Abstract
Complete genome sequences of 30 microbial species have been determined during the past five years, and work in progress indicates that the complete sequences of more than 100 further microbial species will be available in the next two to four years. These results have revealed a tremendous amount of information on the physiology and evolution of microbial species, and should provide novel approaches to the diagnosis and treatment of infectious disease.
Similar content being viewed by others
Main
Microbes were the first organisms on Earth and preceded animals and plants by more than 3 billion years. They are the foundation of the biosphere, from both an evolutionary and an environmental perspective1. It has been estimated that microbial species comprise about 60% of the Earth's biomass. The genetic, metabolic and physiological diversity of microbial species is far greater than that found in plants and animals. But the diversity of the microbial world is largely unknown, with less than one-half of 1% of the estimated 2–3 billion microbial species identified. Of those species that have been described, their biological diversity is extraordinary, having adapted to grow under extremes of temperature, pH, salt concentration and oxygen levels.
Perhaps no other area of research has been so energized by the application of genomic technology than the microbial field. It was only five years ago that The Institute for Genomic Research (TIGR) published the first complete genome sequence for a free-living organism, Haemophilus influenzae2; since that first report another 27 microbial genome sequences have been published, with at least 10–20 other projects at or near completion (for details see http://www.tigr.org/tdb/mdb/mdb.html). This progress represents, on average, one completed genome sequence every two months and all indications are that this pace will continue to accelerate. Included in the first completed microbial projects are many important human pathogens, the simplest known free-living organism, ‘model’ organisms, Escherichia coli and Bacillus subtilis, thermophilic bacterial species that might represent some of the deepest-branching members of the bacterial lineage, five representatives of the archaeal domain, and the first eukaryote, Saccharomyces cerevisiae. All of the organisms that have been studied by whole-genome analysis are species that can be grown either in the laboratory or in animal cells. It is important to remember that the vast majority of microbial species cannot be cultivated at all, and these organisms, which live in microbial communities, are essential to the overall ecology of the planet. Nevertheless, the study of ‘laboratory-adapted’ microbes has had a profound impact on our understanding of the biology and the evolutionary relationships between microbial species.
Methods for whole-genome analysis
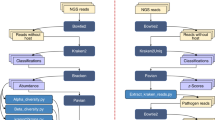
The method that was successfully used to determine the complete genome sequence of H. influenzae is a shotgun sequencing strategy ( Fig. 1). Before 1995, the largest genome sequenced with a random strategy was that of bacteriophage lambda with a genome size of 48,502 base pairs (bp), completed by Sanger et al. in 1982 (ref. 3). Despite advances in DNA-sequencing technology, the sequencing of whole genomes had not progressed beyond lambda-sized clones (about 40 kbp) because of the lack of sufficient computational approaches that would enable the efficient assembly of a large number of independent random sequences into a single contig.
For the H. influenzae and subsequent projects, we have used a computational method that was developed to create assemblies from hundreds of thousands of complementary DNA sequences 300–500-bp long4. This approach has proved to be a cost-effective and efficient approach to sequencing megabase-sized segments of genomic DNA. This strategy does not require an ordered set of cosmids or other subclones, thus significantly reducing the overall cost per base pair of producing a finished sequence, while providing high redundancy for accuracy and minimizing the effort required to obtain the whole genome sequence. The availability of improved technologies for longer sequence lengths (more than 700 bp) reduces problems associated with repetitive elements in the final assembly.
Microbial gene finding and annotation
The identification of genes in prokaryotic genomes has advanced to the stage at which nearly all protein-coding regions can be identified with confidence. Computational gene finders using Markov modelling techniques now routinely find more than 99% of protein-coding regions5 and RNA genes6. Once the protein-coding genes have been located, the most challenging problem is to determine their function. Typically, about 40–60% of the genes in a newly sequenced bacterial genome display a detectable sequence similarity to protein sequences whose function is at least tentatively known. This sequence similarity is the primary basis for assigning function to new proteins, but the transfer of functional assignments is fraught with difficulties.
