Abstract
Most cells contain the same set of genes and yet they are extremely diverse in appearance and functions. It is the selective expression and repression of genes that determines the specific properties of individual cells. Nevertheless, even when fully differentiated, any cell can potentially be reprogrammed back to totipotency, which in turn results in re-differentiation of the full repertoire of adult cells from a single original cell of any kind. Mechanisms that regulate this exceptional genomic plasticity and the state of totipotency are being unravelled, and will enhance our ability to manipulate stem cells for therapeutic purposes.
Similar content being viewed by others
Main
Development is a remarkably orderly process — it begins with a totipotent zygote and ends with an array of specific, differentiated cell types in adults. About 40,000 genes are needed to build a human being possessing ∼200 histologically distinct cell types, and these categories can be subdivided further into a myriad of specialized cell types. They fulfil precise functions that are as diverse as mounting a defence against diseases, regulating energy input–output and building neural networks, so allowing us to interact with our environment.
Once a cell is fully differentiated, this state is strikingly stable. Regardless of how different a neuron is from a hepatocyte, most cells retain an intact genome with the full complement of genes that are present at the beginning in the zygote. This simple concept of profound significance for development had its origin in the work of Spemann. The distinguishing features of cells arise from an orderly selection of genes that are expressed while the rest are switched off.
The genetic network that controls developmental decisions is beginning to be defined. The ability to acquire and inherit gene-expression patterns efficiently is also crucial to the individual history of cell differentiation. There are potential mechanisms that can allow differentiated cells to perpetuate the 'molecular memory' of the developmental decisions that created it. We know that this occurs without alterations or deletion of any DNA sequences, but rather by epigenetic mechanisms, which propagate appropriate patterns of gene expression (Fig. 1). These mechanisms involve heritable but potentially reversible modifications of DNA, primarily methylation of CpG (cytosine–guanine) dinucleotide1. The binding of specific protein complexes to DNA also occurs to form stable and heritable chromatin structures that ensure efficient silencing of genes2,3 that are no longer required for determination of cell fate, allowing expression of only those genes that define properties of specific, differentiated cell types.
Most cells contain the same set of genes, but their phenotype can vary according to which genes are expressed and repressed. Alterations in gene-expression patterns, without changes in DNA sequences, are referred to as epigenetic mechanisms. Epigenetic mechanisms make it possible to restore pluripotency to a differentiated cell, and a differentiated cell can also undergo transdifferentiation resulting in a pronounced change in its appearance and function. Mammalian genomes contain an additional layer of epigenetic information referred to as parental 'imprints'. These imprints are erased and re-initiated normally in the germ line, and passed on to the offspring in which they survive into adulthood. Parental imprints also regulate gene expression and confer functional differences on parental genomes during development. Parental imprints can undergo changes without affecting the fundamental property of pluripotency.
Although the mechanisms that perpetuate cell memory are naturally robust and reliable, they can be erased under some circumstances. The most pronounced manifestation of this erasure occurs when a differentiated somatic nucleus is transplanted back into an oocyte, which results in the restoration of totipotency4,5,6,7. The reconstituted egg can then progress forward to generate a new organism that is a genetic copy or a clone of the individual nuclear donor. While not an efficient process, it is remarkable that it occurs at all; importantly, however, this establishes the principle that epigenetic states are reversible.
Mammalian genomes have an additional layer of epigenetic information referred to as genomic imprints, so called because they carry a molecular memory of their parental origin that is acquired in the germ line (Fig. 1). All our genomes therefore contain these distinct maternal and paternal 'imprints' that are inherited after fertilization by embryos and endure thereafter into adulthood8,9. These modifications, which are recognized as differential methylation of specific DNA sequences in sperm and oocytes, regulate expression of imprinted genes, which confer functional differences between parental genomes during development. Thus parental genomes exhibit an epigenetic asymmetry at fertilization, which persists throughout life. Furthermore, while the overall epigenetic state of the genome changes markedly during development and differentiation of cells, the parental imprints remain relatively stable.
Reprogramming of genomes during imprinting in the germ line requires a stepwise cycle of erasure and re-initiation of imprints. A less well explored but a highly significant consequence of genomic imprinting is that the oocyte cytoplasmic factors have apparently evolved and acquired complex properties in mammals that are required to enhance and maintain the epigenetic asymmetry between parental genomes in the zygote (refs 8, 10–13, and K. Arney et al. unpublished data). These factors could have important consequences for reprogramming of a somatic nucleus to totipotency when transplanted into the oocyte. One key objective in this field is to gain a detailed knowledge of the mechanisms involved in the erasure of existing epigenetic states and establishment of new modifications for totipotency and during imprinting. These studies will allow us to assess more precisely events associated with reprogramming of somatic nuclei to a pluripotent or a totipotent state. The analysis of epigenetic mechanisms involved is also crucial for our ability to manipulate pluripotent stem cells and for the derivation of a range of differentiated cell types from pluripotent embryonic stem cells.
Epigenetic asymmetry between parental genomes
One of the main consequences of genomic imprinting and epigenetic asymmetry is that, whereas oocytes are potentially totipotent in many organisms, this is not so in mammals. This is because the maternal genome is epigenetically modified in the germ line to contain only the maternal 'imprints', which will normally result in the repression of certain maternally inherited imprinted genes. A paternal genome is essential to 'rescue' the oocyte, as the maternal genes are imprinted reciprocally to paternal imprints14,15. So both parental genomes are needed for normal development — the paternal genome is relatively more important for development of the extraembryonic tissues, such as the trophectoderm, whereas the maternal genome apparently has a greater influence on development of the embryo proper.
