Abstract
Giant cell tumor of the bone (GCTOB) is a primary bone tumor that occurs mainly in young adults and is capable of locally aggressive growth. Its histologic appearance can resemble a number of benign and malignant tumors but no useful diagnostic marker is known currently. To identify diagnostic markers for this tumor, global gene expression profiling using cDNA microarray was performed on 6 fresh-frozen GCTOB, 3 aneurysmal bone cysts, 4 fibrous dysplasias and 12 giant cell tumors of tendon sheath/diffuse-type giant cell tumors. Unsupervised hierarchical clustering separated the tumors based on their histopathologic types, and significance analysis of microarray identified several genes including TP73L (encoding the p63 protein) that are significantly highly expressed in GCTOB relative to these other tumors. The diagnostic utility of p63 was subsequently confirmed using anti-p63 antibody on a series of 26 GCTOB, 25 aneurysmal bone cysts, 15 chondroblastomas, 13 giant cell reparative granulomas, 13 chondromyxoid fibromas, 4 brown tumors, 4 fibrous dysplasias, 53 giant cell tumors of tendon sheath/diffuse-type giant cell tumors and 385 additional mesenchymal tumors in tissue microarrays. Strong p63 nuclear staining was present in 18 of 26 (69%) GCTOB, 3 of 15 (20%) chondroblastomas and in 1 of 25 (4%) aneurysmal bone cysts while none of the other tumors commonly considered in the differential diagnosis of GCTOB showed any detectable p63 staining. Strong p63 staining is rare in bone and soft-tissue tumors in general. In contrast to the pattern of p63 staining, the majority of the chondroblastomas (70%) demonstrated S-100 immunoreactivity while only a minority of the GCTOB (8%) was immunoreactive for S-100. These findings altogether show that p63 can be used as a diagnostic marker to aid the clinical diagnosis of GCTOB.
Similar content being viewed by others
Main
Giant cell tumor of the bone (GCTOB) is an osteolytic neoplasm that accounts for about 5% of primary bone tumors. It most commonly arises at the epiphyses of long bones like the distal femur, proximal tibia, distal radius and proximal humerus and it predominantly affects young adults after the closure of the epiphyseal growth plate.1 Although the majority of GCTOB follows a benign clinical course, a subset can behave in a locally aggressive manner causing destruction of cortical bone with extension to the adjacent soft tissues. The rate of local recurrence following surgical curettage is relatively high at about 25% and limb–salvage surgery may be required in some of these instances.1 Metastases particularly to the lung can also occur in 2% of the patients and these cases can exhibit otherwise typical histological features.
GCTOB is mainly comprised of three cell types—mononuclear stromal cells, multinucleated osteoclast-like giant cells and monocytes.2, 3 The mononuclear stromal cell represents the neoplastic component of the tumor whereas monocytes represent a minor component of the mononuclear cell population.1, 2, 4 Histologically, these neoplastic mononuclear stromal cells can assume a number of appearances ranging from round polygonal cells to spindle-shaped cells growing in a storiform pattern. There may also be areas of hemorrhage, fibrosis, ossification, necrosis and xanthomatous change. In addition, areas showing secondary aneurysmal bone cysts (ABC) changes can be found in about 10% of GCTOB.1 These variations can add to the challenge of the histologic differentiation of GCTOB from other neoplasms such as ABC, brown tumors, giant cell reparative granulomas (GCRG), chondroblastomas, chondromyxoid fibromas, undifferentiated pleomorphic sarcomas and osteosarcomas. Although radiological assessment is frequently used to aid in the diagnosis of GCTOB, there remain cases that are difficult to determine. Furthermore, there is currently no well-accepted diagnostic marker available for GCTOB. Given that some GCTOB can behave in an aggressive manner, distinction from more benign-behaving tumors like ABC, GCRG and chondromyxoid fibromas is important clinically.
In this study, we used global gene expression analysis to examine the gene expression profiles of GCTOB relative to other primary bone tumors and tumors characterized by abundant giant cells in an effort to identify novel diagnostic markers for GCTOB. This led to the identification and immunohistochemical confirmation of p63 by tissue microarray as a diagnostic marker that can be used to aid in the diagnosis of GCTOB.
