Abstract
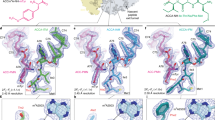
The 30S ribosomal subunit has two primary functions in protein synthesis. It discriminates against aminoacyl transfer RNAs that do not match the codon of messenger RNA, thereby ensuring accuracy in translation of the genetic message in a process called decoding. Also, it works with the 50S subunit to move the tRNAs and associated mRNA by precisely one codon, in a process called translocation. Here we describe the functional implications of the high-resolution 30S crystal structure presented in the accompanying paper, and infer details of the interactions between the 30S subunit and its tRNA and mRNA ligands. We also describe the crystal structure of the 30S subunit complexed with the antibiotics paromomycin, streptomycin and spectinomycin, which interfere with decoding and translocation. This work reveals the structural basis for the action of these antibiotics, and leads to a model for the role of the universally conserved 16S RNA residues A1492 and A1493 in the decoding process.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Green, R. & Noller, H. F. Ribosomes and translation. Annu. Rev. Biochem. 66, 679–716 (1997).
Agrawal, R. K. et al. Direct visualization of A-, P-, and E-site transfer RNAs in the Escherichia coli ribosome. Science 271, 1000–1002 (1996).
Stark, H. et al. Arrangement of tRNAs in pre- and posttranslocational ribosomes revealed by electron cryomicroscopy. Cell 88, 19–28 (1997).
Kurland, C. G. Translational accuracy and the fitness of bacteria. Annu. Rev. Genet. 26, 29–50 ( 1992).
Pape, T., Wintermeyer, W. & Rodnina, M. V. Complete kinetic mechanism of elongation factor Tu-dependent binding of aminoacyl-tRNA to the A site of the E. coli ribosome. EMBO J. 17, 7490–7497 ( 1998).
Pape, T., Wintermeyer, W. & Rodnina, M. Induced fit in initial selection and proof-reading of aminoacyl-tRNA on the ribosome. EMBO J. 18, 3800–3807 (1999).
Pape, T., Wintermeyer, W. & Rodnina, M. V. Conformational switch in the decoding region of 16S rRNA during aminoacyl-tRNA selection on the ribosome. Nature Struct. Biol. 7, 104–107 ( 2000).
Gabashvili, I. S. et al. Major rearrangements in the 70S ribosomal 3D structure caused by a conformational switch in 16S ribosomal RNA. EMBO J. 18, 6501–6507 (1999).
Nierhaus, K. H. The allosteric three-site model for the ribosomal elongation cycle: features and future. Biochemistry 29, 4997– 5008 (1990).
Allen, P. N. & Noller, H. F. Mutations in ribosomal proteins S4 and S12 influence the higher order structure of 16 S ribosomal RNA. J. Mol. Biol. 208, 457–468 (1989).
Lodmell, J. S. & Dahlberg, A. E. A conformational switch in Escherichia coli 16S ribosomal RNA during decoding of messenger RNA. Science 277, 1262– 1267 (1997).
Wimberly, B. T. et al. Structure of the 30S ribosomal subunit. Nature 407, 327–339 ( 2000).
Mueller, F. & Brimacombe, R. A new model for the three-dimensional folding of Escherichia coli 16 S ribosomal RNA. I. Fitting the RNA to a 3D electron microscopic map at 20 Å. J. Mol. Biol. 271, 524–544 (1997).
Cate, J. H., Yusupov, M. M., Yusupova, G. Z., Earnest, T. N. & Noller, H. F. X-ray crystal structures of 70S ribosome functional complexes. Science 285, 2095–2104 (1999).
Frank, J. & Agrawal, R. K. A ratchet-like inter-subunit reorganization of the ribosome during translocation. Nature 406, 319–322 (2000).
Stark, H., Rodnina, M. V., Wieden, H. J., van Heel, M. & Wintermeyer, W. Large-scale movement of elongation factor G and extensive conformational change of the ribosome during translocation. Cell 100, 301–309 (2000).
Moazed, D. & Noller, H. F. Binding of tRNA to the ribosomal A and P sites protects two distinct sets of nucleotides in 16 S rRNA. J. Mol. Biol. 211, 135–145 (1990).
Prince, J. B., Taylor, B. H., Thurlow, D. L., Ofengand, J. & Zimmermann, R. A. Covalent crosslinking of tRNA1Val to 16S RNA at the ribosomal P site: identification of crosslinked residues. Proc. Natl Acad. Sci. USA 79, 5450– 5454 (1982).
von Ahsen, U. & Noller, H. F. Identification of bases in 16S rRNA essential for tRNA binding at the 30S ribosomal P site. Science 267, 234–237 ( 1995).
Mueller, F., Stark, H., van Heel, M., Rinke-Appel, J. & Brimacombe, R. A new model for the three-dimensional folding of Escherichia coli 16 S ribosomal RNA. III. The topography of the functional centre. J. Mol. Biol. 271, 566– 587 (1997).
Garrett, R. A. (eds) The Ribosome. Structure, Function, Antibiotics and Cellular Interactions (ASM, Washington DC, 2000).
Kurland, C. G., Hughes, D., Ehrenberg, M. in Limitations of Translational Accuracy (eds Neidhardt, F. C. et al.) 979–1003 (ASM, Washington DC, 1996).
Bilgin, N., Richter, A. A., Ehrenberg, M., Dahlberg, A. E. & Kurland, C. G. Ribosomal RNA and protein mutants resistant to spectinomycin. EMBO J. 9, 735 –739 (1990).
Moazed, D. & Noller, H. F. Interaction of antibiotics with functional sites in 16S ribosomal RNA. Nature 327, 389–394 (1987).