So far, about 45 imprinted genes have been identified in mice and humans8,9,16. Imprinted genes may regulate some of the crucial aspects of mammalian physiology associated with reproduction, placentation, energy homeostasis, lactation and behaviour16,17,18. For example, Igf2, which encodes a fetal insulin-like growth factor 2, is repressed in the maternal genome and active only in the paternal genome19. Other genes are repressed in the paternal genome and active in the maternal genome8,9,16 (for full details, see http://www.mgu.har.mrc.ac.uk). Some of the anomalies encountered in cloned embryos suggest disruption of imprinted gene expression20.
Imprinted genes are often organized in clusters, sometimes in the megabase-range chromosomal regions containing key control elements — the differentially methylated regions (DMRs)8,9,16. DMRs are CpG rich and subject to epigenetic modifications. These imprinting control regions are often complex with multiple functions acting to repress genes when methylated, or serving as boundary elements when unmethylated (the boundary element21,22 indirectly affects expression of neighbouring genes). Some DMRs also function as silencer elements when unmethylated23, a function that is apparently abolished when the DMR is methylated. In other instances, a DMR is associated with the expression of an antisense transcript whose expression in turn ensures repression of the upstream gene24. The result, in all cases, is to ensure monoallelic expression of imprinted genes. Such complex organization of imprinted genes means that any disruption of such clusters through chromosomal translocations or epigenetic mutations can cause anomalous expression of many genes within the cluster, and this accounts for some of the diseases associated with growth, neurogenetic disorders and diabetes8,9,16.
Genomic imprinting probably accompanied mammalian evolution25, and the evolution of placentation and viviparity in mammals resulted in significant changes in early development. One striking aspect is the emergence of trophectoderm cells, which are essential for blastocyst implantation and as such are the first differentiated cell type to form during development. Also within the blastocyst are the pluripotent epiblast cells, the precursor of embryonic stem (ES) cells (Fig. 2). Gastrulation commences relatively late after implantation in response to signals that emanate from the trophectoderm and primary endoderm cells. Another feature of mammalian development is that the oocytes are relatively small and lack the cytoplasmic determinants of development commonly encountered in other organisms, including those for the germ cell lineage. As a result, there is relatively early activation of the embryonic genome, which in the mouse occurs at the two-cell stage (Fig. 2).
Development commences with the totipotent zygote following fertilization with reciprocally 'imprinted' parental genomes. Imprinting confers developmental asymmetry on parental genomes, so that both are essential for normal development. The mouse embryonic genome is activated at the two-cell stage, and is followed by development of a blastocyst with an inner cell mass and trophectoderm cells. The epiblast cells within the inner cell mass are pluripotent, and give rise to all the somatic cells in the fetus, as well as germ cells. PGCs retain pluripotency as shown by the ability to generate EG cells, which may lack parental imprints.
The initiation of imprinting is confined to the germ line, first with the erasure of existing imprints in primordial germ cells (PGCs)26,27,28,29, followed by the initiation of a new set of imprints in the male and female germ lines. It should be noted that whereas methylation of DMRs for some genes results in their repression, in other instances (for example, the Igf2r gene), methylation is essential for gene activation8,9,16. In most instances, methylation of DMRs occurs predominantly in the female germ line. Nuclear transplantation studies between developing oocytes have shown that the maternal imprints are acquired at the time when oocytes resume growth from a quiescent state prior to ovulation30,31,32. At this time, the X chromosome also acquires an imprint for the non-random inactivation of the paternal X chromosome in trophectoderm and primary endoderm cells29. The precise mechanism by which de novo methylation of DMRs occurs is not yet known, but it should involve some germline-specific factors acting in conjunction with DNA methyltransferase enzymes.
Genomic reprogramming in the germ line
Two important properties of PGCs are that they retain pluripotency (reviewed by Donovan and Gearhart, pages 92–97), and they are endowed with an exceptional capacity for epigenetic modifications of the genome. A unique feature is their ability to erase parental imprints, which shows that epigenetic modifications associated with imprinting can occur independently of the genomic status concerning pluripotency.
The pluripotent germ line
Elaborate transcriptional regulation is generally associated with the founding of germ cells to prevent them from acquiring a somatic cell fate. In mice, germ cells originate from the proximal epiblast cells of the embryonic day 6.5 (E6.5) egg cylinder. Cells migrate to the posterior proximal region where a founder population of ∼45 PGCs is detected by E7.2 (refs 33, 34). It is the proximal location of the epiblast cells that is critical for germ cell fate, rather than any intrinsic properties of these cells35. The specification of germ cell lineage depends on signals, such as bone morphogenetic protein (BMP)-4 and BMP8b (refs 36, 37), originating from the extraembryonic ectoderm in contact with the proximal epiblast. PGCs can be generated in vitro by combining epiblast with extraembryonic tissues38 and, in principle, it should also be possible to generate PGCs from ES or embryonic germ (EG) cells. Oct4, a gene expressed in all totipotent and pluripotent cells in mammals39,40, has an enhancer that is required for its expression predominantly in germ cells40. There are, however, additional genes involved in the specification of the germ cell fate, and these are also likely to be crucial for pluripotency in general, including pluripotent stem cells (M. Saitou, S. C. Barton and M.A.S., unpublished data). Nevertheless, PGCs cannot participate in early development if re-introduced into blastocysts, unlike ES cells that can differentiate into a wide variety of somatic cells and germ cells. It may be that PGCs do not respond to signalling molecules, or that they are transcriptionally repressed. That PGCs are pluripotent is illustrated by the ability to derive EG cells from them41,42, which share many common properties with ES cells. Precisely how a PGC reverts to a pluripotent EG cell remains unknown.