Materials and methods
Fresh-Frozen Tumor and Paraffin-Embedded Tumor Samples
Fresh-frozen samples of six cases of giant cell tumor of bone (GCTOB), four cases of fibrous dysplasias (FD) and three cases of ABC were obtained from specimens resected at Vancouver General Hospital between 1996 and 2002. The clinicopathologic features of these tumors are shown in Table 1. The 12 cases of giant cell tumor of tendon sheath/diffuse-type giant cell tumor (GCTTS/DTGCT) were profiled as described previously.5 Additional paraffin-embedded tumor samples from 26 cases of GCTOB, 25 cases of ABC, 15 cases of chondroblastomas, 12 cases of GCRG, 12 cases of chondromyxoid fibromas, 4 cases of hyperparathyroidism associated brown tumors, 4 cases of FD and 53 cases of GCTTS/DTGCT were collected from Stanford University Medical Center and Vancouver General Hospital. All cases of primary bone tumors were diagnosed by experienced subspecialty bone and soft-tissue pathologists at the respective centers and reviewed by a study pathologist (TON or RBW). The collection of the paraffin-embedded tumor samples for other mesenchymal tumors was described previously.6 The studies were performed with the approval from the Institutional Review Boards at Stanford University Medical Center and Vancouver General Hospital.
RNA Extraction and cDNA Microarray Analysis
The microarrays used in the study contain a total of about 42 000 cDNAs representing about 28 000 genes or ESTs printed on polylysine-coated glass slides by the Stanford Functional Genomics Facility (http://www.microarray.org/). Details of microarray construction were described previously.7, 8 After confirmation of the presence of viable tumor by frozen section, the frozen tissue was homogenized in Trizol reagent (Invitrogen, Carlsbad, CA, USA) and total RNA was extracted. Preparation of Cy-3-dUTP-labeled cDNA from reference RNA (Stratagen, Universal human reference RNA, catalog no. 740000) and Cy-5-dUTP-labeled cDNA from each tumor specimen, microarray hybridization and washing of arrays were performed as described by Perou et al.8 Microarrays were scanned on a GenePix 4000 microarray scanner (Axon Instruments, Foster City, CA, USA) and fluorescence ratios (tumor/reference) were calculated using GenePix software. The raw data and the image files are available from the Stanford Microarray Database (http://smd.stanford.edu/).9 Only cDNA spots with a regression correlation of greater than 0.6 and a ratio of signal over background of at least 1.5 in either the Cy3 or Cy5 channel were included. Array and gene centering were applied to the expression values. For subsequent data analysis, a set of standard filtering criteria was employed that included only those genes that had 70% available good data and showed a minimum of four-fold absolute difference in gene expression levels compared to the mean expression level across all samples for each gene in at least two samples. The filtered data set is shown in web supplement Table 1.
Unsupervised hierarchical clustering analysis was used to produce dendrograms that depict the degree of relatedness between tumor specimens based on their gene expression profiles. Significance analysis of microarrays (SAM) was performed to identify differentially expressed genes between groups of tumor specimens.10 Gene set enrichment analysis (GSEA) was performed as described by Subramanian et al11 to identify specific pathways that are significantly affected in the tumor specimens.