Brink, M. F., Brink, G., Verbeet, M. P. & de Boer, H. A. Spectinomycin interacts specifically with the residues G1064 and C1192 in 16S rRNA, thereby potentially freezing this molecule into an inactive conformation. Nucleic Acids Res. 22, 325– 331 (1994).
Wittmann-Liebold, B. & Greuer, B. The primary structure of protein S5 from the small subunit of the Escherichia coli ribosome. FEBS Lett. 95, 91–98 (1978).
Ruusala, T. & Kurland, C. G. Streptomycin preferentially perturbs ribosomal proofreading. Mol. Gen. Genet. 198, 100–104 (1984).
Karimi, R. & Ehrenberg, M. Dissociation rate of cognate peptidyl-tRNA from the A-site of hyper- accurate and error-prone ribosomes. Eur. J. Biochem. 226, 355–360 (1994).
Karimi, R. & Ehrenberg, M. Dissociation rates of peptidyl-tRNA from the P-site of E. coli ribosomes. EMBO J. 15, 1149–1154 (1996).
Bilgin, N. & Ehrenberg, M. Mutations in 23 S ribosomal RNA perturb transfer RNA selection and can lead to streptomycin dependence. J. Mol. Biol. 235, 813–824 (1994).
Gravel, M., Melancon, P. & Brakier-Gingras, L. Cross-linking of streptomycin to the 16S ribosomal RNA of Escherichia coli. Biochemistry 26, 6227–6232 (1987).
Montandon, P. E., Wagner, R. & Stutz, E. E. coli ribosomes with a C912 to U base change in the 16S rRNA are streptomycin resistant. EMBO J. 5, 3705–3708 (1986).
Pinard, R., Payant, C., Melancon, P. & Brakier-Gingras, L. The 5′ proximal helix of 16S rRNA is involved in the binding of streptomycin to the ribosome. FASEB J. 7, 173– 176 (1993).
Melancon, P., Lemieux, C. & Brakier-Gingras, L. A mutation in the 530 loop of Escherichia coli 16S ribosomal RNA causes resistance to streptomycin. Nucleic Acids Res. 16, 9631–9639 ( 1988).
Montandon, P. E., Nicolas, P., Schurmann, P. & Stutz, E. Streptomycin–resistance of Euglena gracilis chloroplasts: identification of a point mutation in the 16S rRNA gene in an invariant position. Nucleic Acids Res. 13, 4299–4310 (1985).
Leclerc, D., Melancon, P. & Brakier-Gingras, L. Mutations in the 915 region of Escherichia coli 16S ribosomal RNA reduce the binding of streptomycin to the ribosome. Nucleic Acids Res. 19, 3973–3977 (1991).
Melancon, P., Boileau, G. & Brakier-Gingras, L. Cross-linking of streptomycin to the 30S subunit of Escherichia coli with phenyldiglyoxal. Biochemistry 23, 6697–6703 (1984).
Powers, T. & Noller, H. F. A functional pseudoknot in 16S ribosomal RNA. EMBO J. 10, 2203– 2214 (1991).
Bock, A., Petzet, A. & Piepersberg, W. Ribosomal ambiguity (ram) mutations facilitate diyhydrostreptomycin binding to ribosomes. FEBS Lett. 104, 317 –321 (1979).
Spahn, C. M. & Prescott, C. D. Throwing a spanner in the works: antibiotics and the translation apparatus. J. Mol. Med. 74, 423–439 (1996).
Fourmy, D., Recht, M. I., Blanchard, S. C. & Puglisi, J. D. Structure of the A site of Escherichia coli 16S ribosomal RNA complexed with an aminoglycoside antibiotic. Science 274, 1367–1371 (1996).
Yoshizawa, S., Fourmy, D. & Puglisi, J. D. Recognition of the codon–anticodon helix by ribosomal RNA. Science 285, 1722– 1725 (1999).
VanLoock, M. S., Easterwood, T. R. & Harvey, S. C. Major groove binding of the tRNA/mRNA complex to the 16 S ribosomal RNA decoding site. J. Mol. Biol. 285, 2069–2078 (1999).
Otwinowski, Z. & Minor, W. in Methods in Enzymology (eds Carter, C. W. J. & Sweet, R. M.) 307– 325 (Academic, New York, 1997).
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr D 54, 905–921 (1998).
Carson, M. Ribbons 2. 0. J. Appl. Cryst. 24, 958– 961 (1991).
Acknowledgements
This work was supported by the Medical Research Council (UK) and a grant from the US National Institutes of Health to V.R. and S.W.White. D.E.B. was supported by an EMBO long-term fellowship, and W.M.C. by an NIH predoctoral fellowship. We thank R. Ravelli for help with data collection, and M. Pacold for help and discussions with the modelling of antibiotic structures.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Carter, A., Clemons, W., Brodersen, D. et al. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 407, 340–348 (2000). https://doi.org/10.1038/35030019
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35030019
This article is cited by
-
Genome-wide identification of Kanamycin B binding RNA in Escherichia coli
BMC Genomics (2023)
-
Global translational control by the transcriptional repressor TrcR in the filamentous cyanobacterium Anabaena sp. PCC 7120
Communications Biology (2023)
-
A mini-review: environmental and metabolic factors affecting aminoglycoside efficacy
World Journal of Microbiology and Biotechnology (2023)
-
Chloroplast Engineering: Fundamental Insights and Its Application in Amelioration of Environmental Stress
Applied Biochemistry and Biotechnology (2023)
-
Reconstitution of microtubule into GTP-responsive nanocapsules
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.