To illustrate this problem, Table 1 contains an example showing the best matches in the database for a 1,344-bp gene from Mycobacterium tuberculosis at the time that the genome was being sequenced. All six of the best matches are kinases, but the specific names differ. A conservative naming strategy might use a family name that includes all six matches. Another strategy might use curated protein families (if they exist) to assign names; for example, the FGGY family named in the fourth line of Table 1 comes from the Pfam database7, a set of 1,815 hidden Markov models based on multiple alignments. By a closer examination of the literature, one could determine which of these protein names were based on laboratory experiments and which on sequence similarity. In any case, the assignment of a function to this protein requires the expertise of a skilled biologist. The rapidly changing nature of genome databases means that database searches must be repeated regularly to keep annotation accurate and up to date.
One possible solution to the annotation problem is to bring more of the resources of the scientific community to bear on each genome. No single centre can annotate all the functions of a living organism; experts from many different areas of biology should be encouraged to contribute to the annotation process. One possible model would be for geographically separated experts to deposit annotation to a central repository, which might also take on a curatorial or editorial role. An alternative model is one in which annotation resides in many different locations (as it does today), but in which new electronic links are created that allow scientists to locate rapidly all the information about a gene, genome or function. This latter model scales more easily and avoids the problem of overdependence on a single source.
What have we learned from genome analysis?
Comparison of the results from 24 completed prokaryotic genome sequences, containing more than 50 Mbp of DNA sequence and 54,000 predicted open reading frames (ORFs), has revealed that gene density in the microbes is consistent across many species, with about one gene per kilobase (Table 2). Almost half of the ORFs in each species are of unknown biological function. When the function of this large subset of genes begins to be explained, it is likely that entirely novel biochemical pathways will be identified that might be relevant to medicine and biotechnology. Perhaps even more unexpected is the observation that about a quarter of the ORFs in each species studied so far are unique, with no significant sequence similarity to any other available protein sequence. Although this might at present be an artefact of the small number of microbial species studied by whole-genome analysis, it nevertheless supports the idea that there is tremendous biological diversity between microorganisms. Taken together, these data indicate that much of microbial biology has yet to be understood and suggest that the idea of a ‘model’ organism in the microbial world might not be appropriate, given the vast differences between even related species.
Our molecular picture of evolution for the past 20 years has been dominated by the small-subunit ribosomal RNA phylogentic tree that proposes three non-overlapping groups of living organisms: the bacteria, the archaea and the eukaryotes8. Although the archaea possess bacterial cell structures, it has been suggested that they share a common ancestor exclusive of bacteria.
Analysis of complete genome sequences is beginning to provide great insight into many questions about the evolution of microbes. One such area has encompassed the occurrence of genetic exchanges between different evolutionary lineages, a phenomenon known as horizontal, or lateral, gene transfer. The occurrence of horizontal gene transfer, such as that involving genes from organellar genomes to the nucleus, or of antibiotic resistance genes between bacterial species, has been well established for many years (see, for example, ref. 9). This phenomenon causes problems for studying the evolution of species because it means that some species are chimaeric, with different histories for different genes. Before the availability of complete genome sequences, studies of horizontal gene transfer had been limited because of the incompleteness of the data sets being analysed. Analyses of complete genome sequences have led to many recent suggestions that the extent of horizontal gene exchange is much greater than was previously realized10,11,12. For example, an analysis of the genomes of two thermophilic bacterial species, Aquifex aeolicus and Thermotoga maritima, revealed that 20–25% of the genes in these species were more similar to genes from archaea than those from bacteria13,14. This led to the suggestion of possible extensive gene exchanges between these species and archaeal lineages. But before one jumps to this conclusion it is important to consider the difficulties in inferring the occurrence of gene transfer. For example, the high percentage of genes with best matches to archaea in A. aeolicus and T. maritima could also be due to a high rate of evolution in the mesophilic bacteria (which would cause thermophilic and archaeal genes to have high levels of similarity despite their not having a common ancestry) or the loss of these genes from mesophilic bacteria15. For T. maritima, many lines of additional evidence support the assertion of gene transfer, including the observation that many of the archaeal-like genes occur in clusters in the genome, are in regions of unusual nucleotide composition, and branch in phylogenetic trees most closely to archaeal genes14. Most of the lines of evidence leading to assertions of horizontal gene transfer can have other causes. For example, unusual nucleotide composition can also arise from selection16, and differences in phylogenetic trees can be caused by convergence, inaccurate alignments17, long-branch attraction18 or sampling of different species19. It is therefore important to assess the evidence carefully and to find multiple types of evidence. This has yet to be done systematically, so we believe that it is too early to assign quantitative values to the extent of gene exchange between species.