Tissue Microarray
Five tissue microarrays (TA-92, TA-137, TA-186, TA-216 and TA03-008) were constructed as previously published using a manual tissue arrayer (Beecher instruments, Silver Spring, MD, USA).12 For each specimen one to two 2-mm tissue cores were taken from areas in formalin-fixed, paraffin-embedded tumor blocks that were representative of the tumor. Known positive and negative controls were added to the TMA, consisting of normal squamous epithelium, transitional epithelium and lymph node. The construction of the tissue microarrays (TA-38 and TA-39) for a large series of 385 mesenchymal tumors was described previously.6
Immunohistochemical Analysis
Monoclonal mouse antibody raised against human p63 protein (Dako antibody, M7247, clone 4A4, Carpenteria, CA, USA) was used at 1:200 dilution with citrate pH 6.0 antigen retrieval in a pressure cooker and Dako automated staining and polyclonal rabbit antibody against S-100 protein (Dako antibody, Z311, Carpenteria, CA, USA) was used at 1:4000 dilution with Ventana Benchmark stainer. For p63, staining was interpreted as negative when less than 10% of the tumor cells showed light nuclear staining. A score of ‘weak positive’ was given for faint nuclear staining above background in at least 10% of the tumor cells. A score of ‘strong positive’ was given for strong nuclear staining in at least 10% of the tumor cells. The intensity of the strong nuclear p63 staining observed in the positive cases is similar to that seen in the positive controls (strong p63 nuclear staining present in basal layers of squamous epithelium or transitional epithelium). Only nuclear staining was evaluated. For S-100, a score of ‘weak positive’ was given for faint cytoplasmic/nuclear staining in at least 10% of the tumor cells while a score of ‘strong positive’ was given for strong cytoplasmic/nuclear staining in at least 10% of the tumor cells. The higher score was used in duplicate cores that demonstrated different scores of immunostaining. Cores in which no diagnostic material was present or with equivocal staining results were omitted from further analysis. Scoring results were combined using Deconvoluter and Combiner programs.13, 14
Results
Gene Expression Profiling Study
Global gene expression analysis was performed for 6 giant cell tumors of the bone (GCTOB), 3 ABC, 4 FD and 12 giant cell tumors of tendon sheath/diffuse-type giant cell tumors (GCTTS/DTGCT). The 3 ABC, 4 FD and 12 GCTTS/DTGCT were included for comparison to identify genes that were differentially expressed in GCTOB vs other benign primary bone tumors and other mesenchymal tumors enriched with multinucleated giant cells. The gene expression profiles of the 12 GCTTS/DTGCT have been published previously.5 The clinicopathologic features the six GCTOB, three ABC and four FD are shown in Table 1. All of these bone tumors represented primary lesions. The six cases of GCTOB occurred at the ends of long bones and showed classic histologic features.
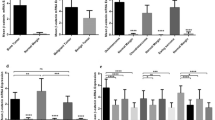
The dendrogram produced by unsupervised hierarchical clustering using a set of 1018 genes filtered by the criteria described in the Materials and methods section is shown in Figure 1. The hierarchical clustering separated all primary bone tumors from the 12 cases of GCTTS/DTGCT. Within the primary bone tumor cluster, the 6 GCTOB are clustered separately from the 3 ABC and 4 FD. SAM was performed to identify the differentially expressed genes that distinguish one tumor type from the others. A selected list of the differentially expressed genes with a false discovery rate (FDR) of less than 5% is shown in Table 2 with the complete list shown in web supplement Table 2. GCTOB showed a high level of expression for 85 genes and decreased level of expression for 52 genes (FDR<5%) relative to other tumors. When compared to the other specimens, many of the highly expressed genes like CA2, MMP13, CKB, TNFSF11, VNN1 and COL10A1 were found in earlier studies to be highly expressed in GCTOB.15, 16 TNFSF11 or ligand for receptor activator of NFκB (RANKL) is known to be expressed by the mononuclear stromal cells in GCTOB and promotes osteoclast formation.17 Furthermore, several other ligands including a receptor–ligand pair—FGFR2 and FGF9—were found to be highly expressed in GCTOB. In comparison to other tumors, ABC showed relative upregulation of 55 genes but no relative downregulation of genes with an FDR<5%. A total of 838 genes were found to be differentially expressed in FD relative to other tumors in this series. Many of the overexpressed genes such as RUNX2, BMP2, SPARC, TNFRSF11B and ALPL are known to be important in the process of bone resorption and formation.18 GSEA was performed to identify specific pathways that are upregulated in individual tumor types and the NFκB pathway showed relative upregulation in GCTOB although statistical significance was not reached (P=0.08). This may account in part for the abundance of osteoclast-like multinucleated giant cells usually seen in GCTOB because NFκB together with c-fos and RANKL regulate the differentiation of osteoclasts from preosteoclasts.17, 18
Interestingly, TP73L whose protein product is p63 was among the significantly upregulated genes identified for GCTOB even though none of the earlier gene expression profiling studies on GCTOB identified p63 as being highly expressed.15, 16, 19 A closer examination of the TP73L expression in this series of tumors revealed that its expression was the highest in four of the six GCTOB whereas the other two GCTOB showed similar levels of TP73L expression as seen in ABC and FD. GCTTS/DTGCT showed a much lower level of TP73L in contrast to GCTOB.
