Abstract
In response to DNA damage and blocks to replication, eukaryotes activate the checkpoint pathways that prevent genomic instability and cancer by coordinating cell cycle progression with DNA repair1,2,3,4,5. In budding yeast, the checkpoint response requires the Mec1-dependent activation of the Rad53 protein kinase3,4,6. Active Rad53 slows DNA synthesis when DNA is damaged7 and prevents firing of late origins of replication8,9. Further, rad53 mutants are unable to recover from a replication block10. Mec1 and Rad53 also modulate the phosphorylation state of different DNA replication and repair enzymes6,11,12,13. Little is known of the mechanisms by which checkpoint pathways interact with the replication apparatus when DNA is damaged or replication blocked. We used the two-dimensional gel technique14 to examine replication intermediates in response to hydroxyurea-induced replication blocks. Here we show that hydroxyurea-treated rad53 mutants accumulate unusual DNA structures at replication forks. The persistence of these abnormal molecules during recovery from the hydroxyurea block correlates with the inability to dephosphorylate Rad53. Further, Rad53 is required to properly maintain stable replication forks during the block. We propose that Rad53 prevents collapse of the fork when replication pauses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Elledge, S. J. Cell cycle checkpoints: preventing an identity crisis. Science 274, 1664–1672 (1996).
Weinert, T. DNA damage checkpoints update: getting molecular. Curr. Opin. Genet. Dev. 8, 185–193 (1998).
Lowndes, N. F. & Murguia, J. R. Sensing and responding to DNA damage. Curr. Opin. Genet. Dev. 10, 17–25 (2000).
Foiani, M. et al. DNA damage checkpoints and DNA replication controls in Saccharomyces cerevisiae. Mutat. Res. 21, 286–294 (2000).
Weinert, T. Yeast checkpoints controls and relevance to cancer. Cancer Surv. 29, 109–132 (1997).
Pellicioli, A. et al. Activation of Rad53 kinase in response to DNA damage and its effect in modulating phosphorylation of the lagging strand DNA polymerase. EMBO J. 18, 6561–6572 (1999).
Paulovich, A. G. & Hartwell, L. H. A checkpoint regulates the rate of progression through S-phase in S. cerevisiae in response to DNA damage. Cell 82, 841–847 (1995).
Santocanale, C. & Diffley, J. F. X. A Mec1- and Rad53-dependent checkpoint controls late-firing origins of DNA replication. Nature 395, 615–618 (1998).
Shirahige, K. et al. Regulation of DNA-replication origins during cell-cycle progression. Nature 395, 618–621 (1998).
Desany, B. A., Alcasabas, A. A., Bachant, J. B. & Elledge, S. J. Recovery from DNA replicational stress is the essential function of the S-phase checkpoint pathway. Genes Dev. 12, 2956–2970 (1998).
Brush, G., Morrow, D. M., Heiter, P. & Kelly, T. J. The ATM homolog MEC1 is required for phosphorylation of replication protein A in yeast. Proc. Natl Acad. Sci. USA 93, 15075–15080 (1996).
Liberi, G. et al. Srs2 DNA helicase is involved in checkpoint response and its regulation requires a functional Mec1-dependent pathway and CDK1 activity. EMBO J. 19, 5027–5038 (2000).
Bashkirov, V. I. et al. DNA repair protein Rad55 is a terminal substrate of the DNA damage checkpoints. Mol. Cell. Biol. 20, 4393–4404 (2000).
Brewer, B. J. & Fangman, W. L. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell 51, 463–471 (1987).
Zhao, X., Muller, E. G. & Rothstein, R. A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol. Cell 2, 329–340 (1998).
Newlon, C. S. et al. Analysis of replication origin function on chromosome III of Saccharomyces cerevisiae. Cold Spring Harb. Symp. Quant. Biol. 58, 415–423 (1993).
Friedman, K. L. & Brewer, B. J. Analysis of replication intermediates by two-dimensional agarose gel electrophoresis. Methods Enzymol. 262, 613–627 (1995).
Lindsay, H. D. et al. S-phase-specific activation of Cds1 kinase defines a subpathway of the checkpoint response in Schizosaccharomyces pombe. Genes Dev. 12, 382–395 (1998).
Edwards, R. J., Bentley, N. J. & Carr, A. M. A Rad3–Rad26 complex response to DNA damage independently of other checkpoint proteins. Nature Cell Biol. 1, 393–398 (1999).
