Abstract
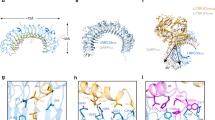
Recognition by integrin proteins on the cell surface regulates the adhesive interactions between cells and their surroundings1,2. The structure of the 'I' domain that is found in some but not all integrins, has been determined3,4. However, the only integrin ligands for which structures are known, namely fibronectin and VCAM-1 (refs 5–7), are recognized by integrins that lack I domains. The intercellular adhesion molecules ICAM-1, 2 and 3 are, like VCAM-1, members of the immunoglobulin superfamily (IgSF), but they are recognized by an I domain-containing integrin, lymphocyte-function-associated antigen 1 (LFA-1, or CD 11 a/CD 18). Here we present the crystal structure of the extracellular region of ICAM-2. The glutamic acid residue at position 37 is critical for LFA-1 binding and is proposed to coordinate the Mg2+ ion in the I domain; this Glu 37 is surrounded by a relatively flat recognition surface and lies in a β-strand, whereas the critical aspartic acid residue in VCAM-1 and fibronectin lie in protruding loops. This finding suggests that there are differences in the architecture of recognition sites between integrins that contain or lack I domains. A bend between domains 1 and 2 of ICAM-2 and a tripod-like arrangement of N-linked glycans in the membrane-proximal region of domain 2 may be important for presenting the recognition surface to LFA-1. A model of ICAM-1 based on the ICAM-2 structure provides a framework for understanding its recognition by pathogens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Hynes, R. O. Integrins: Versatility, modulation, and signaling in cell adhesion. Cell 69, 11–25 (1992).
Springer, T. A. Traffic signals for lymphocyte recirculation and leukocyte emigration: The multi-step paradigm. Cell 76, 301–314 (1994).
Lee, J.-O., Rieu, P., Arnaout, M. A. & Liddington, R. Crystal structure of the A domain from the α-subunit of integrin CR3 (CD11b/CD18). Cell 80, 631–638 (1995).
Qu, A. & Leahy, D. J. Crystal structure of the I-domain from the GD11a/CD18 (LFA-1, a1b2) integrin. Proc. Natl Acad. Sci. USA 92, 10277–10281 (1995).
Leahy, D. J., Aukhil, I. & Erickson, H. P. 2.0 Å crystal structure of a four-domain segment of human fibronectin encompassing the RGD loop and synergy region. Cell 84, 155–164 (1996).
Jones, E. Y. et al. Crystal structure of an integrin-binding fragment of vascular cell adhesion molecule-1 at 1.8 Å resolution. Nature 373, 539–544 (1995).
Wang, J.-h. et al. The crystal structure of an N-terminal two domain fragment of VCAM-1: A cyclic peptide based on the domain 1 C-D loop can inhibit VCAM-l/α4 integrin. Proc. Natl Acad. Sci. USA 92, 5714–5718 (1995).
Stanley, P. Chinese hamster ovary cell mutants with multiple glycosylation defects for production of glycopoteins with minimal carbohydrate heterogeneity. Mol. Cell. Biol. 9, 377–383 (1989).
Staunton, D. E., Dustin, M. L., Erickson, H. P. & Springer, T. A. The arrangement of the immunoglobulin-like domains of ICAM-1 and the binding sites for LFA-1 and rhinovirus. Cell 61, 243–254 (1990).
Holness, C. L. et al. Analysis of the binding site on intercellular adhesion molecule 3 for the leukocyte integrin lymphocyte function-associated antigen 1. J. Biol. Chem. 270, 877–884 (1995).
Klickstein, L. B., York, M. B., de Fougerolles, A. R. & Springer, T. A. Localization of the binding site on intercellular adhesion molecule-3 (ICAM-3) for lymphocyte function-associated antigen-1 (LFA-1). J. Biol. Chem. 271, 23920–23927 (1996).
Sadhu, C. et al. LFA-1 binding site in ICAM-3 contains a conserved motif and non-contiguous amino acids. Cell Adhes Commun. 2, 429–440 (1994).
Harpaz, Y. & Chothia, C. Many of the immunoglobulin superfamily domains in cell adhesion molecules and surface receptors belong to a new structural set which is close to that containing variable domains. J. Mol. Biol. 238, 528–539 (1994).
