we extend our analysis of this cross by adding new genetic markers, now defining a 10 cM interval on rat chromosome 8 associated with ambulatory and exploratory activities. Then we present a replication of this QTL detection, at least for exploratory activity, by a new genetic mapping analysis of an activity QTL in an F2 cross between the WKHA and Brown Norway (BN) rat strains. Overall, the results provide compelling evidence for the presence of gene(s) influencing activity at this locus. The QTL interval has been refined such that the human orthologous region could be defined and tested in human populations for association with ADHD. Ultimately, the improved dissection of this genomic locus should allow the identification of the causal genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Thapar A, Holmes J, Poulton K, Harrington R . Genetic basis of attention deficit and hyperactivity. Br J Psychiatry 1999; 174: 105–111.
Moisan MP, Courvoisier H, Bihoreau MT, Gauguier D, Hendley ED, Lathrop M et al. A major quantitative trait locus influences hyperactivity in the WKHA rat. Nat Genet 1996; 14: 471–473.
Steen RG, Kwitek-Black AE, Glenn C, Gullings-Handley J, Van Etten W, Atkinson OS et al. A high-density integrated genetic linkage and radiation hybrid map of the laboratory rat. Genome Res 1999; 9: AP1–8, insert.
Dracheva SV, Remmers EF, Chen S, Chang L, Gulko PS, Kawahito Y et al. An integrated genetic linkage map with 1,137 markers constructed from five F2 crosses of autoimmune disease-prone and -resistant inbred rat strains. Genomics 2000; 63: 202–226.
Watanabe TK, Ono T, Okuno S, Mizoguchi-Miyakita A, Yamasaki Y, Kanemoto N et al. Characterization of newly developed SSLP markers for the rat. Mamm Genome 2000; 11: 300–305.
Bihoreau MT, Sebag-Montefiore L, Godfrey RF, Wallis RH, Brown JH, Danoy PA et al. A high-resolution consensus linkage map of the rat, integrating radiation hybrid and genetic maps. Genomics 2001; 75: 57–69.
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE et al. MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1987; 1: 174–181.
Canzian F . Phylogenetics of the laboratory rat Rattus norvegicus. Genome Res 1997; 7: 262–267.
Hendley ED, Ohlsson WG . Two new inbred rat strains derived from SHR: WKHA, hyperactive, and WKHT, hypertensive, rats. Am J Physiol 1991; 261: H583–H589.
Hendley ED . WKHA rats with genetic hyperactivity and hyperreactivity to stress: a review. Neurosci Biobehav Rev 2000; 24: 41–44.
Gershenfeld HK, Paul SM . Mapping quantitative trait loci for fear-like behaviors in mice. Genomics 1997; 46: 1–8.
Crabbe JC, Wahlsten D, Dudek BC . Genetics of mouse behavior: interactions with laboratory environment [see comments]. Science 1999; 284: 1670–1672.
Zhang QY, Dene HY, Deng AY, Garrett MR, Jacob HJ, Rapp JP . Interval mapping and congenic strains for a blood pressure QTL on rat chromosome 13. Mamm Genome 1997; 8: 636–641.
Dukhanina OI, Dene HY, Deng AY, Choi CR, Hoebee B, Rapp JP . Linkage map and congenic strains to localize blood pressure QTL on rat chromosome 10. Mamm Genome 1997; 8: 229–235.
Podolin PL, Denny P, Armitage N, Lord CJ, Hill NJ, Levy ER et al. Localization of two insulin-dependent diabetes (Idd) genes to the Idd10 region on mouse chromosome 3. Mamm Genome 1998; 9: 283–286.
Frantz S, Clemitson JR, Bihoreau MT, Gauguier D, Samani NJ . Genetic dissection of region around the Sa gene on rat chromosome 1: evidence for multiple loci affecting blood pressure. Hypertension 2001; 38: 216–221.
Darvasi A . Experimental strategies for the genetic dissection of complex traits in animal models. Nat Genet 1998; 18: 19–24.
Watanabe TK, Bihoreau MT, McCarthy LC, Kiguwa SL, Hishigaki H, Tsuji A et al. A radiation hybrid map of the rat genome containing 5,255 markers [see comments]. Nat Genet 1999; 22: 27–36.
Miner LL, Marley RJ . Chromosomal mapping of the psychomotor stimulant effects of cocaine in BXD recombinant inbred mice. Psychopharmacology (Berlin) 1995; 122: 209–214.
Mathis C, Neumann PE, Gershenfeld H, Paul SM, Crawley JN . Genetic analysis of anxiety-related behaviors and responses to benzodiazepine-related drugs in AXB and BXA recombinant inbred mouse strains. Behav Genet 1995; 25: 557–568.
Grisel JE, Belknap JK, O'Toole LA, Helms ML, Wenger CD, Crabbe JC . Quantitative trait loci affecting methamphetamine responses in BXD recombinant inbred mouse strains. J Neurosci 1997; 17: 745–754.
Lander E, Kruglyak L . Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 1995; 11: 241–247.
Faraone SV, Doyle AE, Mick E, Biederman J . Meta-analysis of the association between the 7-repeat allele of the dopamine D(4) receptor gene and attention deficit hyperactivity disorder. Am J Psychiatry 2001; 158: 1052–1057.
Roman T, Schmitz M, Polanczyk GV, Eizirik M, Rohde LA, Hutz MH . Further evidence for the association between attention-deficit/hyperactivity disorder and the dopamine-[beta]-hydroxylase gene. Am J Med Genet 2002; 114: 154–158.
Courvoisier H, Moisan MP, Sarrieau A, Hendley ED, Mormède P . Behavioral and neuroendocrine profile of stress reactivity in the WKHA/WKY inbred rat strains: a multifactorial and genetic analysis. Brain Res 1996; 743: 77–85.
Manly KF, Cudmore Jr RH, Meer JM . Map Manager QTX, cross-platform software for genetic mapping. Mamm Genome 2001; 12: 930–932.
Acknowledgements
This work was supported by the European Community Program Biotechnology, Grant BIO4CT 96562 to MPM. We thank Y Mellerin and M Duriez for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Moisan, MP., Llamas, B., Cook, M. et al. Further dissection of a genomic locus associated with behavioral activity in the Wistar–Kyoto hyperactive rat, an animal model of hyperkinesis. Mol Psychiatry 8, 348–352 (2003). https://doi.org/10.1038/sj.mp.4001234
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.mp.4001234
Keywords
This article is cited by
-
The influence of sex and estrous cycle on QTL for emotionality and ethanol consumption
Mammalian Genome (2011)
-
QTL mapping for traits associated with stress neuroendocrine reactivity in rats
Mammalian Genome (2005)
-
Lineage is an Epigenetic Modifier of QTL Influencing Behavioral Coping with Stress
Behavior Genetics (2005)