Key Points
-
Genomic imprinting leads to allele-specific expression depending on the parent of origin of the allele.
-
The most consistent difference between the alleles of an imprinted gene is in DNA methylation. But imprinted genes are also characterized by differences in chromatin conformation, histone modification, replication timing and recombination rate. The primary imprint causing these differences is unknown.
-
The life cycle of imprints involves erasure during early germ cell development, establishment later in germ cell development or after fertilization, and maintenance during embryonic development, and begins again with erasure in the germ cells of the embryo.
-
Genomic imprinting leads to imprinted gene expression. The mechanisms underlying this reading of the imprint can involve different aspects of gene expression: promoter methylation, the regulation of antisense transcripts, boundary elements, and silencers.
-
Imprinted genes are often found in clusters. These can be regulated by imprinting centres.
-
Dysregulation of imprinting has been found in many diseases: growth and behavioural defects (Beckwith–Wiedemann syndrome, Prader–Willi syndrome and Angelman syndrome) and cancer (Wilms tumour).
-
Imprinted genes are implicated in fetal–maternal physiology, and one theory for the evolution of imprinting (the conflict theory) is that it reflects the competing interests of the maternal and paternal genomes in the developing embryo.
Abstract
Genomic imprinting affects several dozen mammalian genes and results in the expression of those genes from only one of the two parental chromosomes. This is brought about by epigenetic instructions — imprints — that are laid down in the parental germ cells. Imprinting is a particularly important genetic mechanism in mammals, and is thought to influence the transfer of nutrients to the fetus and the newborn from the mother. Consistent with this view is the fact that imprinted genes tend to affect growth in the womb and behaviour after birth. Aberrant imprinting disturbs development and is the cause of various disease syndromes. The study of imprinting also provides new insights into epigenetic gene modification during development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Surani, M. A. H., Barton, S. C. & Norris, M. L. Development of reconstituted mouse eggs suggests imprinting of the genome during gametogenesis. Nature 308, 548–550 (1984).
McGrath, J. & Solter, D. Completion of mouse embryogenesis requires both the maternal and paternal genomes. Cell 37, 179–183 (1984). References 1 and 2 are key papers that uncovered the existence of genomic imprinting. Nuclear transplantation in mouse embryos showed that embryos with only maternal or paternal chromosomes could not develop normally, despite being diploid.
Mann, J. R. & Lovell-Badge, R. H. Inviability of parthenogenones is determined by pronuclei, not egg cytoplasm. Nature 310, 66–67 (1984).
Lyon, M. F. & Glenister, P. H. Factors affecting the observed number of young resulting from adjacent-2 disjunction in mice carrying a translocation . Genet. Res. 29, 83–92 (1977).
Searle, A. G. & Beechey, C. V. Complementation studies with mouse translocations. Cytogenet. Cell Genet. 20, 282–303 (1978).
Cattanach, B. M. & Kirk, M. Differential activity of maternally and paternally derived regions in mice. Nature 315, 496–498 (1985). This paper showed that specific chromosome regions could function differently depending on their inheritance from mother or father. It established marked opposing effects of the parental genomes on fetal growth and postnatal behaviour.
DeChiara, T. M., Robertson, E. J. & Efstratidiadis, A. Parental imprinting of the mouse insulin-like growth factor II gene. Cell 64, 849– 859 (1991).
Barlow, D. P., Stoeger, R., Herrmann, B. G., Saito, K. & Schweifer, N. The mouse insulin-like growth factor type-2 receptor is imprinted and closely linked to the Tme locus . Nature 349, 84–87 (1991).
Bartolomei, M. S., Zemel, S. & Tilghman, S. M. Parental imprinting of the H19 gene. Nature 351, 153–155 ( 1991).References 7 – 9 describe the first paternally expressed ( Igf2 ) and maternally expressed ( Igf2r, H19 ) imprinted genes. All three genes are involved in fetal growth control.
Nicholls, R. D., Knoll, J. H., Butler, M. G., Karam, S. & Lalande, M. Genetic imprinting suggested by maternal heterodisomy in nondeletion Prader–Willi syndrome. Nature 342, 281–285 ( 1989).
Henry, I. et al. Uniparental parental disomy in a genetic cancer-predisposing syndrome. Nature 351, 665– 667 (1991).