Epigenetic modifications in germ cells
When PGCs begin migration into the genital ridge at E9.5–10.5, they contain genomic imprints, and one of the two X chromosomes is inactive in female PGCs (ref. 43, and P. Hajkova et al., unpublished data). Pronounced epigenetic modifications commence with the entry of PGCs into the genital ridge (Fig. 3). There is rapid and possibly active genome-wide demethylation in both male and female PGCs, resulting in the erasure of imprints (refs 28, 30, 32, 44, and P. Hajkova et al., unpublished data). This is accompanied by reactivation of the inactive X chromosome in female germ cells43. The same mechanism also erases any aberrant epigenetic modifications, so preventing the inheritance of epimutations, which consequently occurs very rarely45. The precise mechanism responsible for epigenetic erasure and demethylation in PGCs is as yet unclear.
Primordial germ cells in E9.5 embryos have the full complement of parental imprints. Upon the entry of PGCs into the genital ridge at E10.5–E11.5, extensive epigenetic modifications commence, perhaps in response to signalling molecules (purple arrows) from somatic cells. These epigenetic modifications include genome-wide demethylation, reactivation of the inactive X chromosome and erasure of imprints. By E13.5, definitive male and female gonads are formed and the PGCs are devoid of parental imprints. New sex-specific imprints are introduced later during gametogenesis, and are detected in mature sperm and oocytes.
Concerning the timing of epigenetic modifications in PGCs, there are at least two possibilities. First, these epigenetic modifications may be triggered in PGCs by a signal from somatic cells when they enter the genital ridge at E10.5–E11.5, which at this stage of development is undifferentiated and identical in both male and female embryos. Alternatively, the erasure of imprints might occur at a specific time and be regulated by a developmental clock. Whatever determines the timing of these events, PGCs by E13.5 possess an equivalent epigenetic state with erased imprints26,27,44, and male and female gonads become distinguishable with distinct phenotypes. The erasure of imprints is also observed in EG cells28. Here it occurs precociously, being found in EG cells derived from PGCs before their entry into the genital ridge, and could be due to culture of PGCs in vitro. It is important to note that ES cells do not show the same property for the erasure of imprints (see below). When the imprint-free EG cells are introduced into blastocysts to generate chimaeras, they can cause developmental anomalies, such as aberrant growth and skeletal abnormalities28. The initiation of new imprints occurs subsequently during gametogenesis, and primarily during oogenesis.
Other forms of genomic modifications probably occur in PGCs, including restoration of telomeres to their optimum size and DNA repair generally46. In single-cell organisms such as yeast, these functions reside within all cells, but in multicellular organisms, some of these functions occur either optimally or exclusively in the germ line. For example, the levels of telomerase are low in somatic cells, and this contributes to their ageing and senescence. Further work is needed to determine the capacity of the mammalian germ line to deal with DNA damage.
Genomic reprogramming in the zygote
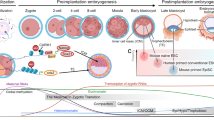
Following imprinting in the germ line, the parental genomes exhibit epigenetic asymmetry at fertilization. These epigenetic differences are both maintained and enhanced in the zygote. During 0–5 hours post fertilization (h.p.f.), parental chromosomes can interact directly with maternally inherited cytoplasmic factors in the oocyte. Thereafter, the pronuclear membrane forms which can regulate access of oocyte cytoplasmic factors to the parental genomes (Fig. 4). During the initial 0–5 h.p.f., the parental genomes exhibit dramatic epigenetic differences. The paternal genome undergoes marked demethylation while the maternal genome, which contains most of the methylation marks associated with imprints, undergoes further de novo methylation11,12,47. Species where imprinting is unknown do not show such differential methylation of parental genomes during early development. Demethylation of the paternal genome may be essential to make it compatible for early activation of the embryonic genome. Alternatively, demethylation of the paternal genome may have accompanied evolution of developmental asymmetry between parental genomes13.
Cytoplasm factors stored in the oocyte commence interactions with parental genomes after fertilization. Between 0 and 5 hours post fertilization (h.p.f.), the parental genomes display marked differences in epigenetic modifications, with the paternal genome undergoing demethylation and the maternal genome showing de novo methylation. This accentuates the epigenetic asymmetry between parental genomes. After the formation of pronuclei at approximately 6 h.p.f., further epigenetic modifications are regulated by the entry of cytoplasmic factors into the nuclei. For example, DNMT1 is excluded from entry into the nuclei. These cytoplasmic factors and others have the potential to modify the epigenetic state of somatic nuclei transplanted into the oocyte. PB, polar body.
The oocyte cytoplasm has often been deployed to discriminate against the paternal genome during the evolution of diverse reproductive strategies and for generating interspecific barriers, which may have been the case during the evolution of genomic imprinting. Disruption of imprinting in interspecific mammalian hybrids of the deermouse, Peromyscus maniculatus, may be due to nuclear–cytoplasmic incompatibility48. Furthermore, certain oocyte cytoplasmic modifiers can induce epigenetic modifications of target loci, subsequently rendering them inactive by DNA methylation49. Such incompatibility may also arise with transplantation of the somatic nucleus into oocytes. The response of a somatic nucleus is likely to be dictated both by its original state, and by how it reacts to the complex oocyte cytoplasmic factors.