Despite the apparent occurrence of extensive gene transfers in the history of microbes, it does seem that there might be a ‘core’ to each evolutionary lineage that retains some phylogenetic signal. The best evidence for this comes from the construction of ‘whole genome trees’ based on the presence and absence of particular homologues or orthologues in different complete genomes20. It is important to note that gene-content trees are averages of patterns produced by phylogeny, gene duplication and loss, and horizontal transfer; they are therefore not real phylogenetic trees. Nevertheless, the fact that these trees are very similar to phylogenetic trees of genes such as ribosomal RNA and RecA suggests that although horizontal gene transfer might be extensive, it is somehow constrained by phylogenetic relationships. Other evidence for a ‘core’ of particular lineages comes from the finding of a conserved core of euryarchaeal genomes21,22 and another finding that some types of gene might be more prone to gene transfer than others23. It therefore seems likely that horizontal gene transfer has not completely obliterated the phylogenetic signal in microbial genomes. Careful studies in which the phylogenetic trees of some of these core genes are compared across all genomes need to be done to see whether or not the core has a consistent phylogeny. Initial studies suggest that it does, at least for the major microbial groups14.
Although our ability to resolve patterns of the relationships among microbes is still limited, analysis of the genomes of closely related species is revealing much about genome evolution24,25. For example, a comparison of the genomes of four chlamydial species has revealed the occurrence of frequent tandem gene duplication and gene loss, as well as large chromosomal inversions25. Comparisons of closely related species should also reveal much about mutation processes, codon usage and other features that evolve rapidly16.
Design of new antimicrobial agents and vaccines
One of the expected benefits of genome analysis of pathogenic bacteria is in the area of human health, particularly in the design of more rapid diagnostic reagents and the development of new vaccines and antimicrobial agents. These goals have become more urgent with the continuing spread of antibiotic resistance in important human pathogens. Moreover, results from the whole-genome analysis of human pathogens has suggested that there are mechanisms for generating antigenic variation in proteins expressed on the cell surface that are encoded within the genomes of these organisms. These mechanisms include the following: (1) slipped-strand mispairing within DNA sequence repeats found in 5′-intergenic regions and coding sequences as described for H. influenzae2, Helicobacter pylori26 and M. tuberculosis27, (2) recombination between homologous genes encoding outer-surface proteins as described for Mycoplasma genitalium28, Mycoplasma pneumoniae29 and Treponema pallidum30, and (3) clonal variability in surface-expressed proteins as described for Plasmodium falciparum31 and possibly Borrelia burgdorferi 32. Experimental evidence from studies of clinical isolates of some species has demonstrated phenotypic variation in the relevant cell-surface proteins33, suggesting that, at least for human pathogens, the evolution of antigenic proteins probably occurs in real time, as cell populations divide. The ability of human pathogens to alter their antigenic potential and thereby evade the immune system has the potential to hinder vaccine development by conventional methods.
Progress during the past year has supported the idea that complete genome sequence information can be exploited in the design of new vaccines and antimicrobial compounds. As an example, the identification of new vaccine candidates against serogroup B Neisseria meningitidis (MenB) was reported by Pizza et al. using a genomics-based approach34 ( Fig. 2). With the use of the entire genome sequence of a virulent serogroup B strain35, 570 putative cell-surface-expressed or secreted proteins were identified; the corresponding DNA sequences were cloned and expressed in E. coli. Of the putative targets, 61% were expressed successfully and used to immunize mice. Immune sera were screened for bactericidal activity and for the ability to bind to the surface of MenB cells. Seven representative proteins were selected for further study and were evaluated for their degree of sequence variability among multiple isolates and serogroups of N. meningitidis . Two highly conserved vaccine candidates emerged from this large-scale screening effort, which occurred in parallel with the completion of the genome sequence of N. meningitidis. These results provide the first definitive demonstration of the potential of genomic information to expand and accelerate the development of vaccines against pathogenic organisms.
