Abstract
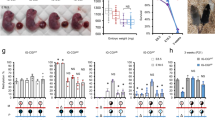
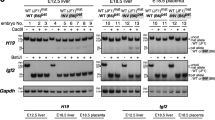
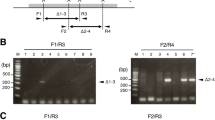
The imprinted H19 gene, which encodes an untranslated RNA, lies at the end of a cluster of imprinted genes in the mouse. Imprinting of the insulin-2 and insulin-like growth factor 2 genes, which lie about 100 kilobases upstream of H19, can be disrupted by maternal inheritance of a targeted deletion of the H19 gene and its flanking sequence. Animals inheriting the H19 mutation from their mothers are 27% heavier than those inheriting it from their fathers. Paternal inheritance of the disruption has no effect, which presumably reflects the normally silent state of the paternal gene. The somatic overgrowth of heterozygotes for the maternal deletion is attributed to a gain of function of insulin-like growth factor 2, rather than a loss of function of H19.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Solter, D. A. Rev. Genet. 22, 127–146 (1988).
Efstratiadis, A. Curr. Opin. genet. Dev. 4, 265–280 (1994).
Johnson, J. E., Birren, S. J. & Anderson, D. J. Nature 346, 858–861 (1990).
Guillemot, F. et al. Nature Genet. 9, 235–241 (1995).
Guillemot, F., Nagy, A., Auerbach, A., Rossant, J. & Joyner, A. L. Nature 371, 333–336 (1994).
Giddings, S. J., King, C. D., Harman, K. W., Flood, J. F. & Carnaghi, L. R. Nature Genet. 6, 310–313 (1994).
DeChiara, T. M., Robertson, E. J. & Efstratiadis, A. Cell 64, 849–859 (1991).
Pachnis, V., Brannan, C. I. & Tilghman, S. M. EMBO J. 7, 673–681 (1988).
Brannan, C. I., Dees, E. C., Ingram, R. S. & Tilghman, S. M. Molec. cell. Biol. 10, 28–36 (1990).
Bartolomei, M. S., Zemel, S. & Tilghman, S. M. Nature 351, 153–155 (1991).
Lee, J. E., Pintar, J. & Efstratiadis, A. Development 110, 151–159 (1990).
Poirier, F. et al. Development 113, 1105–1114 (1991).
Bartolomei, M. S., Webber, A. L., Brunkow, M. E. & Tilghman, S. M. Genes Dev. 7, 1663–1673 (1993).
Bartolomei, M. & Tilghman, S. M. Sem. devl Biol. 3, 107–117 (1992).
Ferguson-Smith, A. C., Sasaki, H., Cattanach, B. M. & Surani, M. A. Nature 362, 751–755 (1993).
Li, E., Beard, C. & Jaenisch, R. Nature 366, 362–365 (1993).
Brandeis, M. et al. EMBO J. 12, 3669–3677 (1993).
Tremblay, K. D., Saam, J., Ingram, R. S., Tilghman, S. M. & Bartolomei, M. S. Nature Genet. 9, 407–413 (1995).
McBurney, M. W. et al. Nucleic Acids Res. 19, 5755–5761 (1991).
Adra, C. N., Boer, P. H. & McBurney, M. W. Gene 60, 65–74 (1987).
Brunkow, M. E. & Tilghman, S. M. Genes Dev. 5, 1092–1101 (1991).
DeChiara, T. M., Efstratiadis, A. & Robertson, E. J. Nature 345, 78–80 (1990).
Hao, Y., Crenshaw, T., Moulton, T., Newcomb, E. & Tycko, B. Nature 365, 764–767 (1993).
Sutcliffe, J. S. et al. Nature Genet. 8, 52–58 (1994).
Reis, A. et al. Am. J. hum. Genet. 54, 741–747 (1994).
Yoo-Warren, H., Pachnis, V., Ingram, R. S. & Tilghman, S. M. Molec. cell. Biol. 8, 4707–4715 (1988).
Steenman, M. J. C. et al. Nature Genet. 7, 433–439 (1994).
Moulton, T. et al. Nature Genet. 7, 440–447 (1994).
Edlund, T., Walker, M. D., Barr, P. J. & Rutter, W. J. Science 230, 912–916 (1985).
Ohlsson, H., Thor, S. & Edlund, T. Molec. Endocr. 5, 897–904 (1991).
Choi, O.-R. B. & Engel, J. D. Cell 55, 17–26 (1988).
Foley, K. P. & Engel, J. D. Genes Dev. 6, 730–744 (1992).
Kellum, R. & Schedl, P. Cell 64, 941–950 (1991).
Roseman, R. R., Pirotta, V. & Geyer, P. K. EMBO J. 12, 435–442 (1993).
Brown, C. J. et al. Cell 71, 527–542 (1992).
Brockdorff, N. et al. Cell 71, 515–526 (1992).
Hendrich, B. D., Brown, C. J. & Willard, H. F. Hum. molec. Gen. 2, 663–672 (1993).
Kay, G. F. et al. Cell 72, 171–182 (1993).
Wevrick, R., Kerns, J. A. & Francke, U. Hum. molec. Genet. 3, 1877–1882 (1994).
Pfeifer, K. & Tilghman, S. M. Genes Dev. 8, 1867–1874 (1994).
Zachar, V., Thomas, R. A. & Goustin, A. S. Nucleic Acids Res. 21, 2017–2018 (1993).
Zemel, S., Bartolomei, M. S. & Tilghman, S. M. Nature Genet. 2, 61–65 (1992).
Soriano, P., Montgomery, C., Geske, R. & Bradley, A. Cell 64, 693–702 (1991).
Robertson, E., Bradley, A., Kuehn, M. & Evans, M. Nature 323, 445–448 (1986).
Southern, E. M. J. molec. Biol. 98, 503–517 (1975).
Feinberg, A. P. & Vogelstein, B. Analyt. Biochem. 132, 6–13 (1983).
Thomas, P. S. Proc. natn. Acad. Sci. U.S.A. 77, 5201–5205 (1980).
Auffray, C. & Rougeon, F. Eur. J. Biochem. 107, 303–314 (1980).
Dudov, K. P. & Perry, R. P. Cell 37, 457–468 (1984).
Chirgwin, J. M., Przybyla, R. J., Macdonald, R. J. & Rutter, W. J. Biochemistry 18, 5294–5299 (1979).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Leighton, P., Ingram, R., Eggenschwiler, J. et al. Disruption of imprinting caused by deletion of the H19 gene region in mice. Nature 375, 34–39 (1995). https://doi.org/10.1038/375034a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/375034a0
This article is cited by
-
Association Between rs217727 and rs2839698 H19 Polymorphisms and Obesity
Biochemical Genetics (2023)
-
Possible transfer of lncRNA H19-derived miRNA miR-675-3p to adjacent H19-non-expressing trophoblast cells in near-term mouse placenta
Histochemistry and Cell Biology (2023)
-
The admixed brushtail possum genome reveals invasion history in New Zealand and novel imprinted genes
Nature Communications (2023)
-
Analysis of long intergenic non-coding RNAs transcriptomic profiling in skeletal muscle growth during porcine embryonic development
Scientific Reports (2021)
-
Reverse-genetics studies of lncRNAs—what we have learnt and paths forward
Genome Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.