There are a number of maternally inherited oocyte cytoplasmic factors with the potential to modify the epigenetic states of parental genomes and of the transplanted somatic nuclei. One such factor is the heterochromatin protein HP1, which may interact differentially with parental genomes, and with somatic nuclei depending on their existing epigenetic state (ref. 10, and K. Arney, unpublished data). HP1 can bind methylated histone H3 (meH3) via a chromodomain, and there is growing evidence indicating that this interaction can lead to de novo DNA methylation. Such interactions may account for the differential methylation of parental genomes in the zygote (K. Arney et al., unpublished data). HP1–meH3 interaction also provides a mechanism for the inheritance of the newly established epigenetic states, a mechanism that is apparently widely conserved in many organisms, including yeast2,3. The histone methyltransferase activity is intrinsic to the SET domain of Su(var)3-9, a Polycomb Group (Pc-G) gene3. This mechanism might also be involved in the initiation of parental imprints during oocyte development.
The oocyte cytoplasm contains several other Pc-G proteins, with the products of Ezh2 and eed seeming particularly crucial for early mammalian development10,50. EZH2 and EED, together with YY1 (mammalian homologue of Drosophila pleiohomiotic), form a complex with histone deacetylases (HDACs) 1 and 2 that has the potential to mediate transcriptional repression51. It is significant that loss of function of any of the three Pc-G genes results in very early embryonic lethality50,52,53. It is particularly striking that EZH2, EED and YY1 are present in the oocyte cytoplasm and translocate into pronuclei after fertilization (ref. 10, and S. Erhardt and K. Arney, unpublished data). EZH2, which also has the conserved SET domain53, may potentially be involved in histone methylation. The role of meH3–HP1 interactions in inducing de novo methylation and as a heritable epigenetic mechanism has the potential to be important in pluripotent stem cells and their subsequent differentiation. For example, the loss of function of Ezh2 is incompatible with the derivation of ES cells53, which is particularly remarkable since ES cells lacking DNA methyltransferases, such as DNMT1, can survive and proliferate extensively54.
DNMT1, which is also present in the oocyte, is known to be excluded from entry into pronuclei, which is why demethylation of the embryonic genome continues throughout pre-implantation development until the blastocyst stage26,47,55. Genome-wide de novo methylation occurs later in early post-implantation embryos. Despite these marked changes in genomic DNA methylation, the parental imprints are preserved, possibly because of the specific properties of the DMRs. The transient entry of DNMT1 into the nuclei at the eight-cell stage seems to help to maintain the imprints55.
Some aspects of transcriptional regulation may be shared between early embryos and PGCs. It is likely, for example, that Ezh2 has a significant role in transcriptional regulation in the germ line. There are only two Pc-G genes known in Caenorhabditis elegans; both are homologues of Ezh and eed, and are critical for transcriptional repression. Loss of function of either of the two genes results in the loss of germ cell lineage in worms. Genome-wide demethylation is also observed in PGCs and in the paternal genome in the zygote11,12,56, although it is important to note that whereas the imprints are erased in PGCs, this is not the case during embryonic development.
Reprogramming somatic nuclei in the oocyte
Transplantation of a somatic nucleus into the oocyte has the potential to restore totipotency to the somatic nucleus57. This transformation probably involves erasure of the existing epigenetic state and a reversion to an embryonic pattern of gene expression. Both the erasure of DNA methylation and the initiation of new epigenetic modifications seem possible, based on studies on the zygote. The imprints have the potential to survive during reprogramming of somatic nuclei to totipotency11,12,47,58, but are occasionally erased as well57.
The interactions between oocyte cytoplasmic factors, such as HP1, and a somatic nucleus will depend in part on the epigenetic state of the donor nucleus when transplanted into oocytes. A donor nucleus is usually transplanted into a non-activated oocyte, which would initially involve direct interactions between the donor chromosomes and the oocyte cytoplasmic factors. Such interactions may result in both de novo methylation and demethylation of specific loci of donor nuclei, which may occur stochastically, resulting subsequently in aberrant patterns of gene expression and failure of development. In one recent study, the methylation status of the donor nucleus at the blastocyst stage was found to be either unchanged or aberrant compared to the control59. Reprogramming of the X chromosome can occur, however, as the inactive X chromosome can be reactivated in the somatic nucleus, although the molecular memory of the inactive X chromosome is retained, resulting in its preferential inactivation in the trophectoderm60. Although imprinted genes may be expected to remain largely unaffected during reprogramming of somatic nuclei to totipotency, there are nevertheless some instances where imprints may be erased, which leads to fetal and placental growth anomalies57. Many other phenotypic anomalies have been noted after live birth (for example, respiratory problems20,61), although it is unclear if this is due to genetic or epigenetic anomalies.
Several additional factors influence survival after birth61, including the genetic background of the donor nucleus. A variety of somatic cell types of different ages have been examined for cloning efficiency, but this still remains at less than 3% (refs 62, 63). Clearly, the oocyte cytoplasmic factors are designed primarily to modify the distinct epigenetic states of parental genomes. A better understanding of both the mechanism of these epigenetic changes and the selection and prior modifications of donor nuclei might improve the outcome.