p63 Expression in GCTOB
A commercial antibody against p63 is routinely used in most surgical pathology laboratories for a variety of diagnostic purposes but no detailed analysis of p63 expression in mesenchymal tumors has been published. To verify the gene expression findings, tissue microarrays that contained 26 GCTOB, 25 ABC, 4 FD, 53 GCTTS/DTGCT as well as 15 chondroblastomas, 12 GCRG, 12 chondromyxoid fibromas and 4 brown tumors were created. The 26 GCTOB included all 6 cases that were analyzed by gene arrays. The clinicopathologic features for the bone tumors in this TMA are shown in web supplement Table 3. The 26 GCTOB were obtained from 25 patients and included 23 primary and 3 recurrent tumors with two cases from the same patient (one primary and one recurrent tumor). The average age at the time of diagnosis was 34.5 years with a male-to-female ratio of 3:2. The tumors occurred at the ends of the long bones (n=15), the small bones of the distal extremities (n=6) and the axial skeleton (n=4). The 25 cases of ABC contained 22 primary tumors and 3 recurrent tumors. The most common sites of involvement by ABC in our current series were the tibia and femur. The 15 chondroblastomas, 12 GCRG and 12 chondromyxoid fibromas all represented primary lesions. The most common sites of involvement for chondroblastomas in our current series were the proximal femur, radius and humerus. All except three cases of GCRG occurred in the mandible. A total of 7 out of the 12 chondromyxoid fibromas occurred in the long bones of the limbs with the remainder arising from the pelvic bone predominantly.
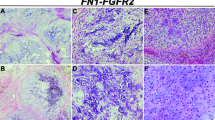
The results of the immunohistochemical analysis are shown in Table 3. Strong p63 nuclear staining was observed in 18 of 26 GCTOB (69%), 3 of 15 chondroblastomas (20%), 1 of 25 ABC (4%), 0 of 12 GCRG (0%), 0 of 12 chondromyxoid fibromas (0%), 0 of 4 brown tumors (0%), 0 of 4 FD (0%) and 0 of 53 GCTTS/DTGCT (0%). The staining was observed only in the nucleus of the mononuclear cells and no staining was present in the multinucleated giant cells (Figure 2). There was one case of GCTOB for which both the primary and the recurrent tumors from the same site were available and both tumor samples expressed strong p63 nuclear staining. Of the six cases of GCTOB that were analyzed by gene arrays, the same four GCTOB that showed high p63 gene expression also demonstrated positive p63 expression by immunohistochemistry (strong positive in three cases and weak positive in one case) whereas the two GCTOB with low p63 gene expression were both negative for p63 protein expression by immunohistochemistry. There was a predominance of male patients among the eight cases of GCTOB that exhibited weak/absent nuclear p63 staining but no other significant clinicopathologic differences were noted between the group with strong p63-positive staining and the group with weak/absent p63 staining. The single case of ABC that demonstrated strong nuclear p63 staining showed no unusual features histologically or radiologically and was considered a classic case of ABC. All three cases of chondroblastomas that showed strong p63 nuclear staining also demonstrated positive S-100 immunostaining in the tumor cells (Figure 2g and h). In contrast, only 2 of 26 (8%) GCTOB in our series demonstrated weak positive staining for S-100 (web supplement Table 4). Furthermore, the two GCTOB specimens that showed S-100 immunoreactivity represent primary and recurrent diseases from the same patient and both tumors show strong p63 nuclear staining. Overall, in keeping with our current understanding,1 70% of the chondroblastomas included in our study demonstrated positive S-100 immunostaining.