Reik, W., Collick, A., Norris, M. L., Barton, S. C. & Surani, M. A. Genomic imprinting determines methylation of parental alleles in transgenic mice. Nature 328, 248–251 (1987).
Sapienza, C., Peterson, A. C., Rossant, J. & Balling, R. Degree of methylation of transgenes is dependent on gamete of origin. Nature 328, 251–254 ( 1987).
Swain, J. L., Stewart, T. A. & Leder, P. Parental legacy determines methylation and expression of an autosomal transgene: A molecular mechanism for parental imprinting. Cell 50, 719–727 ( 1987).
Chaillet, J. R., Vogt, T. F., Beier, D. R. & Leder, P. Parental-specific methylation of an imprinted transgene is established during gametogenesis and progressively changes during gametogenesis. Cell 66, 77–83 ( 1991).
Sasaki, H. et al. Inherited type of allelic methylation variations in a mouse chromosome region where an integrated transgene shows methylation imprinting . Development 111, 573– 581 (1991).
Li, E., Beard, C. & Jaenisch, R. Role for DNA methylation in genomic imprinting. Nature 366, 362–365 ( 1993).This paper showed that DNA methylation has a key role in maintaining imprinting. Imprinted expression of several genes was lost in a knockout of Dnmt1 , the maintenance methyltransferase gene.
Moore, T. & Haig, D. Genomic imprinting in mammalian development: a parental tug-of-war. Trends Genet. 7, 45–49 (1991).
Hurst, L. D. & McVean, G. T. Growth effects of uniparental disomies and the conflict theory of genomic imprinting. Trends Genet. 13, 436–443 ( 1997).
Feil, R., Khosla, S., Cappai, P. & Loi, P. Genomic imprinting in ruminants: allele-specific gene expression in parthenogenetic sheep. Mamm. Genome 9, 831–834 (1998).
McLaren, R. J. & Montgomery, G. W. Genomic imprinting of the insulin-like growth factor 2 gene in sheep. Mamm. Genome 10, 588–591 (1999).
Killian, J. K. et al. M6P/IGF2R imprinting evolution in mammals. Mol. Cell 5, 707–716 ( 2000).
Paulsen, M. et al. Sequence conservation and variability of imprinting in the Beckwith–Wiedemann syndrome gene cluster in human and mouse. Hum. Mol. Genet. 9, 1829–1841 (2000).
Neumann, B., Kubicka, P. & Barlow, D. P. Characteristics of imprinted genes. Nature Genet. 9, 12–13 ( 1995).
Stöger, R. et al. Maternal-specific methylation of the imprinted mouse Igf2r locus identifies the expressed locus as carrying the imprinting signal . Cell 73, 61–71 (1993).
Olek, A. & Walter, J. The pre-implantation ontogeny of the H19 methylation imprint. Nature Genet. 17 , 275–276 (1997).
Tremblay, K. D., Duran, K. L. & Bartolomei, M. S. A 5′ 2-kilobase-pair region of the imprinted mouse H19 gene exhibits exclusive paternal methylation throughout development . Mol. Cell. Biol. 17, 4322– 4329 (1997).
Shemer, R., Birger, Y., Riggs, A. D. & Razin, A. Structure of the imprinted mouse snrpn gene and establishment of its parental-specific methylation pattern. Proc. Natl Acad. Sci. USA 94, 10267–10272 (1997).
Feil, R., Walter, J., Allen, N. D. & Reik, W. Developmental control of allelic methylation in the imprinted mouse Igf2 and H19 genes . Development 120, 2933– 2943 (1994).
Ferguson, S. A., Sasaki, H., Cattanach, B. M. & Surani, M. A. Parental-origin-specific epigenetic modification of the mouse H19 gene . Nature 362, 751–755 (1993).
Bartolomei, M. S., Webber, A. L., Brunkow, M. E. & Tilghman, S. M. Epigenetic mechanisms underlying the imprinting of the mouse H19 gene . Genes Dev. 7, 1663–1673 (1993).
Feil, R., Boyano, M. D., Allen, N. D. & Kelsey, G. Parental chromosome-specific chromatin conformation in the imprinted U2af1-rs1 gene in the mouse. J. Biol. Chem. 272, 20893–20900 (1997).
