Abstract
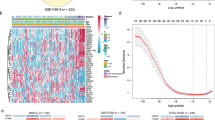
Prostate cancer is the most frequently diagnosed cancer in American men1,2. Screening for prostate-specific antigen (PSA) has led to earlier detection of prostate cancer3, but elevated serum PSA levels may be present in non-malignant conditions such as benign prostatic hyperlasia (BPH). Characterization of gene-expression profiles that molecularly distinguish prostatic neoplasms may identify genes involved in prostate carcinogenesis, elucidate clinical biomarkers, and lead to an improved classification of prostate cancer4,5,6. Using microarrays of complementary DNA, we examined gene-expression profiles of more than 50 normal and neoplastic prostate specimens and three common prostate-cancer cell lines. Signature expression profiles of normal adjacent prostate (NAP), BPH, localized prostate cancer, and metastatic, hormone-refractory prostate cancer were determined. Here we establish many associations between genes and prostate cancer. We assessed two of these genes—hepsin, a transmembrane serine protease, and pim-1, a serine/threonine kinase—at the protein level using tissue microarrays consisting of over 700 clinically stratified prostate-cancer specimens. Expression of hepsin and pim-1 proteins was significantly correlated with measures of clinical outcome. Thus, the integration of cDNA microarray, high-density tissue microarray, and linked clinical and pathology data is a powerful approach to molecular profiling of human cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Abate-Shen, C. & Shen, M. M. Molecular genetics of prostate cancer. Genes Dev. 14, 2410–2434 (2000).
Ruijter, E. et al. Molecular genetics and epidemiology of prostate carcinoma. Endocr. Rev. 20, 22–45 (1999).
Barry, M. J. Prostate-specific-antigen testing for early diagnosis of prostate cancer. N. Engl. J. Med. 344, 1373–1377 (2001).
Liotta, L. & Petricoin, E. Molecular profiling of human cancer. Nature Rev. Genet. 1, 48–56 (2000).
Elek, J., Park, K. H. & Narayanan, R. Microarray-based expression profiling in prostate tumors. In Vivo 14, 173–182 (2000).
Emmert-Buck, M. R. et al. Molecular profiling of clinical tissue specimens: feasibility and applications. Am. J. Pathol. 156, 1109–1115 (2000).
Golub, T. R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
Bittner, M. et al. Molecular classification of cutaneous malignant melanoma by gene expression profiling. Nature 406, 536–540 (2000).
Perou, C. M. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000).
Alizadeh, A. A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
Eisen, M. B., Spellman, P. T., Brown, P. O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863–14868 (1998).
Tomita, K. et al. Cadherin switching in human prostate cancer progression. Cancer Res. 60, 3650–3654 (2000).
Shurbaji, M. S., Kalbfleisch, J. H. & Thurmond, T. S. Immunohistochemical detection of a fatty acid synthase (OA-519) as a predictor of progression of prostate cancer. Hum. Pathol. 27, 917–921 (1996).
Kononen, J. et al. Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nature Med. 4, 844–847 (1998).
Perrone, E. E. et al. Tissue microarray assessment of prostate cancer tumor proliferation in African-American and white men. J. Natl Cancer. Inst. 92, 937–939 (2000).
Tsuji, A. et al. Hepsin, a cell membrane-associated protease. Characterization, tissue distribution, and gene localization. J. Biol. Chem. 266, 16948–16953 (1991).
Pound, C. R. et al. Natural history of progression after PSA elevation following radical prostatectomy. Jama 281, 1591–1597 (1999).
Gleason, D. F. Histologic grading of prostate cancer: a perspective. Hum. Pathol. 23, 273–279 (1992).
Buttyan, R., Sawczuk, I. S., Benson, M. C., Siegal, J. D. & Olsson, C. A. Enhanced expression of the c-myc protooncogene in high-grade human prostate cancers. Prostate 11, 327–337 (1987).
Rubin, M. A. et al. Rapid (‘warm’) autopsy study for procurement of metastatic prostate cancer. Clin. Cancer Res. 6, 1036–10345 (2000).
Acknowledgements
We are indebted to P. Ward and the University of Michigan Microarray Network for support and encouragement in the development of the Pathology Microarray Node. We thank M. Sanda and J. Wei for support of the clinical database; E. Cushenberry for her help with collecting the prostate tissue samples, The University of Michigan Comprehensive Cancer Center Histology and Immunoperoxidase Core; and the instructors of the 1999 Cold Spring Harbor Workshop on DNA Microarrays. This work is supported by developmental grants (A.M.C. and M.A.R.) from the Specialized Program of Research Excellence in Prostate Cancer, National Cancer Institute.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplemental Figure 1
Same as Fig. 1b of the manuscript except for depiction of dendrograms for gene expression.