Another example illustrates the potential of genomics to accelerate the development of novel antimicrobial agents. Jomaa et al.36 identified two genes in P. falciparum from sequence data from the malaria genome consortium that encode key enzymes in the 1-deoxy-d-xylulose-5-phosphate (DOXP) pathway that are required for the synthesis of isoprenoids such as cholesterol37. The DOXP pathway functions in some bacteria, algae and higher plants to produce isopentenyl diphosphate, a precursor of isoprenoids. In P. falciparum, the enzymes of the DOXP pathway are probably associated with a specialized organelle derived from algae called the apicoplast; they are expressed when the parasite is growing within red blood cells. Inhibitors of one of the key enzymes in the DOXP pathway, DOXP reductoisomerase, had previously been identified and had been shown to inhibit the bacterial enzyme and the growth of some bacterial species. Jomaa et al.36 demonstrated that two inhibitors of DOXP reductoisomerase, fosidomycin and FR900098, were able to inhibit the growth of P. falciparum in vitro and cure mice infected with a related species of Plasmodium . Both of these compounds exhibit low toxicity and high stability and are relatively inexpensive to produce, suggesting that they might be the basis of a potentially important new class of anti-malarial drugs.
Conclusions
So far, studies in genomics have only scratched the surface of microbial diversity and have revealed how little is known about microbial species. In the next few years, more than 100 projects for sequencing microbial genomes should be completed, providing the scientific community with information on more than 300,000 predicted genes. A significant number of these genes will be novel and of unknown function. These novel genes represent exciting new opportunities for future research and potential sources of biological resources to be explored and exploited. The benefits of comparative genomics in understanding biochemical diversity, virulence and pathogenesis, and the evolution of species has been unequivocally demonstrated and the usefulness of comparative techniques will improve as more genomes become available. One of the major challenges is to develop techniques for assessing the function of novel genes on a large scale and integrating information on how genes and proteins interact at the cellular level to create and maintain a living organism. It is not unreasonable to expect that, by expanding our understanding of microbial biology and biodiversity, great strides can be made in the diagnosis and treatment of infectious diseases and in the identification of useful functions in the microbial world that could be applied to agricultural and industrial processes.
References
Staley, J. J. et al. The Microbial World. The Foundation of the Biosphere (American Society for Microbiology, Washington DC, 1997).
Fleischmann, R. D. et al. Whole-genome random sequencing and assembly of Haemophilus influenzae. Science 269, 496– 512 (1995).
Sanger, F., Coulson, A. R., Hong, G. F., Hill, D. F. & Petersen, S. Nucleotide sequence of bacteriophage lambda DNA. J. Mol. Biol. 162, 729 (1982).
Sutton, G. G., White, O., Adams, M. D. & Kerlavage, A. R. TIGR Assembler: a new tool for assembling large shotgun sequencing projects. Genome Sci. Technol. 1, 9–19 ( 1995).
Delcher, A. L., Harmon, D., Kasif, S., White, O. & Salzberg, S. L. Improved microbial gene identification with Glimmer . Nucleic Acids Res. 27, 4636– 4641 (1999).
Lowe, T. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 ( 1997).
Bateman, A. et al. The Pfam Protein Families Database. Nucleic Acids Res. 28, 263–266 ( 2000).
Woese, C. R. & Fox, G. E. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc. Natl Acad. Sci. USA 74, 5088–5090 (1977).
Davies, J. Origins and evolution of antibiotic resistance. Microbiologia 12, 9–16 (1996).
Lawrence, J. G. & Ochman, H. Molecular archaeology of the Escherichia coli genome. Proc. Natl. Acad. Sci. USA 95, 9413–9417 ( 1998).
Doolittle, W. F. Phylogenetic classification and the universal tree. Science 284, 2124–2129 (1999).
Martin, W. Mosaic bacterial chromosomes: a challenge en route to a tree of genomes. Bioessays 21, 99–104 ( 1999).
Aravind, L., Tatusov, R. L., Wolf, Y. I., Walker, D. R. & Koonin, E. V. Evidence for massive gene exchange between archaeal and bacterial hyperthermophiles. Trends Genet. 14, 442–444 ( 1998).
Nelson, K. E. et al. Genome sequencing of Thermotoga maritima: evidence for lateral gene transfer between bacteria and archaea. Nature 399, 323–329 ( 1999).
Kyrpides, N. C. & Olsen, G. J. Archaeal and bacterial hyperthermophiles: horizontal gene exchange or common ancestry? Trends Genet 15, 298–299 (1999).
Lafay, B. et al. Proteome composition and codon usage in spirochaetes: species-specific and DNA strand-specific mutational biases. Nucleic Acids Res. 27, 1642–1649 (1999).