The oocyte-derived components must also contain other modifiers of chromatin. In amphibians, at least one chromatin-remodelling factor, the ATPase ISWI (a member of the SW12/SNF2 superfamily), erases TATA-binding protein from association with the nuclear matrix of somatic nuclei64. This and other kinds of chromatin-remodelling activities must occur in mice as well. There are also factors, such as OCT4, that could have an essential role in restoring totipotency, whereas other factors might be required for early development until at least the activation of the embryonic genome, and possibly during pre-implantation development.
Another measure of reprogramming of somatic nuclei is the potential to restore the telomere size and to overcome senescence65,66,67,68. Although there is some evidence that telomere length is restored in somatic nuclei after transplantation into oocytes, most DNA mutations in the somatic nucleus cannot be repaired. Although such mutations may be tolerated in a differentiated cell, they can be lethal at any stage of early development. This might also contribute to the low efficiency of development following transplantation of somatic nuclei into oocytes. It is significant that transplantation of nuclei from pluripotent ES cells into oocytes results in higher rates of development to term69. Development of procedures to select and modify the epigenetic status of the donor nucleus might improve the frequency of normal development.
Whereas most attention has focused on how a somatic nucleus may acquire a totipotent state, it has been largely overlooked that the nucleus must also be reprogrammed to undergo rapid transdifferentiation to generate the highly specialized trophectoderm cells by the fourth cleavage division in the mouse. This remarkable example of genomic plasticity is crucial for the subsequent embryonic development. The extraembryonic tissues in mammals are critical as the source of signalling molecules and must function optimally for differentiation of both embryonic somatic cells and for the establishment of the germ line.
Are imprints essential for development?
Because imprints and DNA methylation of the somatic nucleus may affect development, is it possible to obtain normal development if the asymmetry between parental genomes is removed by complete erasure of imprints? Evidence suggests that an imprint-free (and undermethylated) PGC nucleus transplanted to an oocyte is incapable of development to term, even though PGCs are pluripotent30. A larger proportion of these reconstituted zygotes do develop to the blastocyst stage, but the resulting conceptuses fail to progress. In part, this results from an aberrant placental phenotype, which to some extent can be attributed to the loss of function of Mash2, a maternally expressed imprinted gene. Loss of function of other key genes such as Igf2 occurs with the loss of imprints30. There is clear evidence to show that the oocyte cannot initiate imprints if they are erased; imprints can only be initiated in the germ line30.
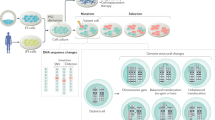
The ability to derive ES cells from blastocysts after nuclear transfer can be used as a measure of nuclear reprogramming (Fig. 5). Individual blastocysts from various somatic cells have been used to derive ES cells, but the success rate is about 4%, compared to an average of 40–50% from normal blastocysts of the 129/Sv strain70. Derivation of ES cells is apparently unaffected by the lack of imprints, as these cells were obtained from blastocysts generated with the imprint-free (and substantially demethylated) PGC nuclei30. Remarkably, the frequency of ES cell derivation in this case was close to 100%, perhaps because PGCs are themselves pluripotent and substantially unmethylated (Y. Kato and M.A.S., unpublished data). It seems that the ability to restore overall pluripotency to somatic nuclei is feasible with aberrant or complete lack of imprints. Additionally, many specific mutations may have no effect on reprogramming of somatic nuclei to pluripotency and the subsequent derivation of ES cells, if they do not affect early development. The consequences of some genetic and epigenetic mutations may become evident only later when these cells are allowed to undergo differentiation towards specific cell types.
When transplanted into an oocyte, a somatic nucleus may respond to the cytoplasmic factors and be reprogrammed back to totipotency. These cytoplasmic factors must be capable of erasing the 'molecular memory' that gives somatic cells their characteristic properties. It would also be necessary for the reprogrammed nucleus to switch off specific genes that are expressed by the somatic nucleus and initiate embryo-specific genes at the two-cell stage in the mouse. The reprogrammed genome generates pluripotent epiblast cells, and undergoes rapid transdifferentiation to generate trophectoderm cells. The extent of reprogramming can be judged by the development of the conceptus in vivo, as well as by the derivation of ES cells from blastocysts. These ES cells can potentially be induced to differentiate to generate the entire repertoire of adult cell types and germ cells.
Reprogramming factors in pluripotent stem cells
The distinguishing feature of pluripotent ES or EG cells is that they exist in a transcriptionally permissive state71. Potentially, this allows indefinite self-renewal without the imposition of restrictions, at least until they are permitted to differentiate into diverse cell types. Then, they undergo progressive restrictions during differentiation, as occurs during development from the totipotent zygote.
Pluripotent stem cells themselves must possess factors with the ability to reprogramme somatic nuclei to pluripotency. As mammalian oocytes are immensely complex, and small in size and numbers, ES and EG cells may provide an alternative system to identify the critical factors and mechanisms necessary for pluripotency using a variety of biochemical, genetic and cell biological approaches. Fusion between pluripotent and somatic cells is the first step towards elucidating the potential of EG and ES cells to modify a somatic nucleus (Fig. 6). This approach, involving fusion between different cell types, has been used previously to examine aspects of gene regulation and genomic plasticity.
