Abstract
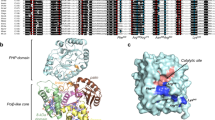
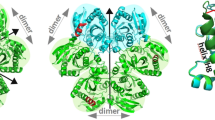
The three-domain structure of porphobilinogen deaminase, a key enzyme in the biosynthetic pathway of tetrapyrroles, has been defined by X-ray analysis at 1.9 Å resolution. Two of the domains structurally resemble the transferrins and periplasmic binding proteins. The dipyrromethane cofactor is covalently linked to domain 3 but is bound by extensive salt-bridges and hydrogen-bonds within the cleft between domains 1 and 2, at a position corresponding to the binding sites for small-molecule ligands in the analogous proteins. The X-ray structure and results from site-directed mutagenesis provide evidence for a single catalytic site. Interdomain flexibility may aid elongation of the polypyrrole product in the active-site cleft of the enzyme.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Jordan, P. M. in New Comprehensive Biochemistry Vol. 19 (eds Neuberger, A. & Van Deenen, L. L. M.) 1–66 (Elsevier, Amsterdam, 1991).
Thomas, S. D. & Jordan, P. M. Nucleic Acids Res. 14, 6215–6226 (1986).
Petricek, M., Rutberg, L., Schroder, I. & Hederstedt, L. J. Bact 172, 2250–2258 (1990).
Sharif, A. L., Smith, A. G. & Abell, C. Eur. J. Biochem. 184, 353–359 (1989).
Stubnicer, A. C., Picat, C. & Grandchamp, B. Nucleic Acids Res. 16, 3102 (1988).
Beaumont, C. et al. J. biol. Chem. 264, 14829–14834 (1989).
Raich, N. et al. Nucleic Acids Res. 14, 5955–5968 (1986).
Kappas, A., Sassa, S., Galbraith, R. A. & Nordmann, Y. in The Metabolic Basis of Inherited Disease (eds Scriver, C. R., Beaudet, A., Sly, W. S. & Valle, D.) 1305–1365 (McGraw-Hill, New York, 1989).
Jordan, P. M. & Warren, M. J. FEBS Lett. 225, 87–92 (1987).
Hart, G. J., Miller, A. D., Leeper, F. J. & Battersby, A. R. J. chem. Soc. chem. Commun. 23, 1762–1765 (1987).
Warren, M. J. & Jordan, P. M. Biochemistry 27, 9020–9029 (1988).
Jones, C., Jordan, P. M. & Akhtar, M. A. J. chem. Soc. Perkin Trans. 1, 2625–2633 (1984).
Schauder, J.-R., Jendrezejewski, S., Abell, A., Hart, G. J. & Battersby, A. R. J. chem. soc. Chem. Commun. 6, 436–438 (1987).
Richardson, J. S. Adv. Protein Chem. 34, 167–339 (1981).
Sutcliffe, M. J., Haneef, I., Carney, D. & Blundell, T. L. Protein Engng 1, 377–384 (1987).
Lander, M. et al. Biochem. J. 275, 447–452 (1991).
Jordan, P. M. & Woodcock, S. C. Biochem. J. 280, 445–449 (1991).
Hadener, A., Alefounder, P. R., Hart, G. J. & Battersby, A. R. Biochem. J. 271, 487–491 (1990).
Louie, G. V. & Brayer, G. D. J. molec. Biol. 214, 527–542 (1990).
Scott, A. I. et al. FEBS Lett. 242, 319–324 (1989).
Jordan, P. M. et al. J. molec. Biol. 224, 269–271 (1992).
Spurlino, J., Lu, G.-Y. & Quiocho, F. A. J. biol. Chem. 266, 5202–5219 (1991).
Pflugrath, J. W. & Quiocho, F. A. J. molec. Biol. 200, 163–180 (1988).
Luecke, H. & Quiocho, F. A. Nature 347, 402–406 (1990).
Baker, E. N., Rumball, S. V. & Anderson, B. F. Trends biochem. Sci. 12, 350–353 (1987).
Sarra, R., Garratt, R., Gorinsky, B., Jhoti, H. & Lindley, P. Acta Crystallogr. B46, 763–771 (1990).
Anderson, B. F. et al. Nature 344, 784–787 (1990).
Mauzerall, D. J. Am. chem. Soc. 82, 2605–2609 (1960).
Delfau, M. H. et al. J. din. Invest. 86, 1511–1516 (1990).
Mgone, C. S., Lanyon, W. G., Moore, M. R. & Connor, J. M. Hum. Genet. (in the press).
Wang, B. C. Meth. Enzym. 115, 90–111 (1985).
Jones, T. A. J. appl. Crystallogr. 11, 268–272 (1978).
Haneef, I., Moss, D. S., Stanford, M. J. & Borkakoti, N. Acta Crystallogr. A41, 426–433 (1985).
Brunger, A. T., Kuriyan, J. & Karplus, M. Science 235, 458–460 (1987).
Ramakrishnan, C. & Ramachandran, G. N. Biophys. J. 5, 909–933 (1965).
Bernstein, F. C. et al. J. molec. Biol. 112, 533–542 (1977).
Evans, S. V. J. molec. Graphics (in the press).
Connolly, M. L. Science 221, 709–713 (1983).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Louie, G., Brownlie, P., Lambert, R. et al. Structure of porphobilinogen deaminase reveals a flexible multidomain polymerase with a single catalytic site. Nature 359, 33–39 (1992). https://doi.org/10.1038/359033a0
Issue Date:
DOI: https://doi.org/10.1038/359033a0
This article is cited by
-
Propionibacterium spp.—source of propionic acid, vitamin B12, and other metabolites important for the industry
Applied Microbiology and Biotechnology (2018)
-
Schiff base bis(5-nitrosalycilaldehyde)ethylenediamine as colorimetric sensor for fluoride
Research on Chemical Intermediates (2015)
-
Recent advances in the biosynthesis of modified tetrapyrroles: the discovery of an alternative pathway for the formation of heme and heme d 1
Cellular and Molecular Life Sciences (2014)
-
A porphobilinogen deaminase (PBGD) Ran-binding protein interaction is implicated in nuclear trafficking of PBGD in differentiating glioma cells
Oncogene (2003)
-
Acute Intermittent Porphyria: Expression of Mutant and Wild-Type Porphobilinogen Deaminase in COS-1 Cells
Molecular Medicine (2000)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.