Khosla, S., Aitchison, A., Gregory, R., Allen, N. D. & Feil, R. Parental allele-specific chromatin configuration in a boundary-imprinting-control element upstream of the mouse H19 gene. Mol. Cell. Biol. 19, 2556– 2566 (1999).
Hark, A. T. & Tilghman, S. M. Chromatin conformation of the H19 epigenetic mark. Hum. Mol. Genet. 7, 1979–1985 (1998).
Hu, J. F., Oruganti, H., Vu, T. H. & Hoffman, A. R. The role of histone acetylation in the allelic expression of the imprinted human insulin-like growth factor II gene. Biochem. Biophys. Res. Commun. 251, 403–408 (1998).
Pedone, P. V. et al. Role of histone acetylation and DNA methylation in the maintenance of the imprinted expression of the H19 and Igf2 genes. FEBS Lett. 458, 45–50 ( 1999).
Kitsberg, D. et al. Allele-specific replication timing of imprinted gene regions . Nature 364, 459–463 (1993).
Knoll, J. H., Cheng, S. D. & Lalande, M. Allele specificity of DNA replication timing in the Angelman/Prader–Willi syndrome imprinted chromosomal region. Nature Genet. 6, 41–46 ( 1994).
Paldi, A., Gyapay, G. & Jami, J. Imprinted chromosomal regions of the human genome display sex-specific meiotic recombination frequencies. Curr. Biol. 5, 1030–1035 (1995).
Robinson, W. P. & Lalande, M. Sex-specific meiotic recombination in the Prader–Willi/Angelman syndrome imprinted region . Hum. Mol. Genet. 4, 801– 806 (1995).
Brandeis, M. et al. The ontogeny of allele-specific methylation associated with imprinted genes in the mouse. EMBO J. 12, 3669–3677 (1993).
Tada, T. et al. Epigenotype switching of imprintable loci in embryonic germ cells . Dev. Genes Evol. 207, 551– 561 (1998).
Tada, M., Tada, T., Lefebvre, L., Barton, S. C. & Surani, M. A. Embryonic germ cells induce epigenetic reprogramming of somatic nucleus in hybrid cells. EMBO J. 16, 6510–6520 (1997).
Ueda, T. et al. The paternal methylation imprint of the mouse H19 locus is acquired in the gonocyte stage during foetal testis development. Genes Cells 5, 649–659 ( 2000).
Davis, T. L., Yang, G. J., McCarrey, J. R. & Bartolomei, M. S. The H19 methylation imprint is erased and reestablished differentially on the parental alleles during male germ cell development. Hum. Mol. Genet. 9, 2885–2894 (2000).
Labosky, P. A., Barlow, D. P. & Hogan, B. L. Mouse embryonic germ (EG) cell lines: transmission through the germline and differences in the methylation imprint of insulin-like growth factor 2 receptor (Igf2r) gene compared with embryonic stem (ES) cell lines. Development 120, 3197– 3204 (1994)
Obata,Y. et al. Disruption of primary imprinting during oocyte growth leads to the modified expression of imprinted genes during embryogenesis. Development 125,1553–1560 (1998).
Kato, Y. et al. Developmental potential of mouse primordial germ cells. Development 126, 1823–1832 (1999).References 47 and 48 showed that germ cell nuclei, when transplanted to zygotes, had a characteristically restricted potential for development. This is explained, at least in part, by the erasure of imprints in primordial germ cells.
Simon, I. et al. Asynchronous replication of imprinted genes is established in the gametes and maintained during development. Nature 401, 929–932 (1999). This work showed that asynchrony in DNA replication of imprinted regions is erased in germ cells.
Kafri, T. et al. Developmental pattern of gene-specific DNA methylation in the mouse embryo and germ line. Genes Dev. 6, 705–714 (1992).
Tada, T. et al. Imprint switching for non-random X-chromosome inactivation during mouse oocyte growth. Development 127, 3101 –3105 (2000).
Mertineit, C. et al. Sex-specific exons control DNA methyltransferase in mammalian germ cells. Development 125, 889– 897 (1998).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
Tucker, K. L. et al. Germ-line passage is required for establishment of methylation and expression patterns of imprinted but not of nonimprinted genes. Genes Dev. 10, 1008–1020 (1996).