A somatic nucleus when fused with an ES or EG cell undergoes extensive reprogramming to pluripotency. The Oct4 gene, which is silent and methylated in somatic cells, is also reactivated as it undergoes demethylation. However, fusion with EG cells, but not ES cells, results in genome-wide demethylation and erasure of imprints from somatic nuclei, a property that is inherited from the precursor primordial germ cells. Pluripotent cells therefore contain factors for extensive modifications of somatic cells, which can confer pluripotency on somatic cells.
Reactivation of the inactive X chromosome occurs when thymocytes are fused with pluripotent embryonal carcinoma cells72. Both ES and EG cells apparently show dominant activities concerning reprogramming of somatic nuclei after cell fusion, presumably in response to trans-acting factors from the pluripotent cells. For example, fusion of a somatic nucleus to an ES cell results in reactivation of Oct4 in the somatic nucleus73 (P. Western and M.A.S., unpublished data). Hence, when thymocytes carrying the Oct4-GFP transgene are fused with ES cells, the reporter expression was observed in hybrid cells within 24–48 h (possibly after 1–2 cell divisions), illustrating that transcriptional reactivation of Oct4 occurs rapidly. As the Oct4 gene is methylated in somatic cells, its expression in ES–somatic cell hybrids is accompanied by demethylation of the promoter region73. Expression of the Oct4 gene is confined to totipotent and pluripotent cells40,74, and the expression of Oct4 from the somatic nucleus is a clear reflection of reprogramming of this somatic nucleus. A similar response is observed when other types of somatic cells are fused with ES cells.
The dominant activity of an EG cell fused with a somatic cell results in the induction of similar epigenetic changes in the somatic cell, together with the restoration of the pluripotent state75. However, EG cells also have the ability to erase parental imprints, a property they inherit from their germline precursor cells. This property is absent in ES cells, oocytes and early embryos. Thus, when fused with a thymocyte, EG cells induce erasure of parental imprints from the somatic nucleus. Furthermore, a silent imprinted allele of the Mest gene is reactivated after demethylation75. Other changes to the somatic nucleus include repression of the thymocyte-specific transcript Thy-1.2 and reactivation of the inactive X chromosome. The hybrid cells also exhibit pluripotency when introduced into blastocysts73,75. The ability of ES and EG cells to reprogramme a somatic nucleus therefore reflects the properties inherited from their precursor cells, the epiblast and germ cells, respectively.
Perspective
Germ cells, stem cells and early embryos all exhibit pluripotency, but each cell type also displays certain unique properties. It is essential to identify the precise nature of the pluripotent state shared by these different cell types, including how it is attained and propagated and which genes confer pluripotency. At the same time, it is imperative to elucidate the mechanisms by which the epigenetic states are changed, so resulting in profound effects on cell potency. Alterations to epigenetic modifications allow a switch in patterns of gene expression, which are central to genomic plasticity and transdifferentiation. While emphasizing the intrinsic nature of the switch, environmental factors also play a fundamental role in vivo, and the impact of these factors on epigenetic modifications must be determined. Many of the advances in understanding the intrinsic mechanisms by which embryonic cell fate is determined during development will prove informative for manipulating cell fates. The immense potential of stem cells will rely on understanding many of the critical mechanisms that regulate the appropriate selection of genes available in all cells.
References
Bird, A. P. & Wolffe, A. P. Methylation-induced repression—belts, braces and chromatin. Cell 99, 451–454 (1999).
Bannister, A. J. et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410, 120–124 (2001).
Jenuwein, T. Re-SET-ting heterochromatin by histone methyltransferases. Trends Cell Biol. 11, 266–273 (2001).
Gurdon, J. The developmental capacity of nuclei taken from intestinal epithelial cells of feeding tadpoles. J. Embryol. Exp. Morphol. 10, 622–640 (1962).
Wilmut, I., Schnieke, A. E., McWhir, J., Kind, A. J. & Campbell, K. H. S. Viable offspring derived from fetal and adult mammalian cells. Nature 385, 810–813 (1997).
Wakayama, T., Zuccotti, M., Johnson, K. R., Perry, A. C. F. & Yanagimachi, R. Full term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature 394, 369–374 (1998).
Kato, Y. et al. Eight calves cloned from somatic cells of a single adult. Science 282, 2095–2098 (1998).
Ferguson-Smith, A. C. & Surani, M. A. Imprinting and the epigenetic asymmetry between parental genomes. Science 293, 1086–1089 (2001).
Reik, W. & Walter, J. Genomic imprinting: parental influence on the genome. Nature Rev. Genet. 2, 21–32 (2001).
Arney, K. L., Erhardt, S., Drewell, R. A. & Surani, M. A. Epigenetic reprogramming of the genome—from the germ line to the embryo and back again. Int. J. Dev. Biol. 45, 533–540 (2001).
Mayer, W., Niveleau, A., Walter, J., Fundele, R. & Haaf, T. Demethylation of the zygotic paternal genome. Nature 403, 501–502 (2000).
Oswald, J. et al. Active demethylation of the paternal genome in the mouse zygote. Curr. Biol. 10, 475–478 (2000).
Reik, W. & Walter, J. Evolution of imprinting mechanisms: the battle of the sexes begins in the zygote. Nature Genet. 27, 255–256 (2001).
Surani, M. A. H., Barton, S. C. & Norris, M. L. Development of reconstituted mouse eggs suggests imprinting of the genome during gametogenesis. Nature 308, 548–550 (1984).