Representative results of immunostaining in GCTOB, ABC, GCRG and chondroblastomas. (a and b) Strong nuclear p63 staining observed in two cases of GCTOB. The nuclear staining is restricted to the mononuclear stromal cells and not present in the multinucleated giant cells. (c) Weak nuclear p63 staining observed in a case of GCTOB. (d) Weak nuclear p63 staining observed in a case of ABC. (e) Negative nuclear p63 staining observed in a case of ABC. (f) Negative nuclear p63 staining observed in a case of GCRG. (g) Strong nuclear p63 staining observed in a case of chondroblastoma. (h) Strong S-100 staining observed in the same case of chondroblastoma.
p63 Expression in Other Mesenchymal Tumors
For a more comprehensive examination of the diagnostic utility of p63 for GCTOB, we also examined other primary bone tumors such as osteosarcomas and other soft-tissue tumors such as undifferentiated pleomorphic sarcomas and leiomyosarcoma that can either directly arise in bone or involve the bone secondarily. We studied immunohistochemical expression of p63 in 385 mesenchymal tumors that encompassed 32 different types of soft-tissue, bone and cartilage tumors (Table 4). Nuclear p63 immunoreactivity was infrequent in this series of tumors. Strong nuclear p63 staining was found only in 2 cases (1 of 60 undifferentiated pleomorphic sarcomas and 1 of 19 synovial sarcomas) whereas weak nuclear p63 staining was found in 8 additional cases that included 2 of 13 osteosarcomas (15%). The current series included one case of giant cell rich osteosarcoma, which showed no p63 immunostaining. Occasional cases of undifferentiated pleomorphic sarcomas and leiomyosarcomas showed faint cytoplasmic staining in the tumor cells but only nuclear staining was considered in our current analysis. These findings confirm that strong p63 nuclear staining is specific for GCTOB in mesenchymal tumors.
Discussion
GCTOB is classified as a benign bone tumor but has a tendency to recur and can behave in an aggressive manner locally or even metastasize to the lungs in a small subset of cases. The differential diagnosis includes a range of diagnostic entities from benign tumors like ABC to highly malignant tumors like undifferentiated pleomorphic sarcomas. Although both morphologic and radiologic examination can lead to the correct diagnosis in the majority of cases, the diagnosis of GCTOB can be difficult in some cases and no useful diagnostic marker is currently available clinically to aid in its diagnosis. Combining global gene expression and immunohistochemical analysis, we have shown that strong nuclear p63 expression in the mononuclear stromal tumor cells is present in about two-thirds of GCTOB and is rare in other bone and soft-tissue tumors.
The finding of strong TP73L gene and p63 protein expression in the neoplastic stromal cells of GCTOB is rather unexpected as it has no known role in the development of normal cartilage or bone. TP73L (encoding the p63 protein) belongs to the p53 gene family and possess several splice variants.20, 21, 22 It is required developmentally for limb and skin formation.21, 23 Unlike p53, which is an important tumor suppressor that is frequently inactivated in many tumor types, the precise role(s) of p63 in oncogenesis is less clear. There is however increasing realization that p63 may be a bifunctional cancer gene in that it can suppress tumorigenesis in some cell types while promoting tumorigenesis in others.20 The specific role of p63 in GCTOB, if any, therefore remains to be determined.
p63 expression was not reported as a useful marker for GTOB in any of the earlier gene array analyses.15, 16, 19 Thomas and co-workers have recently published the gene expression profile of 9 GCTOB together with a series of undifferentiated pleomorphic sarcomas, liposarcomas, leiomyosarcomas and synovial sarcomas for comparison.15 Although they found relatively high expression of TP73L in 3 of the 9 GCTOB, they also noted expression in 3 of 11 undifferentiated pleomorphic sarcomas, 3 of 9 leiomyosarcomas and 1 of 4 synovial sarcomas with similarly high expression levels for TP73L. TP73L was therefore not identified as a significant marker for GCTOB in their study. We did not compare the gene array findings of GTOB to pleomorphic sarcoma, LMS and SS but our immunohistochemical data showed strong p63 nuclear staining in only 1 of 60 undifferentiated pleomorphic sarcomas and 1 of 19 synovial sarcomas, and weak p63 nuclear staining in 3 of 42 leiomyosarcomas. The faint cytoplasmic immunohistochemical p63 staining observed by us in some cases of undifferentiated pleomorphic sarcomas and leiomyosarcomas may contribute at least partially to the difference between gene expression and immunohistochemical data.