Gatesy, J., Desalle, R. & Wheller, W. Alignment-ambiguous nucleotide sites and the exclusion of systematic data. Mol. Phylog. Evol. 2, 152–157 (1993).
Philippe, H. & Forterre, P. The rooting of the universal tree of life is not reliable. J. Mol. Evol. 49, 509–523 (1999).
Eisen, J. A. The RecA protein as a model molecule for molecular systematic studies of bacteria: comparison of trees of RecAs and 16s rRNAs from the same species. J. Mol. Evol. 41, 1105–1123 (1995).
Snel, B., Bork, P. & Huynen, M. A. Genome phylogeny based on gene content. Nature Genet. 21, 108–110 (1999).
Makarova, K. S. et al. Comparative genomics of the Archaea (Euryarchaeota): evolution of conserved protein families, the stable core, and the variable shell. Genome Res. 9, 608–628 ( 1999).
22. Graham, D. E., Overbeek, R., Olsen, G. J. & Woese, C. R. An archaeal genomic signature. Proc. Natl. Acad. Sci. USA 97, 3304–3308 (2000).
Jain, R., Rivera, M. C. & Lake, J. A. Horizontal gene transfer among genomes: the complexity hypothesis. Proc. Natl. Acad. Sci. USA 96, 3801–3806 (1999).
Alm, R. A. et al. Genomic-sequence comparison of two unrelated isolates of the human gastric pathogen, Helicobacter pylori. Nature 397,176–180 (1999).
Read, T. D. et al. Genome sequences of Chlamydia trachomatis MoPn and Chlamydia pneumoniae AR39. Nucleic Acids Res. 28 ,1397–1406 (2000).
Tomb, J. F. et al. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 388, 539– 547 (1997).
Cole, S. T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Fraser, C. M. et al. The minimal gene complement of Mycoplasma genitalium. Science 270, 397–403 (1995).
Himmelreich, R. et al. Complete sequence analysis of the genome of the bacterium, Mycoplasma pneumoniae. Nucleic Acids Res. 24, 4420–4429 (1996).
Fraser, C. M. et al. Complete genome sequence of Treponema pallidum, the syphilis spirochete. Science 281, 375– 388 (1998).
Gardner, M. J. et al. Chromosome 2 sequence of the human malaria parasite Plasmodium falciparum. Science 282, 1126– 1132 (1998).
Fraser, C. M. et al. Genomic sequence of a Lyme disease spirochaete, Borrelia burgdorferi. Nature 390, 580– 586 (1997).
Peterson, S. N. et al. Characterization of repetitive DNA in the Mycoplasma genitalium genome: possible role in the generation of antigenic variation. Proc. Natl Acad. Sci. USA 92,11829– 11833 (1995).
Pizza, M. et al. Identification of vaccine candidates against serogroup B meningococcus by whole genome sequencing. Science 287, 1816–1820 (2000).
Tettelin, H. et al. Complete genome sequence of Neisseria meningitidis serogroup B strain MC58. Science 287, 1809 –1815 (2000).
Jomaa, H. et al. Inhibitors of the nonmevalonate pathway of isoprenoid biosynthesis as antimalarial drugs. Science 285, 1573 –1576 (1999).
Ridley, R. G. Planting the seeds of new antimalarial drugs. Science 285, 1502–1503 (1999).
Author information
Authors and Affiliations
Rights and permissions
This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fraser, C., Eisen, J. & Salzberg, S. Microbial genome sequencing. Nature 406, 799–803 (2000). https://doi.org/10.1038/35021244
Published:
Issue Date:
DOI: https://doi.org/10.1038/35021244
This article is cited by
-
Influence of nitric acid on biodegradation of polystyrene and low-density polyethylene by Cephalosporium species
Archives of Microbiology (2022)
-
Prokaryotic and eukaryotic diversity in hydrothermal continental systems
Archives of Microbiology (2021)
-
Estimate of the sequenced proportion of the global prokaryotic genome
Microbiome (2020)
-
Vibrio spp. infections
Nature Reviews Disease Primers (2018)
-
Pcal_0111, a highly thermostable bifunctional fructose-1,6-bisphosphate aldolase/phosphatase from Pyrobaculum calidifontis
Extremophiles (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.