McGrath, J. & Solter, D. Completion of mouse embryogenesis requires both the maternal and paternal genomes. Cell 37, 179–183 (1984).
Tilghman, S. M. The sins of the fathers and mothers: genomic imprinting in mammalian development. Cell 96, 185–193 (1999).
Lefebvre, L. et al. Abnormal maternal behaviour and growth retardation associated with loss of the imprinted gene Mest. Nature Genet. 20, 163–169 (1998).
Li, L. et al. Regulation of maternal behavior and offspring growth by paternally expressed Peg3. Science 284, 330–333 (1999).
DeChiara, T. M., Robertson, E. J. & Efstratidiadis, A. Parental imprinting of the mouse insulin-like growth factor II gene. Cell 64, 849–859 (1991).
Tamashiro, K. L. K., Wakayama, T., Blanchard, R. J., Blanchard, D. C. & Yanagimachi, R. Postnatal growth and behavioural development of mice cloned from adult cumulus cells. Biol. Reprod. 63, 328–334 (2000).
Bell, A. C. & Felsenfeld, G. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature 405, 482–485 (2000).
Hark, A. T. et al. CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus. Nature 405, 486–489 (2000).
Constancia, M. et al. Deletion of a silencer element in Igf2 results in loss of imprinting independent of H19. Nature Genet. 26, 203–206 (2000).
Lyle, R. et al. The imprinted antisense RNA at the Igf2r locus overlaps but does not imprint Mas1. Nature Genet. 25, 19–21 (2000).
Killian, J. K. et al. M6P/IGF2R imprinting evolution in mammals. Mol. Cell 5, 707–716 (2000).
Brandeis, M. et al. The ontogeny of allele-specific methylation associated with imprinted genes in the mouse. EMBO J. 12, 3669–3677 (1993).
Kafri, T. et al. Developmental pattern of gene-specific DNA methylation in the mouse embryo and germ line. Genes Dev. 6, 705–714 (1992).
Tada, T. et al. Epigenotype switching of imprintable loci in embryonic germ cells. Dev. Genes Evol. 207, 551–561 (1998).
Tada, T. et al. Imprint switching for non-random X-chromosome inactivation during mouse oocyte growth. Development 127, 3101–3105 (2000).
Kato, Y. et al. Developmental potential of mouse primordial germ cells. Development 126, 1823–1832 (1999).
Kono, T., Obata, Y., Yoshimzu, T., Nakahara, T. & Carroll, J. Epigenetic modifications during oocyte growth correlates with extended parthenogenetic development in the mouse. Nature Genet. 13, 91–94 (1996).
Obata, Y. et al. Disruption of primary imprinting during oocyte growth leads to the modified expression of imprinted genes during embryogenesis. Development 125, 1553–1560 (1998).
Lawson, K. A. & Hage, W. J. Clonal analysis of the origin of primordial germ cells in the mouse. Ciba Found. Symp. 182, 68–84 (1994).
McLaren, A. Signaling for germ cells. Genes Dev. 13, 373–376 (1999).
Tam, P. P. & Zhou, S. X. The allocation of epiblast cells to ectodermal and germ-line lineages is influenced by the position of the cells in the gastrulating mouse embryo. Dev. Biol. 178, 124–132 (1996).
Lawson, K. A. et al. Bmp4 is required for the generation of primordial germ cells in the mouse embryo. Genes Dev. 13, 424–436 (1999).
Ying, Y., Liu, X. M., Marble, A., Lawson, K. A. & Zhao, G. Q. Requirement of Bmp8b for the generation of primordial germ cells in the mouse. Mol. Endocrinol. 14, 1053–1063 (2000).
Yoshimizu, T., Obinata, M. & Matsui, Y. Stage-specific tissue and cell interactions play key roles in mouse germ cell specification. Development 128, 481–490 (2001).
Nichols, J. et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct-4. Cell 95, 379–391 (1998).
Pesce, M., Gross, M. K. & Scholer, H. R. In line with our ancestors: Oct-4 and the mammalian germ. Bioessays 20, 722–732 (1998).
Matsui, Y., Zsebo, K. & Hogan, B. L. Derivation of pluripotential embryonic stem cells from murine primordial germ cells in culture. Cell 70, 841–847 (1992).
Resnick, J. L., Bixler, L. S., Cheng, L. & Donovan, P. Long-term proliferation of mouse primordial germ cells in culture. Nature 359, 550–551 (1992).
Tam, P. L., Zhou, S. X. & Tan, S.-S. X-chromosome activity of the mouse primordial germ cells revealed by the expression of an X-linked lacZ transgene. Development 120, 2925–2932 (1994).
Mann, J. R. Imprinting in the germ line. Stem Cells 19, 289–294 (2001).
Morgan, H. D., Sutherland, H. G., Martin, D. I. & Whitelaw, E. Epigenetic inheritance at the agouti locus in the mouse. Nature Genet. 23, 314–318 (1999).
Meeker, A. K. & Coffey, D. S. Telomerase: a promising marker of biological immortality of germ, stem, and cancer cells. A review. Biochemistry 62, 1323–1331 (1997).
Monk, M., Boubelik, M. & Lehnert, S. Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ cell lineages during mouse embryo development. Development 99, 371–382 (1987).
Vrana, P. B., Guan, X. J., Ingram, R. S. & Tilghman, S. M. Genomic imprinting is disrupted in interspecific Peromyscus hybrids. Nature Genet. 20, 362–365 (1998).