Supplemental Figure 2
Same as Fig. 1c of the manuscript except for depiction of dendrograms for gene expression.
Supplemental Figure 3
Comparison of normal adjacent prostate tissue (NAP) with the normal prostate tissue reference (CP). The same convention for representing changes in transcript levels was used as in Fig. 1 of the manuscript. The cluster was obtained by selecting for genes with at least a 2.5-fold variation in any two of the samples of each class, namely the normal tissues versus the NAP pool and normal tissue versus the CP pool at a 50% filter. Of the genes analyzed 59 were selected with this criteria. Genes that were found to be up-regulated in the NAP pool in comparison with CP pool, included connective tissue growth factor, EGR-1 (Early Growth Response 1), matrilysin (MMP7), CFLAR/I-FLICE (caspase 8 and FADD-like apoptosis regulator), lumican, serum glucocorticoid regulated kinase, lens epithelium derived growth factor, PAI1 (plasminogen activator inhibitor type I), JUN and FOS B, among others. Vascular endothelial growth factor (VEGF), growth arrest specific (GAS1), cholecystokinin (CCK), amiloride binding protein (ABP1) were among the down-regulated genes in the normal adjacent prostate pool when compared to the commercial pool. The gene expression differences between normal prostate adjacent to PCA (NAP) and normal prostate tissue from individuals without prostate pathology (CP) may be attributable to a "field effect" induced by PCA itself.
Supplementary Figure 4
A focused cluster of PCA-related genes. The same convention for representing changes in transcript levels was used as in Fig. 1 of the manuscript. This cluster of 231 genes was generated by selecting for a 3.5-fold variation in at least 2 of any class, and ratio measurements present in 75% of the samples. Classes included: PCA vs. NAP, MET vs. NAP, PCA vs. CP and MET vs. CP.
Supplemental Figure 5
Gene selection based on computed t-statistics for each gene. Two groups were used in the analysis: PCA/MET and benign (NAP/BPH). a, Analysis of NAP pool data. b, Analysis of CP pool data. Selected genes are named and 200 genes for each data set are shown. Refer to manuscript text for formal statistical method. Gene selection based on each method is included as a supplementary text file. Selected gene names or symbols (as specified by Human genome organization (HUGO) gene nomenclature) are shown.
Supplemental Figure 6
Functional clusters of select genes differentially expressed in prostate cancer. Gene names or symbols (as specified by Human genome organization (HUGO) gene nomenclature) are shown. The same convention for representing changes in transcript levels was used as in Fig.1. The sample order from Fig.1 was preserved for clarity.
Supplemental Figure 7
"Executive Summary." Representative genes differentially expressed in PCA identified by DNA microarray analysis. Genes are grouped functionally and arrows represent up- or down- regulation in metastatic hormone-refractory PCA (MET) and/or localized PCA (PCA) relative to normal prostate epithelium. See Fig.2 of the manuscript for gene expression levels. The following paragraphs refer to descriptions of and hypotheses generated from our gene expression profiling of PCA. Gene names or symbols (as specified by Human genome organization (HUGO) gene nomenclature) are shown.
Supplemental Figure 8
Additional prostate tissue specimens profiled against a commercial prostate reference pool (CPP). The same convention for representing changes in transcript level was used as in Figure 1 of the manuscript. A total of 53 prostate specimens were profiled against the commercial pool. They include, 4 normal adjacent prostate tissue (NAP), 14 benign prostatic hyperplasia (BPH), 1 prostatitis, 14 localized prostate cancer (PCA) and 20 hormone refractory metastatic PCA (MET). Prior to hierarchial average-linkage clustering, the data was filtered for at least 3-fold change in Cy5/Cy3 ratios and measurements present in 75% of the samples. By this method 1325 genes were selected. This figure expands on Figure 1c with an additional 40 samples, which include all from Figure 1b, and also includes 28 additional prostate specimens.
Rights and permissions
About this article
Cite this article
Dhanasekaran, S., Barrette, T., Ghosh, D. et al. Delineation of prognostic biomarkers in prostate cancer. Nature 412, 822–826 (2001). https://doi.org/10.1038/35090585
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35090585
This article is cited by
-
PIM1 drives lipid droplet accumulation to promote proliferation and survival in prostate cancer
Oncogene (2024)
-
Bioinformatics in urology — molecular characterization of pathophysiology and response to treatment
Nature Reviews Urology (2024)
-
Liquid-liquid phase separation throws novel insights into treatment strategies for skin cutaneous melanoma
BMC Cancer (2023)
-
Accurate prognosis for localized prostate cancer through coherent voting networks with multi-omic and clinical data
Scientific Reports (2023)
-
METTL3 regulatory TROAP can regulate the progression of non-small cell lung cancer through PI3K/AKT and EMT signaling pathway
Medical Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.