Williams, A. F. A year in the life of the immunoglobulin superfamily. Immunol. Today 8, 298–303 (1987).
Wang, J.-h. et al. Structure of a functional fragment of VCAM-1 refined at 1.9 Å resolution. Acta Crystallogr. 52, 369–379 (1996).
Bodian, D. L., Jones, E. Y., Harlos, K., Stuart, D. I. & Davis, S. J. Crystal structure of the extracellular region of the human cell adhesion molecule CD2 at 2.5 Å resolution. Structure 2, 755–766 (1995).
Musafia, B., Buchner, V. & Arad, D. Complex salt bridges in proteins: Statistical analysis of structure and function. J. Mol. Biol. 254, 761–770 (1995).
Huang, C. & Springer, T. A. A binding interface on the I domain of lymphocyte function associated antigen-1 (LFA-1) required for specific interaction with intercellular adhesion molecule 1 (ICAM-1). j. Biol. Chem. 270, 19008–19016 (1995).
Springer, T. A. Folding of the N-terminal, ligand-binding region of integrin α-subunits into a β-propeller domain. Proc. Natl Acad. Sci. USA 94, 65–72 (1997).
Greve, J. M. et al. The major human rhinovirus receptor is ICAM-1. Cell 56, 839–847 (1989).
Staunton, D. E. et al. A cell adhesion molecule, ICAM-1, is the major surface receptor for rhinoviruses. Cell 56, 849–853 (1989).
Berendt, A. R., Simmons, D. L., Tansey, J., Newbold, C. I. & Marsh, K. Intercellular adhesion molecule-1 is an endothelial cell adhesion receptor for Plasmodium falciparum. Nature 341, 57–59 (1989).
McClelland, A. et al. Identification of monoclonal antibody epitopes and critical residues for rhinovirus binding in domain 1 of intercellular adhesion molecule 1. Proc. Natl Acad. Sci. USA 88, 7993–7997 (1991).
Register, R. B., Uncapher, C. R., Naylor, A. M., Lineberger, D. W. & Colonno, R. J. Human-murine chimeras of ICAM-1 identify amino acid residues critical for rhinovirus and antibody binding. J. Virol. 65, 6589–6596 1991).
Ockenhouse, C. F., Betageri, R., Springer, T. A. & Staunton, D. E. Plasmodium falciparum-infected erythrocytes bind ICAM-1 at a site distinct from LFA-1, Mac-1, and human rhinovirus. Cell 68, 63–69 (1992).
Berendt, A. R. et al. The binding site on ICAM-1 for plasmodium falciparum-infected erythorcytes overlaps, but is distinct from, the LFA-1-binding site. Cell 68, 71–81 (1992).
Staunton, D. E., Dustin, M. L. & Springer, T. A. Functional cloning of ICAM-2, a cell adhesion ligand for LFA-1 homologous to ICAM-1. Nature 339, 61–64 (1989).
Casasnovas, J. M. & Springer, T. A. Kinetics and thermodynamics of virus binding to receptor: Studies with rhinovirus, intercellular adhesion molecule-1 (ICAM-1), and surface plasmon resonance. J. Biol. Chem. 270, 13216–13224 (1995).
Carson, M. Ribbon models of macromolecules. J. Mol. Graph. 5, 103–106 (1987).
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association; insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Sali, A. Thesis, Univ. London, 1991.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Casasnovas, J., Springer, T., Liu, Jh. et al. Crystal structure of ICAM-2 reveals a distinctive integrin recognition surface. Nature 387, 312–315 (1997). https://doi.org/10.1038/387312a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/387312a0
This article is cited by
-
Direction of actin flow dictates integrin LFA-1 orientation during leukocyte migration
Nature Communications (2017)
-
ICAM-2 confers a non-metastatic phenotype in neuroblastoma cells by interaction with α-actinin
Oncogene (2015)
-
Crystal structure and its bearing towards an understanding of key biological functions of EpCAM
Nature Communications (2014)
-
N-glycosylation of ICAM-2 is required for ICAM-2-mediated complete suppression of metastatic potential of SK-N-AS neuroblastoma cells
BMC Cancer (2013)
-
Solution structure of the X4 protein coded by the SARS related coronavirus reveals an immunoglobulin like fold and suggests a binding activity to integrin I domains
Journal of Biomedical Science (2006)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.