Allen, N. D., Norris, M. L. & Surani, M. A. Epigenetic control of transgene expression and imprinting by genotype-specific modifiers. Cell 61, 853–861 (1990).
Schumacher, A., Faust, C. & Magnuson, T. Positional cloning of a global regulator of anterior–posterior patterning in mice. Nature 384, 648 (1996).
Satijn, D. P., Hamer, K. M., den Blaauwen, J. & Otte, A. P. The polycomb group protein EED interacts with YY1, and both proteins induce neural tissue in Xenopus embryos. Mol. Cell. Biol. 21, 1360–1369 (2001).
Donohoe, M. E. et al. Targeted disruption of mouse Yin Yang 1 transcription factor results in peri-implantation lethality. Mol. Cell. Biol. 19, 7237–7244 (1999).
O'Carroll, D. et al. The Polycomb group gene Ezh2 is required for early mouse development. Mol. Cell. Biol. 21, 4330–4336 (2001).
Li, E., Beard, C. & Jaenisch, R. Role for DNA methylation in genomic imprinting. Nature 366, 362–365 (1993).
Howell, C. Y. et al. Genomic imprinting disrupted by a maternal effect mutation in the Dnmt1 gene. Cell 104, 829–838 (2001).
Monk, M. & Harper, M. I. Sequential X chromosome inactivation coupled with cellular differentiation in early mouse embryos. Nature 281, 311–313 (1979).
Solter, D. Mammalian cloning: advances and limitations. Nature Rev. Genet. 1, 199–207 (2000).
Olek, A. & Walter, J. The pre-implantation ontogeny of the H19 methylation imprint. Nature Genet. 17, 275–276 (1997).
Kang, Y.-K. et al. Aberrant methylation of donor genome in cloned bovine embryos. Nature Genet. 28, 173–177 (2001).
Eggan, K. et al. X-Chromosome inactivation in cloned mouse embryos. Science 290, 1578–1581 (2000).
Eggan, K. et al. Hybrid vigor, fetal overgrowth, and viability of mice derived by nuclear cloning and tetraploid embryo complementation. Proc. Natl Acad. Sci. USA 98, 6209–6214 (2001).
Wakayama, T. & Yanagimachi, R. Mouse cloning with nucleus donor cells of different age and type. Mol. Reprod. Dev. 58, 376–383 (2001).
Wakayama, T. & Yanagimachi, R. Effect of cytokinesis inhibitors, DMSO and the timing of oocyte activation on mouse cloning using cumulus cell nuclei. Reproduction 122, 49–60 (2001).
Kikyo, N., Wade, P. A., Guschin, D., Ge, H. & Wolffe, A. P. Active remodeling of somatic nuclei in egg cytoplasm by the nucleosomal ATPase ISWI. Science 289, 2360–2362 (2000).
Betts, D. et al. Reprogramming of telomerase activity and rebuilding of telomere length in cloned cattle. Proc. Natl Acad. Sci. USA 98, 1077–1082 (2001).
Lanza, R. P. et al. Extension of cell life-span and telomere length in animals cloned from senescent somatic cells. Science 288, 665–669 (2000).
Shiels, P. G. et al. Analysis of telomere lengths in cloned sheep. Nature 399, 316–317 (1999).
Wakayama, T. et al. Cloning of mice to six generations. Nature 407, 318–319 (2000).
Rideout, W. M. et al. Generation of mice from wild-type and targeted ES cells by nuclear cloning. Nature Genet. 24, 109–110 (2000).
Wakayama, T. et al. Differentiation of embryonic stem cell lines generated from adult somatic cells by nuclear transfer. Science 292, 740–743 (2001).
Smith, A. G. Embryo-derived stem cells: of mice and men. Annu. Rev. Cell Dev. Biol. (in the press).
Takagi, N., Yoshida, M. A., Sugawara, O. & Sasaki, M. Reversal of X-inactivation in female mouse somatic cells hybridized with murine teratocarcinoma stem cells in vitro. Cell 34, 1053–1062 (1983).
Tada, M. Nuclear reprogramming of somatic cells by in vitro hybridisation with ES cells. Curr. Biol. (in the press).
Yeom, Y. I. et al. Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development 122, 881–894 (1996).
Tada, M., Tada, T., Lefebvre, L., Barton, S. C. & Surani, M. A. Embryonic germ cells induce epigenetic reprogramming of somatic nucleus in hybrid cells. EMBO J. 16, 6510–6520 (1997).
Acknowledgements
I thank members of my laboratory for their valuable contributions and critical comments during the course of our studies, Barry Keverne for critical comments on the manuscript and the Wellcome Trust for generously supporting my work. Space constraints has prevented citation of a large number of key papers.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Surani, M. Reprogramming of genome function through epigenetic inheritance. Nature 414, 122–128 (2001). https://doi.org/10.1038/35102186
Issue Date:
DOI: https://doi.org/10.1038/35102186
This article is cited by
-
Exploring the evidence for epigenetic regulation of environmental influences on child health across generations
Communications Biology (2021)
-
Dynamics of DNA Methylation and DNMT Expression During Gametogenesis and Early Development of Scallop Patinopecten yessoensis
Marine Biotechnology (2019)
-
Programming of Essential Hypertension: What Pediatric Cardiologists Need to Know
Pediatric Cardiology (2015)
-
Paternal heterochromatin formation in human embryos is H3K9/HP1 directed and primed by sperm-derived histone modifications
Nature Communications (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.