Engemann, S. et al. Sequence and functional comparison in the Beckwith–Wiedeman region: implications for a novel imprinting centre and extended imprinting . Hum. Mol. Genet. 9, 2691– 2706 (2000).
Dorer, D. R. & Henikoff, S. Expansions of transgene repeats cause heterochromatin formation and gene silencing in Drosophila. Cell 77, 993–1002 ( 1994).
Robertson, K. D. et al. The human DNA methyltransferases (DNMTs) 1, 3a and 3b: coordinate mRNA expression in normal tissues and overexpression in tumors. Nucleic Acids Res. 27, 2291–2298 (1999).
Monk, M., Boubelik, M. & Lehnert, S. Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ cell lineages during mouse embryo development . Development 99, 371–382 (1987).
Oswald, J. et al. Active demethylation of the paternal genome in the mouse zygote . Curr. Biol. 10, 475–478 (2000).
Mayer, W., Niveleau, A., Walter, J., Fundele, R. & Haaf, T. Demethylation of the zygotic paternal genome. Nature 403, 501–502 ( 2000).
Howlett, S. K. & Reik, W. Methylation levels of maternal and paternal genomes during preimplantation development. Development 113, 119–127 (1991).
Rougier, N. et al. Chromosome methylation patterns during mammalian preimplantation development. Genes Dev. 12, 2108– 2113 (1998).References 58 – 62 showed that there is genome-wide demethylation in mouse preimplantation embryos followed by remethylation after implantation. The paternal genome is rapidly demethylated in the zygote, presumably by an active mechanism. The maternal genome is largely demethylated passively.
Carlson, L. L., Page, A. W. & Bestor, T. H. Properties and localization of DNA methyltransferase in preimplantation mouse embryos: implications for genomic imprinting. Genes Dev. 6, 2536–2541 (1992).
Bourc'his, D. et al. Abnormal methylation does not prevent X inactivation in ICF patients. Cytogenet. Cell Genet. 84, 245 –252 (1999).
Jouvenot, Y., Poirier, F., Jami, J. & Paldi, A. Biallelic transcription of Igf2 and H19 in individual cells suggests a post-transcriptional contribution to genomic imprinting. Curr. Biol. 9, 1199–1202 (1999).
Reik, W. & Maher, E. R. Imprinting in clusters: lessons from Beckwith–Wiedemann syndrome. Trends Genet. 13, 330–334 (1997).
Thorvaldsen, J. L., Duran, K. L. & Bartolomei, M. S. Deletion of the H19 differentially methylated domain results in loss of imprinted expression of H19 and Igf2. Genes Dev. 12, 3693–3702 (1998).Deletion of the differentially methylated region upstream of H19 results in loss of repression of the linked Igf2 on the maternal allele, and loss of H19 methylation and silencing on the paternal allele.
Bird, A. P. & Wolffe, A. P. Methylation-induced repression — belts, braces, and chromatin. Cell 99, 451–454 (1999).
Lee, J. T. Disruption of imprinted X inactivation by parent-of-origin effects at Tsix. Cell 103, 17– 27 (2000).
Lyle, R. et al. The imprinted antisense RNA at the Igf2r locus overlaps but does not imprint Mas1. Nature Genet. 25, 19–21 (2000).
Smilinich, N. J. et al. A maternally methylated CpG island in KvLQT1 is associated with an antisense paternal transcript and loss of imprinting in Beckwith–Wiedemann syndrome. Proc. Natl Acad. Sci. USA 96, 8064–8069 (1999).
Lee, M. P. et al. Loss of imprinting of a paternally expressed transcript, with antisense orientation to KVLQT1, occurs frequently in Beckwith–Wiedemann syndrome and is independent of insulin-like growth factor II imprinting. Proc. Natl Acad. Sci. USA 96, 5203– 5208 (1999).
Wutz, A. et al. Imprinted expression of the Igf2r gene depends on an intronic CpG island. Nature 389, 745– 749 (1997).Deletion of the intronic DMR in Igf2r leads to loss of imprinted repression on the paternal allele. The DMR contains a promoter for the antisense transcript Air that is not transcribed from the methylated maternal allele.
