Abstract
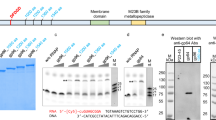
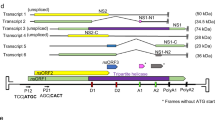
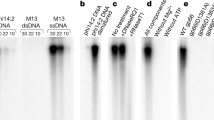
Drosophila cells contain virus-like particles (VLPs) containing 5 kilobases (kb) of RNA (VLP H-RNA) homologous to the transposable element copia1. The identity between VLP H-RNA and copia DNA has previously been confirmed at the nucleotide sequence level2 and reverse transcriptase activity is also detected in the VLPs1. These results suggest that VLPs and copia are derivatives of viral particle and provirus forms, respectively, of the copia retrovirus-like particle. If the copia retrovirus-like particle replicates by a mechanism similar to the mechanism of vertebrate retroviral replication, a cellular transfer RNA would prime synthesis of the first DNA strand. We show that this is indeed so but that copia retrovirus-like particle has a novel type of priming mechanism; the first DNA extension does not start from the 3′ end of a tRNA, but from an internal site (two nucleotides after the anticodon loop) of the Drosophila initiator methionine tRNA (tRNAiMet)
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Shiba, T. & Saigo, K. Nature 302, 119–124 (1983).
Emori, Y. et al. Nature 316, 773–776 (1985).
Varmus, H. & Swanstrom, R. in Molecular Biology of Tumor Viruses 2nd edn RNA Tumor Viruses (eds Weiss, R., Teich, N., Varmus, H. & Coffin, J.) 369–512 (Cold Spring Harbor Laboratory, New York, 1982).
Dahlberg, J. E. et al. J. Virol. 13, 1126–1133 (1974).
Harada, F., Sawyer, R. C. & Dahlberg, J. E. J. biol. Chem. 250, 3487–3497 (1975).
Harada, F., Peters, G. G. & Dahlberg, J. E. J. biol. Chem. 254, 10979–10985 (1979).
Peters, G. et al. J. Virol. 21, 1031–1041 (1977).
Taylor, J. M. Biochim. biophys. Acta 473, 57–71 (1977).
Peters, G. & Glover, C. J. Virol. 35, 31–40 (1980).
Ono, M. & Ohishi, H. Nucleic Acids Res. 11, 7169–7179 (1983).
Hodgson, C. P., Elder, P. K., Ono, T., Foster, D. N. & Getz, M. J. Molec. cell. Biol. 3, 2221–2231 (1983).
Ou, C., Boone, L. R. & Yang, W. K. Nucleic Acids Res. 11, 5603–5620 (1983).
Colicelli, J. & Goff, S. P. J. Virol. 57, 37–45 (1986).
Inouye, S., Saigo, K., Yamada, K. & Kuchino, Y. Nucleic Acids Res. 14, 3031–3043 (1986).
Donis-Keller, H., Maxam, A. M. & Gilbert, W. Nucleic Acids Res. 4, 2527–2538 (1977).
Donis-Keller, H. Nucleic Acids Res. 8, 3133–3142 (1980).
Krupp, G. & Gross, H. J. in The Modified Nucleotides in Transfer RNA II: A Laboratory Manual of Genetic Analysis, Identification and Sequence Determination (eds Agris, P. F. & Kopper, R. A.) 11–58 (Liss, New York, 1983).
Silverman, S. et al. Nucleic Acids Res. 6, 421–433 (1979).
Flavell, A. J. & Ish-Horowicz, D. Cell 34, 415–419 (1983).
Flavell, A. J. & Brierley, C. Nucleic Acids Res. 14, 3659–3669 (1986).
Hung, P. P. Virology 51, 287–296 (1973).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Kikuchi, Y., Ando, Y. & Shiba, T. Unusual priming mechanism of RNA-directed DNA synthesis in copia retrovirus-like particles of Drosophila. Nature 323, 824–826 (1986). https://doi.org/10.1038/323824a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/323824a0
This article is cited by
-
Identification and occurrence of the LTR-Copia-like retrotransposon, PSCR and other Copia-like elements in the genome of Phytophthora sojae
Current Genetics (2009)
-
SplitTester : software to identify domains responsible for functional divergence in protein family
BMC Bioinformatics (2005)
-
RNase P as hyperprocessing enzyme: A model for formation of a biologically functional tRNA fragment
Molecular Biology Reports (1996)
-
Yeast retrotransposon revealed
Nature (1992)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.