Abstract
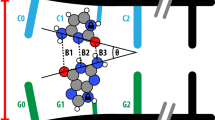
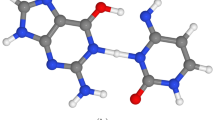
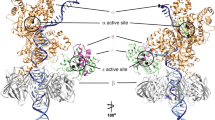
The opening of base pairs of double-stranded DNA is an important process, being a prerequisite for replication and transcription and possibly a factor in the recognition, flexibility and structure of DNA. The kinetics of base-pair opening have, however, been controversial. Base-pair opening can be studied by following the exchange of protons from imino groups with water, a process that seems only to occur from open base pairs. We have recently demonstrated catalysis by proton acceptors of imino proton exchange in nucleic acids. This has enabled us to determine the base-pair lifetimes, which are in the region of 10 ms at room temperature1,2. In earlier reports it had been considered that proton exchange is limited by the rate of base-pair opening, which had led to estimates of base-pair lifetimes that were larger by one or two orders of magnitude3–17. There are also important discrepan-cies between recent and early estimates of the base-pair dissociation constant. Earlier estimates of base-pair lifetimes correspond in fact to the time required for proton exchange in the absence of added catalyst (AAC exchange). This could be a distinct mode of base-pair opening with a very long open lifetime, different from the mode revealed by the effect of catalyst. The evidence reported here suggests on the contrary that there is only a single mode of base-pair opening and that proton exchange in the absence of added catalyst is in fact catalysed by a proton acceptor intrinsic to the nucleic acid, most probably the other base of the open pair.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
1. Leroy, J. L., Broseta, D. & Gueron, M. 7. molec. Biol. 184, 165–178 (1985). 2. Gueron, M., Kochoyan, M. & Leroy, J. L. in Proc. First EBSA Workshop Saltsjobaden, July 1986 (eds Ehrenberg, A., Rigler, R., Graslund, A. & Nilsson, L.) 196–200 (Springer, Berlin, 1987). 3. Teitelbaum, H. & Englander, S. W. J. molec. Biol. 92, 55–78 (1975). 4. Teitelbaum, H. & Englander, S. W. 7. molec. Biol. 92, 79–92 (1975). 5. Mandal, C., Kallenbach, N. R. & Englander, S. W. 7. molec. Biol. 135, 391–411 (1979). 6. Nakanishi, N., Mitane, Y. & Tsuboi, M. Biochim. biophys. Acta 798, 46–52 (1984). 7. Mirau, P. A. & Kearns, D. R. 7. molec. Biol. 77, 207–227 (1984). 8. Mirau, P. A. & Kearns, D. R. Biopolymers 24, 711–724 (1985). 9. Hartmann, B., Leng, M. & Ramstein, J. Biochemistry 25, 3073–3077 (1986). 10. Patel, D. J. & Hilbers, C. W. Biochemistry 14, 2651–2656 (1975). 11. Early, T. A., Kearns, D. R., Hillen, W. & Wells, R. D. Biochemistry 20, 3756–3764 (1981). 12. Pardi, A. & Tinoco, I. Jr Biochemistry 24, 4686–4693 (1982). 13. Chou, S.–H., Wemmer, D. E., Hare, D. R. & Reid, B. R. Biochemistry 23, 2257–2262 (1984). 14. Patel, D. J., Kozlowski, S. A., Weiss, M. & Bhatt, R. Biochemistry 24, 936–944 (1985). 15. Lefevre, J. F., Lane, A. N. & Jardetzky, O. 7. molec. Biol. 185, 689–699 (1985). 16. Englander, S. W. & Kallenbach, N. R. Q. Rev. Biophys. 16, 521–655 (1984). 17. Cantor, C. R. & Schimmel, P. R. Biophysical Chemistry Part III Ch. 22 (Freeman, San Francisco, 1980). 18. Figueroa, N. et al. Proc. natn. Acad. Sci. U.S.A. 80, 4330–4333 (1983). 19. Leroy, J. L., Bolo, N., Figueroa, N., Plateau, P. & Gueron, M. 7. biomolec. struct. Dynam. 2,915–939(1985). 20. Crothers, D. M., Cole, P. E., Hilbers, C. W. & Shulman, R. G. 7. molec. Biol. 87,63–88 (1974). 21. Johnston, P. D. & Redfield, A. G. Nucleic. Acids Res. 5, 3913–3927 (1978). 22. Frank–Kamenetskii, M. D. in Structure and Motion: Membranes, Nucleic Acids and Proteins (eds Clementi, E., Corongin, G., Sarma, M. H. & Sarma, H.), 417–432 (Adenine Press, Guilderland, New York, 1985). 23. Wilcoxon, J. & Schurr, J. M. Biopolymers 22, 2273–2321 (1983). 24. Williams, M. N. & Crothers, D. M. Biochemistry 14, 1944–1951 (1275). 25. Grunwald, E., Lowenstein, A. & Meiboom, S. 7. Chem. Phys. 27, 630–640 (1957). 26. Bianchin, B. et al. in Protons and Ions Involved in Fast Dynamic Phenomena (ed. Lazlo, P.) 265–282 (Elsevier, Amsterdam, 1978). 27. Gralla, J. & Crothers, D. M. 7. molec. Biol. 78, 301–319 (1973). 28. Lukashin, A. V., Vologodskii, A. V., Frank–Kamenetskii, M. D. & Lyubchenko, Y. L. 7. molec. Biol 108, 665–682.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Guéron, M., Kochoyan, M. & Leroy, JL. A single mode of DNA base-pair opening drives imino proton exchange. Nature 328, 89–92 (1987). https://doi.org/10.1038/328089a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/328089a0
This article is cited by
-
Mapping of DNA base-pair sequence from breathing dynamics of hetero-polymeric DNA: A genetic algorithm-based study
Journal of Chemical Sciences (2024)
-
Uracil-DNA glycosylase efficiency is modulated by substrate rigidity
Scientific Reports (2023)
-
Nonlinear physics opens a new paradigm for accurate transcription start site prediction
BMC Bioinformatics (2022)
-
A quantitative model predicts how m6A reshapes the kinetic landscape of nucleic acid hybridization and conformational transitions
Nature Communications (2021)
-
DNA mismatches reveal conformational penalties in protein–DNA recognition
Nature (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.