Dimitrova, D. S. & Gilbert, D. M. Temporally coordinated assembly and disassembly of replication factories in the absence of DNA synthesis. Nature Cell Biol. 2, 686–694 (2000).
Seigneur, M., Bidnenko, V., Ehrlich, S. D. & Michel, B. RuvAB acts at arrested replication forks. Cell 95, 419–430 (1998).
Rothstein, R., Michel, B. & Gangloff, S. Replication fork pausing and recombination or “gimme a break”. Genes Dev. 14, 1–10 (2000).
Martin-Parras, L., Hernandez, P., Martinez-Robles, M. L. & Schwartzman, J. B. Initiation of DNA replication in ColE1 plasmids containing multiple potential origins of replication. J. Biol. Chem. 267, 22496–22505 (1992).
Kalejta, R. F. & Hamlin, J. L. Composite patterns in neutral/neutral two-dimensional gels demonstrate inefficient replication origin usage. Mol. Cell. Biol. 16, 4915–4922 (1996).
Deshpande, A. M. & Newlon, C. S. DNA replication fork pause sites dependent on transcription. Science 272, 1030–1033 (1996).
Ivessa, A. S., Zhou, J. Q. & Zakian, V. A. The Saccharomyces Pif1p DNA helicase and the highly related Rrm3p have opposite effects on replication fork progression in ribosomal DNA. Cell 100, 479–489 (2000).
Thomas, B. J. & Rothstein, R. Elevated recombination rates in transcriptionally active DNA. Cell 56, 619–629 (1989).
Marini, F. et al. Role for DNA primase in coupling DNA replication to DNA damage response. EMBO J. 16, 639–650 (1997).
Pellicioli, A., Lee, S. E., Lucca, C., Foiani, M. & Haber, J. E. Regulation of Saccharomyces Rad53 checkpoint kinase during adaptation from DNA damage-induced G2/M arrest. Mol. Cell 7, 293–300 (2001).
Acknowledgements
We thank S. Alberti, L. Fabiani, A. Gambetta, S. Pintus and J. Theis for technical advice and support. We also thank A. Carr, J. Diffley, J. Haber, R. Rothstein, J. Sogo and all the members of our laboratory for helpful discussions. This work was supported by Associazione Italiana per la Ricerca sul Cancro and partially by grants from Telethon–Italy, Cofinanziamento MURST–Università di Milano, MURST (5%) Biomolecole per la Salute Umana, and CNR Target Project on Biotechnology, by a EU TMR contract, and by a NIH grant to C.S.N.
Author information
Authors and Affiliations
Corresponding author
Supplementary information

Addentum to Figure 1
(JPG 27.8 KB)
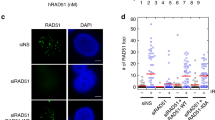
Protein extracts prepared from cells taken at the indicated time points were analysed by western blotting using anti-Rad53 antibodies. The DNA content of wt and rad53 cells was analysed at the indicated time points by FACS analysis.

Addentum to Figure 4
(JPG 10.5 KB)
Protein extracts prepared from cells taken at the indicated time points were analysed by western blotting using anti-DNA polymerase alpha B subunit antibodies. The appearence of the hyperphosphorylated form of the B subunit indicates that the checkpoint has been inactivated (for further details see ref.6). The DNA content of wt and rad53 cells was analysed at the indicated time points by FACS analysis.
Rights and permissions
About this article
Cite this article
Lopes, M., Cotta-Ramusino, C., Pellicioli, A. et al. The DNA replication checkpoint response stabilizes stalled replication forks. Nature 412, 557–561 (2001). https://doi.org/10.1038/35087613
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/35087613
This article is cited by
-
Microbial inoculums improve growth and health of Heteropneustes fossilis via biofloc-driven aquaculture
Microbial Cell Factories (2023)
-
ATR kinase supports normal proliferation in the early S phase by preventing replication resource exhaustion
Nature Communications (2023)
-
Cell cycle control in cancer
Nature Reviews Molecular Cell Biology (2022)
-
Yeast Stn1 promotes MCM to circumvent Rad53 control of the S phase checkpoint
Current Genetics (2022)
-
The yeast Dbf4 Zn2+ finger domain suppresses single-stranded DNA at replication forks initiated from a subset of origins
Current Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.