Abstract
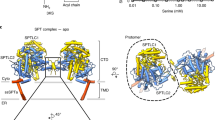
The yeast phosphatidylinositol-transfer protein (Sec14) catalyses exchange of phosphatidylinositol and phosphatidylcholine between membrane bilayers in vitro1,2. In vivo, Sec14 activity is essential for vesicle budding from the Golgi complex3. Here we report a three-dimensional structure for Sec14 at 2.5 Å resolution. Sec14 consists of twelve α-helices, six β-strands, eight 310-helices and has two distinct domains. The carboxy-terminal domain forms a hydrophobic pocket which, in the crystal ructure, is occupied by two molecules of n-octyl-β-D-glucopyranoside and represents the phospholipid-binding domain. This pocket is reinforced by a string motif whose disruption in a sec14 temperature-sensitive mutant results in destabilization of the phospholipid-binding domain. Finally, we have identified an unusual surface helix that may play a critical role in driving Sec14-mediated phospholipid exchange. From this structure, we derive the first molecular clues into how a phosphatidylinositol-transfer protein functions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Bankaitis, V. A., Aitken, J. R., Cleves, A. E. & Dowhan, W. An essential role for a phospholipid transfer protein in yeast Golgi function. Nature 347, 561–562 (1990).
Cleves, A. E., McGee, T. P. & Bankaitis, V. A. Phospholipid transfer proteins: a biological debut. Trends Cell Biol. 1, 30–34 (1991).
Kearns, B. G.et al. An essential role for diacylglycerol in protein transport from the yeast Golgi complex. Nature 387, 101–105 (1997).
Salama, S. R., Cleves, A. E., Malehorn, D. E., Whitters, E. A. & Bankaitis, V. A. Cloning and characterization of Kluyveromyces lactis SEC14, a gene whose product stimulates Golgi secretory function in Saccharomyces cerevisiae. J. Bact. 172, 4510–4521 (1990).
Sato, Y.et al. Primary structure of alpha-tocopherol transfer protein from rat liver. Homology with cellular retinaldehyde protein. J. Biol. Chem. 268, 17705–17710 (1993).
Gu, M., Warshawsky, I. & Majerus, P. W. Cloning and expression of megakaryocyte protein-tyrosine-phosphatase with sequence homology to retinaldehyde binding protein and yeast SEC14p. Proc. Natl Acad. Sci. USA 88, 2980–2984 (1992).
Chinen, K., Takahashi, E. & Nakamura, Y. Isolation and mapping of a human gene (SEC14L), partially homologous to yeast SEC14, that contains a variable number of tandem repeat (VNTR) sites in its 3′ untranslated region. Cytogenet. Cell Genet. 73, 218–223 (1996).
Cleves, A. E.et al. Mutations in the DP-choline pathway for phospholipid biosynthesis bypass the requirement for an essential phospholipid transfer protein. Cell 64, 789–800 (1991).
McGee, T. P., Skinner, H. B., Whitters, E. A., Henry, S. A. & Bankaitis, V. A. Aphosphatidylinositol transfer protein controls the phosphatidylcholine content of yeast Golgi membranes. J. Cell Biol. 124, 273–287 (1994).
Sha, B. D., Phillips, S. E., Bankaitis, V. A. & Luo, M. Crystallization and preliminary X-ray diffraciton studies of the Saccharomyces cerevisiae phospholipid-transfer protein Sec14p. Acta Crystallogr. D 53, 784–786 (1997).
Holm, L. & Sander, C. Searching protein structure databases has come of age. Proteins 19, 165–173 (1994).
Skinner, H. B., Alb, J. B., Whitters, E. A., Helmkamp, G. M. & Bankaitis, V. A. Phospholipid transfer activity is relevant to but not sufficient for the essential function of the yeast SEC14 gene product. EMBO J. 12, 4775–4789 (1993).
Novick, P., Field, C. & Schekman, R. Identification of 23 complementation groups required for post-translational events in the yeast secretory pathway. Cell 21, 205–215 (1980).