Horike, S. et al. Targeted disruption of the human LIT1 locus defines a putative imprinting control element playing an essential role in Beckwith–Wiedemann syndrome. Hum. Mol. Genet. 9, 2075– 2083 (2000).
Bell, A. C. & Felsenfeld, G. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature 405, 482–485 ( 2000).
Hark, A. T. et al. CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus. Nature 405, 486–489 (2000).
Kanduri, C. et al. Functional association of CTCF with the insulator upstream of the H19 gene is parent of origin-specific and methylation-sensitive . Curr. Biol. 10, 853–856 (2000).
Szabo, P., Tang, S. H., Rentsendorj, A., Pfeifer, G. P. & Mann, J. R. Maternal-specific footprints at putative CTCF sites in the H19 imprinting control region give evidence for insulator function. Curr. Biol. 10, 607–610 (2000).Refs 75 – 78 show that the DMR upstream of H19 contains a chromatin boundary that presumably blocks access of enhancers located downstream of H19 to the Igf2 promoters on the maternal chromosome. The unmethylated but not the methylated DMR binds the known boundary protein CTCF.
Sasaki, H. et al. Parental imprinting: potentially active chromatin of the repressed maternal allele of the mouse insulin-like growth factor II (Igf2) gene . Genes Dev. 6, 1843–1856 (1992).The first description of a differentially methylated region (DMR1), located upstream of the Igf2 gene. This work also showed that the maternal Igf2 promoters, although repressed, were neither methylated nor in a closed chromatin conformation.
Ishihara, K. et al. Comparative genomic sequencing identifies novel tissue-specific enhancers and sequence elements for methylation-sensitive factors implicated in Igf2/H19 imprinting. Genome Res. 10, 664–671 (2000).
Kaffer, C. R. et al. A transcriptional insulator at the imprinted H19/Igf2 locus. Genes Dev. 14, 1908– 1919 (2000).
Schmidt, J. V., Matteson, P. G., Jones, B. K., Guan, X. J. & Tilghman, S. M. The Dlk1 and Gtl2 genes are linked and reciprocally imprinted. Genes Dev. 14, 1997–2002 (2000).
Takada, S. et al. Delta-like and Gtl2 are reciprocally expressed, differentially methylated linked imprinted genes on mouse chromosome 12. Curr. Biol. 10, 1135–1138 (2000).
Wylie, A. A., Murphy, S. K., Orton, T. C. & Jirtle, R. L. Novel imprinted DLK1/GTL2 domain on human chromosome 14 contains motifs that mimic those implicated in IGF2/H19 regulation. Genome Res. 10, 1711–1718 ( 2000)
Constancia, M. et al. Deletion of a silencer element in Igf2 results in loss of imprinting independent of H19. Nature Genet. 26, 203–206 (2000).
Ainscough, J. F., John, R. M., Barton, S. C. & Surani, M. A. A skeletal muscle-specific mouse Igf2 repressor lies 40 kb downstream of the gene. Development 127, 3923– 3930 (2000).
Drewell, R. A. et al. Deletion of a silencer element disrupts H19 imprinting independently of a DNA methylation epigenetic switch. Development 127, 3419–3428 ( 2000).Refs 85 – 87 describe silencers in the Igf2 and H19 genes that are important for the regulation of imprinting and expression. These silencers are tissue-specific and may interact with different sets of enhancers. Silencer function may depend on methylation or chromatin modification.
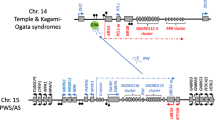
Buiting, K. et al. Inherited microdeletions in the Angelman and Prader–Willi syndromes define an imprinting center on human chromosome 15. Nature Genet. 9, 395–400 (1995).In patients with the imprinting disorders PWS or AS, an imprinting centre (IC) in the chromosome 15 imprinting cluster is deleted. Deletion of the IC alters imprinting of several genes in the whole domain.
Reik, W. et al. Imprinting mutations in the Beckwith–Wiedemann syndrome suggested by altered imprinting pattern in the IGF2-H19 domain. Hum. Mol. Genet. 4, 2379–2385 (1995).