Although the expression of TP73L or p63 in various normal epithelium and epithelial malignancies is well characterized,22 very little is known about the expression of p63 in mesenchymal tumors. Our study of 32 different types of mesenchymal tumors identified only a few rare cases (2 of 385) that demonstrated strong nuclear p63 staining. It is likely that the p63 expression in these rare cases represent an aberrant process that is not related to the predominant oncogenic process occurring in these tumors. As a diagnostic marker for GCTOB, strong nuclear p63 immunopositivity was found in about two-thirds of GCTOB and therefore lacks the ideal sensitivity for diagnosing GCTOB. However, with the exception for chondroblastomas, strong nuclear p63 immunopositivity is rarely present in many benign and malignant tumors that may enter into the differential diagnosis of GCTOB such as ABC, GCRG, FD, brown tumors, chondromyxoid fibromas, osteosarcomas and undifferentiated pleomorphic sarcomas. It is therefore associated with a good specificity for identifying GCTOB. For that reason, the observation of strong nuclear p63 staining in a tumor with a possible diagnosis of GCTOB may lend further support to the diagnosis of GCTOB. If chondroblastoma is a consideration, S-100 can also be of help as the majority of chondroblastomas in our current series, in accordance with the literature, shows positive S-100 immunostaining whereas only occasional weak S-100 immunoreactivity is seen in GCTOB. A combination of strong nuclear p63 staining together with negative S-100 in the mononuclear tumor cells would strongly suggest a diagnosis of GCTOB while the finding of strong S-100 staining only would strongly favor a diagnosis of chondroblastoma.
In summary, we have found through gene expression profiling analysis that p63 is highly expressed in the majority of GCTOB. We validated our findings with p63 immunohistochemical analysis of a larger series of GCTOB and histologically similar lesions and found that strong nuclear p63 immunoreactivity can be seen in about two-thirds of GCTOB and is uncommon in other bone and soft-tissue tumors. The utilization of p63 as a marker for GCTOB may help in the histological diagnosis of GCTOB.
References
Reid R, Banerjee SS, Sciot R . Giant cell tumour. In: Fletcher CDM, Unni KK, Martens F (eds). World Health Organization Classification of Tumours, Pathology and Genetics of Tumours of Soft Tissue and Bone. IARC Press: Lyon, 2002, pp 310–312.
Wulling M, Engels C, Jesse N, et al. The nature of giant cell tumor of bone. J Cancer Res Clin Oncol 2001;127:467–474.
Zheng MH, Robbins P, Xu J, et al. The histogenesis of giant cell tumour of bone: a model of interaction between neoplastic cells and osteoclasts. Histol Histopathol 2001;16:297–307.
Wulling M, Delling G, Kaiser E . The origin of the neoplastic stromal cell in giant cell tumor of bone. Hum Pathol 2003;34:983–993.
West RB, Rubin BP, Miller MA, et al. A landscape effect in tenosynovial giant-cell tumor from activation of CSF1 expression by a translocation in a minority of tumor cells. Proc Natl Acad Sci USA 2006;103:690–695.
Nguyen TT, Schwartz EJ, West RB, et al. Expression of CD163 (hemoglobin scavenger receptor) in normal tissues, lymphomas, carcinomas, and sarcomas is largely restricted to the monocyte/macrophage lineage. Am J Surg Pathol 2005;29:617–624.
Perou CM, Jeffrey SS, van de Rijn M, et al. Distinctive gene expression patterns in human mammary epithelial cells and breast cancers. Proc Natl Acad Sci USA 1999;96:9212–9217.
Perou CM, Sorlie T, Eisen MB, et al. Molecular portraits of human breast tumours. Nature 2000;406:747–752.
