Abstract
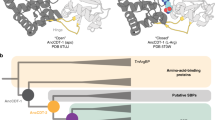
The binding of substrates to lactate dehydrogenases induces a marked rearrangement of the protein structure in which a ‘loop’ of polypeptide (residues 98–110) closes over the active site of the enzyme1,2. In this rearrangement, arginine 109 (a basic residue conserved in all known lactate dehydrogenase sequences and in the homologous malate dehydrogenases3) moves 0.8 nm from a position in the solvent4 to one in the active site1,5,6 where its guanidinium group resides within hydrogen bonding distance of both the reactive carbonyl of pyruvate and imidazole ring of the catalytic histidine 195 (see Fig. 1). Whilst this feature of the enzyme has been commented upon previously1, the function of this mobile arginine residue during catalysis has not been tested experimentally. The advent of protein engineering has now enabled us to define the role of this basic residue by substituting it with the neutral glutamine. Transient kinetic and equilibrium studies of the mutant enzyme indicate that arginine 109 enhances the polarization of the pyruvate carbonyl group in the ground state and stabilizes the transition state. The gross active-site structure of the enzyme is not altered by the mutation since an alternative catalytic function of the enzyme (rate of addition of sulphite to NAD+), which does not require hydride transfer, is insensitive to the arginine→glutamine substitution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Grau, U. M., Trommer, W. E. & Rossmann, M. G. J. molec. Biol. 151, 289–307 (1981).
Parker, D. M. & Holbrook, J. J. Pyridine Nucleotide-Dependent Dehydrogenases 485–502 (ed. Sund, H.) (de Gruyter, Berlin, 1977).
Birktoft, J. J., Fernley, R. T., Bradshaw, R. A. & Banaszak, L. J. Proc. natn. Acad. Sci. U.S.A. 79, 6166–6170 (1982).
Adams, M. J. et al. J. molec. Biol. 41, 159–188 (1969).
Adams, M. J. et al. Proc. natn. Acad. Sci. U.S.A. 70, 1968–1972 (1973).
White, J. L. et al. Molec. Biol. 102, 759–779 (1976).
Barstow, D. A. et al. Gene 46, 47–55 (1986).
Wirz, B., Suter, F. & Zuber, H. Hoppe-Seyler's Z. physiol. Chem. 364, 893–909 (1983).
Clarke, A. R., Atkinson, T., Campbell, J. W. & Holbrook, J. J. Biochim. biophys. Acta 829, 387–396 (1985).
Winter, G., Fersht, A. R., Wilkinson, A. J., Zoller, M. & Smith, M. Nature 299, 756–758 (1982).
Amann, E., Brosius, J. & Ptashne, M. Gene 25, 167–178 (1983).
Gibson, T. J. thesis, Univ. Cambridge (1984).
Clarke, A. R., Waldman, A. D. B., Munro, I. & Holbrook, J. J. Biochim. biophys. Acta 828, 375–379 (1985).
Atkinson, T. et al. in Bioactive Microbial Products (ed. Stowell, J. D.) Vol. 3, 27–43 (Academic, London 1986).
Holbrook, J. J., Liljas, A., Steindel, S. J. & Rossmann, M. G. in The Enzymes (ed. Boyer, P. D.), 3rd Edn Vol. 11, 191–292 (Academic, New York, 1975).
Parker, D. M., Jeckel, D. & Holbrook, J. J. Biochem. J. 201, 465–471 (1982).
Holbrook, J. J. & Stinson, R. A. Biochem. J. 131, 739–748 (1973).
Parker, D. M., Lodola, A. & Holbrook, J. J. Biochem. J. 173, 959–967 (1978).
Johnson, S. L. & Smith, K. W. Biochemistry 15, 553–559 (1976).
Holbrook, J. J. & Gutfreund, H. FEBS Lett. 31, 157–169 (1973).
Klinman, J. P. Adv. Enzym. relat. Areas molec. Biol. 46, 415–493 (1978).
Lodola, A., Shore, J. D., Parker, D. M. & Holbrook, J. J. Biochem. J. 175, 987–998 (1978).
Clarke, A. R., Waldman, A. D. B., Hart, K. W. & Holbrook, J. J. Biochim. biophys. Acta 829, 397–407 (1985).
Winter, A. D. & Schwert, G. W. J. biol. Chem. 234, 1155–1161 (1959).
Ackers, G. K. & Smith, F. R. A. Rev. Biochem. 54, 597–629 (1985).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Clarke, A., Wigley, D., Chia, W. et al. Site-directed mutagenesis reveals role of mobile arginine residue in lactate dehydrogenase catalysis. Nature 324, 699–702 (1986). https://doi.org/10.1038/324699a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/324699a0
This article is cited by
-
The tumor suppressor folliculin inhibits lactate dehydrogenase A and regulates the Warburg effect
Nature Structural & Molecular Biology (2021)
-
LUD, a new protein domain associated with lactate utilization
BMC Bioinformatics (2013)
-
Lactate production yield from engineered yeasts is dependent from the host background, the lactate dehydrogenase source and the lactate export
Microbial Cell Factories (2006)
-
Catalytic reaction mechanism of L-lactate dehydrogenase: anab initio study
Science in China Series B: Chemistry (2000)
-
Simulation of enzyme–substrate encounter with gated active sites
Nature Structural & Molecular Biology (1994)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.