Leighton, P. A., Ingram, R. S., Eggenschwiler, J., Efstratiadis, A. & Tilghman, S. M. Disruption of imprinting caused by deletion of the H19 gene region in mice. Nature 375, 34–39 (1995). A knockout of a region containing an imprinting centre (the H19 DMR) led to altered imprinting of the linked Igf2 and Ins2 genes.
Yang, T. et al. A mouse model for Prader–Willi syndrome imprinting-centre mutations. Nature Genet. 19, 25– 31 (1998).
Bielinska, B. et al. De novo deletions of SNRPN exon 1 in early human and mouse embryos result in a paternal to maternal imprint switch. Nature Genet. 25, 74–78 (2000).
Forne, T. et al. Loss of the maternal H19 gene induces changes in Igf2 methylation in both cis and trans. Proc. Natl Acad. Sci. USA 94, 10243–10248 ( 1997).
Paulsen, M. et al. Syntenic organization of the mouse distal chromosome-7 imprinting cluster and the Beckwith–Wiedemann-syndrome region in chromosome 11p15.5 . Hum. Mol. Genet. 7, 1149– 1159 (1998).
Caspary, T., Cleary, M. A., Baker, C. C., Guan, X. J. & Tilghman, S. M. Multiple mechanisms regulate imprinting of the mouse distal chromosome 7 gene cluster. Mol. Cell. Biol. 18, 3466–3474 (1998).
Maher, E. R. & Reik, W. Beckwith–Wiedemann syndrome: imprinting in clusters revisited. J. Clin. Invest. 105, 247–252 (2000).
Greally, J. M. et al. Conserved characteristics of heterochromatin-forming DNA at the 15q11–q13 imprinting center. Proc. Natl Acad. Sci. USA 96, 14430–14435 ( 1999).
McVean, G. T. & Hurst, L. D. Molecular evolution of imprinted genes: no evidence for antagonistic coevolution. Proc. R. Soc. Lond. B 264, 739–746 ( 1997).
Vrana, P. B., Guan, X. J., Ingram, R. S. & Tilghman, S. M. Genomic imprinting is disrupted in interspecific Peromyscus hybrids . Nature Genet. 20, 362– 365 (1998).
Macleod, D., Clark, V. H. & Bird, A. Absence of genome-wide changes in DNA methylation during development of the zebrafish. Nature Genet. 23, 139–140 (1999).
Efstratiadis, A. Genetics of mouse growth. Int. J. Dev. Biol. 42, 955–976 (1998).
Guillemot, F. et al. Genomic imprinting of Mash2, a mouse gene required for trophoblast development. Nature Genet. 9, 235–242 (1995).
Isles, A. R. & Wilkinson, L. S. Imprinted genes, cognition and behaviour. Trends Cog. Sci. 4, 309– 318 (2000).
Brambilla, R. et al. A role for the Ras signalling pathway in synaptic transmission and long-term memory. Nature 390, 281– 286 (1997).
Jiang, Y. H. et al. Mutation of the Angelman ubiquitin ligase in mice causes increased cytoplasmic p53 and deficits of contextual learning and long-term potentiation . Neuron 21, 799–811 (1998).
Lefebvre, L. et al. Abnormal maternal behaviour and growth retardation associated with loss of the imprinted gene Mest. Nature Genet. 20, 163–169 (1998).
Li, L. et al. Regulation of maternal behavior and offspring growth by paternally expressed Peg3. Science 284, 330– 333 (1999).
Steenman, M. J. et al. Loss of imprinting of IGF2 is linked to reduced expression and abnormal methylation of H19 in Wilms' tumour. Nature Genet. 7, 433–439 ( 1994); erratum in 8, 203 ( 1994).
Moulton, T. et al. Epigenetic lesions at the H19 locus in Wilms' tumour patients. Nature Genet. 7, 440– 447 (1994).
Buiting, K. et al. Sporadic imprinting defects in Prader–Willi syndrome and Angelman syndrome: implications for imprint-switch models, genetic counseling, and prenatal diagnosis. Am. J. Hum. Genet. 63, 170–180 (1998).
Dean, W. et al. Altered imprinted gene methylation and expression in completely ES cell-derived mouse fetuses: association with aberrant phenotypes. Development 125, 2273–2282 (1998).