Bankaitis, V. A., Malehorn, D. E., Emr, S. D. & Greene, R. The Saccharomyces cerevisiae SEC14 gene encodes a cytosolic factor that is required for transport of secretory proteins from the yeast Golgi complex. J. Cell Biol. 108, 1271–1281 (1989).
Cleves, A. E., Novick, P. J. & Bankaitis, V. A. Mutations in the SAC1 gene suppress defects in yeast Golgi and yeast actin function. J. Cell Biol. 109, 2939–2950 (1989).
Bankaitis, V. A., Fry, M. R., Cartee, R. T. & Kagiwada, S. Phospholipid Transfer Proteins: Emerging Roles in Vesicle Trafficking, Signal Transduciton, and Metabolic Regulation (Lanes, Austin, TX, 1996).
Dickeson, S. K.et al. Isolation and sequence of cDNA clones encoding rat phosphatidylinositol transfer protein. J. Biol. Chem. 264, 16557–16564 (1989).
Alb, J. G. J, Melissa, A. K. & Bankaitis, V. A. Phospholipid metabolism and membrane dynamics. Curr. Opin. Cell Biol. 8, 534–541 (1996).
Minor, W. XdisplayF Program (Purdue Univ., West Lafayette, 1993).
Otwinowski, Z. Proc. CCP4 Study Weekend: Data Collection and Processing (compiled by Sawyer, L., Isaacs, N. & Bailey, S.) 56–62 (SER Daresbury Laboratory, England, 1993).
McRee, D. E. Avisual protein crystallographic sofware system for X11/XView. J. Mol. Graphics. 10, 44–46 (1992).
CCP4, The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Matthews, B. W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–497 (1968).
Jones, T. A., Zhou, J. Y., Cowan, S. W. & Kjeldgard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brunger, A. T. X-PLOR, version 3.85. A system for X-ray diffraction: solvation properties of penicillopepsion and neuraminidase crystal structure. J. Mol. Biol. 243, 100–115 (1994).
Jiang, J. S. & Brunger, A. T. Protein hydration observed by X-ray diffraction: solvation properties of penicillopepsion and neuraminidase crystal structure. J. Mol. Biol. 243, 100–115 (1994).
Carson, M. Ribbons 2.0. J. Appl. Crystallogr. 24, 958–961 (1991).
Nichoos, A. GRASP: Graphical Representation & Analysis Surface Properties (Columbia University, New York, 1992).
Huang, C. & Mason, J. T. Geometric packing constraints in egg phosphatidylcholine vesicles. Proc. Natl Acad. Sci. USA 75, 308–310 (1978).
Vaz, W. L. C., Goodsaid-Zalduondo, F. & Jacobson, K. A. Lateral diffusion of lipids and proteins in bilayer membranes. FEBS Lett. 174, 199–207 (1984).
Acknowledgements
We thank J. Collawn for helpful discussions and critical reading of the manuscript and D. Malehorn for help with site-directed mutagenesis. We thank J. Tsao and Y. Luo for their help in data collection. This work was supported by a grant from the NIH to V.A.B. and NASA to M.L. The X-ray crystallographic coordinates have been deposited in the BNL Protein Data Bank (accession number 1AUA).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Sha, B., Phillips, S., Bankaitis, V. et al. Crystal structure of the Saccharomyces cerevisiae phosphatidylinositol- transfer protein. Nature 391, 506–510 (1998). https://doi.org/10.1038/35179
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35179
This article is cited by
-
Phylogenetic analysis of plant multi-domain SEC14-like phosphatidylinositol transfer proteins and structure–function properties of PATELLIN2
Plant Molecular Biology (2020)
-
Lipid transfer proteins: the lipid commute via shuttles, bridges and tubes
Nature Reviews Molecular Cell Biology (2019)
-
Multiplexed precision genome editing with trackable genomic barcodes in yeast
Nature Biotechnology (2018)
-
Self-assembled α-Tocopherol Transfer Protein Nanoparticles Promote Vitamin E Delivery Across an Endothelial Barrier
Scientific Reports (2017)
-
Identifying patellin-like genes in Glycine max and elucidating their response to phosphorus starvation
Acta Physiologiae Plantarum (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.