Sherlock G, Hernandez-Boussard T, Kasarskis A, et al. The Stanford Microarray Database. Nucleic Acids Res 2001;29:152–155.
Tusher VG, Tibshirani R, Chu G . Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 2001;98:5116–5121.
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005;102:15545–15550.
West RB, Corless CL, Chen X, et al. The novel marker, DOG1, is expressed ubiquitously in gastrointestinal stromal tumors irrespective of KIT or PDGFRA mutation status. Am J Pathol 2004;165:107–113.
Liu CL, Montgomery KD, Natkunam Y, et al. TMA-combiner, a simple software tool to permit analysis of replicate cores on tissue microarrays. Mod Pathol 2005;18:1641–1648.
Liu CL, Prapong W, Natkunam Y, et al. Software tools for high-throughput analysis and archiving of immunohistochemistry staining data obtained with tissue microarrays. Am J Pathol 2002;161:1557–1565.
Morgan T, Atkins GJ, Trivett MK, et al. Molecular profiling of giant cell tumor of bone and the osteoclastic localization of ligand for receptor activator of nuclear factor kappaB. Am J Pathol 2005;167:117–128.
Wuelling M, Delling G, Kaiser E . Differential gene expression in stromal cells of human giant cell tumor of bone. Virchows Arch 2004;445:621–630.
Huang L, Xu J, Wood DJ, Zheng MH . Gene expression of osteoprotegerin ligand, osteoprotegerin, and receptor activator of NF-kappaB in giant cell tumor of bone: possible involvement in tumor cell-induced osteoclast-like cell formation. Am J Pathol 2000;156:761–767.
Cohen Jr MM . The new bone biology: pathologic, molecular, and clinical correlates. Am J Med Genet 2006;140:2646–2706.
Guenther R, Krenn V, Morawietz L, et al. Giant cell tumors of the bone: molecular profiling and expression analysis of Ephrin A1 receptor, Claudin 7, CD52, FGFR3 and AMFR. Pathol Res Pract 2005;201:649–663.
Mills AA . p63: oncogene or tumor suppressor? Curr Opin Genet Dev 2006;16:38–44.
Moll UM, Slade N . p63 and p73: roles in development and tumor formation. Mol Cancer Res 2004;2:371–386.
Westfall MD, Pietenpol JA . p63: molecular complexity in development and cancer. Carcinogenesis 2004;25:857–864.
Hermanns P, Lee B . Transcriptional dysregulation in skeletal malformation syndromes. Am J Med Genet 2001;106:258–271.
Acknowledgements
We thank Suzanne Liu for assistance in specimen handling. TON is a scholar of the Michael Smith Foundation for Health Research. The Genetic Pathology Evaluation Centre is supported by unrestricted educational grant from Sanofi-Aventis Canada.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflict of interest
The authors declare that they have no conflict of interest.
Supplementary Information accompanies the paper on Modern Pathology website (http://www.nature.com/modpathol)
Rights and permissions
About this article
Cite this article
Lee, CH., Espinosa, I., Jensen, K. et al. Gene expression profiling identifies p63 as a diagnostic marker for giant cell tumor of the bone. Mod Pathol 21, 531–539 (2008). https://doi.org/10.1038/modpathol.3801023
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3801023
Keywords
This article is cited by
-
Biphasic synovial sarcoma with myoepithelial features: a distinctive variant with a predilection for the foot
Virchows Archiv (2023)
-
A Comparative Analysis of p63 Expression in Giant Cell Tumour (GCT), Central Giant Cell Granuloma (CGCG) and Peripheral Giant Cell Granuloma (PGCG)
Head and Neck Pathology (2020)
-
Prognosis of local recurrence in giant cell tumour of bone: what can we do?
La radiologia medica (2017)
-
Correlation between p63 expression and clinical behavior of giant-cell tumor of bone
Comparative Clinical Pathology (2015)
-
Gene expression profiling of giant cell tumor of bone reveals downregulation of extracellular matrix components decorin and lumican associated with lung metastasis
Virchows Archiv (2014)