Abstract
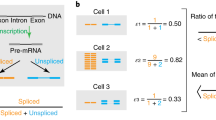
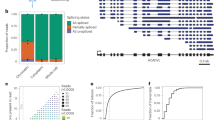
Alternative splicing of eukaryotic messenger RNA precursors is now known to be of widespread importance in generating multiple transcripts from a single gene. This phenomenon has emphasized the problem of the way in which splice sites are selected; recent studies have discussed the role of secondary structure1 or affinity and spatial relationships2 in this selection. Splice site sequences vary widely, although a loose consensus has been derived for the 9 bases around the 5′ splice site and for a longer region around the 3′ splice site3. Mutagenesis experiments have defined the sequences essential for a potential 5′ splice site4,5,6, but, except for some experiments with the E1a gene of adenovirus5,6, these experiments have not examined 5′ splice site sequences for features responsible for site preference where alternative splicing sites exist. Such tests require a choice of site: an appropriate reference site and a constant position at which test sites are introduced. We have begun a series of experiments designed to show whether splice site sequences can be ranked in a hierarchy of preferential use. Here we show that the archetypal consensus sequence is used efficiently, and characterize the cryptic sites of β-globin: sequences alone can explain why these sites are not normally used. We also show with the E1a gene of adenovirus, a simple example of alternative splicing, that one of the two 5′ splice sites used by this gene is intrinsically stronger. We also demonstrate that tandem repeats and secondary structure influence the choice of sites in vivo. We discuss the mechanism of splice site selection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Solnick, D. Cell 43, 667–676 (1985).
Kühne, T., Wieringa, B., Reiser, J. & Weissman, C. EMBO J. 2, 727–733 (1983).
Mount, S. M. Nucleic Acids Res. 10, 459–472 (1982).
Wieringa, B., Meyer, F., Reiser, J. & Weissmann, C. Nature 301, 38–43 (1983).
Montell, C. & Berk, A. J. Nucleic Acids Res. 12, 3821–3827 (1984).
Montell, C., Fisher, E. F., Caruthers, M. H. & Berk, A. J. Nature 295, 380–384 (1982).
Grosveld, G. C., de Boer, E., Shewmaker, C. K. & Flavell, R. A. Nature 295, 120–126 (1982).
Frendewey, D. & Keller, W. Cell 42, 355–367 (1985).
Grabowski, P. J., Seiler, S. R. & Sharp, P. A. Cell 42, 345–353 (1985).
Perkins, K. K., Furneaux, H. M. & Hurwitz, J. Proc natn Acad. Sci. U.S.A. 83, 887–891 (1986).
Estibeiro, J. P. (in preparation).
Sanger, F., Nicklen, S. & Coulson, A. R. Proc. natn. Acad. Sci. U.S.A. 74, 5463–5467 (1977).
Auffray, C. & Rougeon, F. Eur. J. Biochem. 107, 303–314 (1980).
Favaloro, J., Treisman, R. & Kamen, R. Meth. Enzym. 65, 718–749 (1980).
Sanger, F. & Coulson, A. R. FEBS Lett. 87, 107–110 (1978).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Eperon, L., Estibeiro, J. & Eperon, I. The role of nucleotide sequences in splice site selection in eukaryotic pre-messenger RNA. Nature 324, 280–282 (1986). https://doi.org/10.1038/324280a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/324280a0
This article is cited by
-
Human mucin MUC1 RNA undergoes different types of alternative splicing resulting in multiple isoforms
Cancer Immunology, Immunotherapy (2013)
-
Unusual splice sites in the E1A?E1B cotranscripts synthesized in adenovirus type 40-infected A549 cells
Archives of Virology (1994)
-
Applications of gene transfer to study rna splicing in mammalian cell lines
Journal of Tissue Culture Methods (1993)
-
Are vertebrate exons scanned during splice-site selection?
Nature (1992)
-
Scanning from an independently specified branch point defines the 3′ splice site of mammalian introns
Nature (1989)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.