Gurdon, J. B. & Colman, A. The future of cloning . Nature 402, 743–746 (1999).
Young, L. E., Sinclair, K. D. & Wilmut, I. Large offspring syndrome in cattle and sheep. Rev. Reprod. 3, 155–163 (1998).
Lanza, R. P. et al. Extension of cell life-span and telomere length in animals cloned from senescent somatic cells. Science 288, 665–669 (2000).
Kaneko-Ishino,T. et al. Peg1/Mest imprinted gene on chromosome 6 identified by cDNA subtraction hybridization. Nature Genet. 11 , 52–59 (1995).
Hayashizaki, Y. et al. Identification of an imprinted U2af binding protein related sequence on mouse chromosome 11 using the RLGS method. Nature Genet. 6, 33–40 (1994 ).
Peters, J. et al. A cluster of oppositely imprinted transcripts at the Gnas locus in the distal imprinting region of mouse chromosome 2. Proc. Natl Acad. Sci. USA 96, 3830– 3835 (1999).
Shemer, R. et al. The imprinting box of the Prader–Willi/Angelman syndrome domain. Nature Genet. 26, 440– 443 (2000).
Acknowledgements
We thank the many colleagues in our labs who are involved in imprinting work for discussions and comments on the manuscript, in particular W. Dean, M. Constancia, A. Murrell, R. Feil and G. Kelsey. We also thank three anonymous referees for excellent and constructive suggestions. Work in the authors' labs is supported by BBSRC, MRC, CRC, HFSP and DFG.
Author information
Authors and Affiliations
Related links
Related links
DATABASE LINKS
FURTHER INFORMATION
Glossary
- EUTHERIANS
-
Mammals that give birth to live offspring (viviparous) and possess an allantoic placenta.
- CPG ISLAND
-
DNA region of >500 bp that has a high CpG density and is usually unmethylated. CpG islands are found upstream of many mammalian genes.
- LINE-1 ELEMENT
-
A class of repetitive transposable element interspersed between and within genes throughout the genome. Most have degenerated and lost transposition activity.
- SILENCER
-
DNA sequence at which repressor factors bind and mediate silencing of promoters through interaction with the basal transcriptional machinery or the enhancer.
- BOUNDARY ELEMENT
-
DNA sequence that lies between two gene-controlling elements, such as a promoter and enhancer, preventing their communication or interaction. Boundary element function is usually mediated by the binding of specific factors.
- DNASEI HYPERSENSITIVE SITE
-
Chromosomal region highly accessible to cleavage by deoxyribonuclease I (DNAseI). Such sites are associated with open chromatin conformations and transcriptional activity.
- EPIGENOTYPE
-
Methylation and chromatin pattern of a gene or allele.
- MONOTREMES
-
Non-eutherian, egg-laying mammals.
- SPONGIOTROPHOBLAST
-
Junctional zone between the labyrinth and the maternal side of the placenta.
- LABYRINTHINE TROPHOBLAST
-
Placental zone where fetal and maternal exchange takes place.
- HYPERKINETIC, HYPOKINETIC
-
Exceeding (hyper) or reduced (hypo) movement of the body or extremeties.
Rights and permissions
About this article
Cite this article
Reik, W., Walter, J. Genomic imprinting: parental influence on the genome. Nat Rev Genet 2, 21–32 (2001). https://doi.org/10.1038/35047554
Issue Date:
DOI: https://doi.org/10.1038/35047554
This article is cited by
-
Genomic dissection of additive and non-additive genetic effects and genomic prediction in an open-pollinated family test of Japanese larch
BMC Genomics (2024)
-
Deciphering the Epigenetic Landscape: Placental Development and Its Role in Pregnancy Outcomes
Stem Cell Reviews and Reports (2024)
-
EpiTyping: analysis of epigenetic aberrations in parental imprinting and X-chromosome inactivation using RNA-seq
Nature Protocols (2023)
-
Decreased DIO3OS Expression Predicts Poor Prognosis in Hepatocellular Carcinoma and is Associated with Immune Infiltration
Biochemical Genetics (2023)
-
Differential Expression of RNAseq Imprinted Genes from Bovine Females Before and After Puberty
Biochemical Genetics (2023)