Abstract
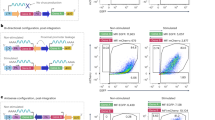
Interleukin-3 (multi-CSF) is a multilineage haematopoietic growth regulator that initiates the proliferation and differentiation of multipotential stem cells1–5. Complementary DNA clones encoding interleukin-3 (IL-3) have recently been isolated6,7 and the structure of the IL-3 gene determined8,9. IL-3 is produced by T lymphocytes or T lymphomas only after stimulation with antigens, mitogens or chemical activators such as phorbol esters. The myelomonocytic leukaemia line WEHI-3B10,11 also produces IL-3 but its production is constitutive12–14 and the WEHI-3B cells do not appear to produce significant levels of any of the other lymphokines normally secreted by T lymphocytes after stimulation. It has been proposed that the genetic change leading to the constitutive expression of IL-3 may have been a key event in the development of this leukaemia2,3,15,16. We report here that the constitutive synthesis of IL-3 by the WEHI-3B cell line is due to the insertion of an endogenous retrovirus-like element close to the 5′ end of the gene. The insertion, an intracisternal A particle (IAP) genome, is positioned with its 5′ long terminal repeat (LTR) close to the promoter region of the IL-3 gene, resulting in constitutive synthesis of IL-3.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Nicola, N. A. & Vadas, M. Immun. Today 5, 76–80 (1984).
Iscove, N. N. & Roitsch, C. Proc. IVth Int. Lymphokine Workshop (Academic, New York, 1985).
Garland, J. M. Lymphokines 9, 153–200 (1984).
Hapel, A. J. et al. Blood 65, 1453–1459 (1985).
Rennick, D. M. et al. J. Immun. 134, 910–914 (1985).
Fung, M. C. et al. Nature 307, 233–237 (1984).
Yokota, T. et al. Proc. natn. Acad. Sci. U.S.A. 81, 1070–1074 (1984).
Campbell, H. D., Ymer, S., Fung, M.-C. & Young, I. G. Eur. J. Biochem. 150, 297–304 (1985).
Miyatake, S., Yokota, T., Lee, F. & Arai, K-I. Proc. natn. Acad. Sci. U.S.A. 82, 316–320 (1985).
Warner, N. L., Moore, M. A. S. & Metcalf, D. J. natn. Cancer Inst. 43, 963–982 (1969).
Metcalf, D. & Nicola, N. A. Int. J. Cancer 30, 773–780 (1982).
Metcalf, D., Moore, M. A. S. & Warner, N. L. J. natn. Cancer Inst. 43, 983–1001 (1969).
Ralph, P., Moore, M. A. S. & Nilsson, K. J exp. Med. 143, 1528–1533 (1976).
Lee, J. C., Hapel, A. J. & Ihle, J. N. J. Immun. 128, 2393–2398 (1982).
Dexter, T. M. & Allen, T. D. J. Path. 141, 415–433 (1983).
Schrader, J. W. & Crapper, R. M. Proc. natn. Acad. Sci. U.S.A. 80, 6892–6896 (1983).
Frischauf, A.-M., Lehrach, H., Poustka, A. & Murray, N. J. molec. Biol. 170, 827–842 (1983).
Benton, W. D. & Davis, R. W. Science 196, 180–182 (1977).
Messing, J. & Vieira, J. Gene 19, 269–276 (1982).
Canaani, E. et al. Proc. natn. Acad. Sci. U.S.A. 80, 7118–7122 (1983).
Christy, R. J., Brown, A. R., Gourlie, B. B. & Huang, R. C. C. Nucleic Acids Res. 13, 289–302 (1985).
Kuff, E. L. et al. Proc. natn. Acad. Sci. U.S.A. 80, 1992–1996 (1983).
Ono, M. & Ohishi, H. Nucleic Acids Res. 11, 7169–7179 (1983).
Graham, F. L. & Bacchetti, S. in Techniques in the Life Sciences Vol. B 5 (Elsevier, Dublin, 1983).
Ihle, J. N. et al. J. Immun. 129, 1377–1383 (1982).
Hapel, A. J., Warren, H. S. & Hume, D. A. Blood 64, 786–790 (1984).
Varmus, H. E. Science 216, 812–820 (1982).
Khoury, G. & Gruss, P. Cell 33, 313–314 (1983).
Horowitz, M., Luria, S., Rechavi, G. & Givol, D. EMBO J. 3, 2937–2941 (1984).
Kuff, E. L. & Fewell, J. W. Molec. cell. Biol. 5, 474–483 (1985).
Augenlicht, L. M., Kobrin, D., Pavlovec, A. & Royston, M. E. J. biol. Chem. 259, 1842–1847 (1984).
Heldin, C.-H. & Westermark, B. Cell 37, 9–20 (1984).
Sporn, M. B. & Roberts, A. B. Nature 313, 745–747 (1985).
Todaro, G. J. & De Larco, J. E. Cancer Res. 38, 4147–5154 (1978).
Derynck, R., Roberts, A. B., Winkler, M. E., Chen, E. Y. & Goeddel, D. V. Cell 38, 287–297 (1984).
Lee, D. C., Rose, T. M., Webb, N. R. & Todaro, G. J. Nature 313, 489–491 (1985).
Waterfield, M. D. et al. Nature 304, 35–39 (1983).
Doolittle, R. F. et al. Science 221, 275–277 (1983).
Downward, J. et al. Nature 307, 521–527 (1984).
Metcalf, D. Science 229, 16–22 (1985).
Sanger, F., Nicklen, S. & Coulson, A. R. Proc. natn. Acad. Sci. U.S.A. 74, 5463–5467 (1977).
Sariger, F., Coulson, A. R., Hong, G. F., Hill, D. F. & Peterson, G. B. J. molec. Biol. 162, 729–773 (1982).
Duckworth, M. L. et al. Nucleic Acids Res. 9, 1681–1706 (1981).
Tanaka, T. & Letsinger, R. L. Nucleic Acids Res. 10, 3249–3260 (1982).
McBride, L. J. & Caruthers, M. H. Tetrahedron Lett. 24, 245–248 (1983).
Staden, R. Nucleic Acids Res. 8, 3673–3694 (1980); 10, 4731–4751 (1982); 12, 499–503 (1984).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ymer, S., Tucker, W., Sanderson, C. et al. Constitutive synthesis of interleukin-3 by leukaemia cell line WEHI-3B is due to retroviral insertion near the gene. Nature 317, 255–258 (1985). https://doi.org/10.1038/317255a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/317255a0
This article is cited by
-
BH3 mimetics efficiently induce apoptosis in mouse basophils and mast cells
Cell Death & Differentiation (2018)
-
IL-4 enhances survival of in vitro-differentiated mouse basophils through transcription-independent signaling downstream of PI3K
Cell Death & Disease (2018)
-
Animal models of acute myelogenous leukaemia – development, application and future perspectives
Leukemia (2005)
-
Stroma-mediated granulocyte-macrophage colony-stimulating factor (GM-CSF) control of myelopoiesis: spatial organisation of intercellular interactions
Cell and Tissue Research (2003)
-
Deregulated c-Myc prematurely recruits both Type I and II CD95/Fas apoptotic pathways associated with terminal myeloid differentiation
Oncogene (2002)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.