Abstract
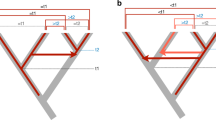
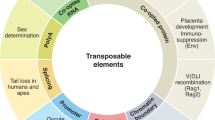
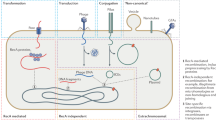
It has recently been demonstrated both emperically1–3 and mathematically4 that transposable elements may spread rapidly throughout a population once introduced even when they dramatically reduce the fitness of individuals that carry them. Such events result in pronounced differences in the phylogenetic distribution of genetic elements capable of rapid genome invasion. Using a simple and general procedure to screen the genome of the white footed mouse Peromyscus leucopus, we have isolated a family of retrovirus-like elements which is apparently absent from the genome of the house mouse Mus domesticus. Here, we report this procedure and an analysis of the organization, phylogenetic distribution and sequence of this family of transposable elements.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Kidwell, M. G. Proc. natn. Acad. Sci. U.S.A. 80, 1655–1659 (1983).
Bregliano, J. C. & Kidwell, M. G. in Mobile Genetic Elements, 363–410 (ed. Shapiro, J. A.) (Academic, New York, 1983).
Engels, W. R. A. Rev. Genet. 17, 315–344 (1983).
Hickey, D. Genetics 101, 519–531 (1982).
Wichman, H. A. thesis, Wesleyan Univ. (1983).
Gilboa, E., Mitra, S. W., Goff, S. & Baltimore, D. Cell 18, 93–100 (1979).
Sprinzl, M. & Gaucs, D. H. Nucleic Acids Res. 10, r1–r55 (1982).
Toh, H., Hayashida, H. & Miyata, T. Nature 305, 827–829 (1983).
Saigo, K. et al. Nature 312, 659–661 (1984).
Lueders, K. K., Segal, S. & Kuff, E. L. Cell 11, 83–94 (1977).
Hawley, M. J., Shulman, M. J. & Hozumi, N. Molec. cell. Biol. 4, 2565–2572 (1984).
Cohen, J. B., Unger, T., Rechavi, G., Cananni, E. & Givol, D. Nature 306, 797–799 (1983).
Augenlicht, L. H., Kobrin, D., Pavlovec, A. & Royston, M. E. J. biol. Chem. 259, 1842–1847 (1984).
Levis, R., Dunsmuir, P. & Rubin, G. M. Cell 21, 581–588 (1980).
Wirth, T. W., Gloggler, K., Baumruker, T., Schmidt, M. & Horak, I. Proc. natn. Acad. Sci. U.S.A. 80, 3327–3330 (1983).
Wirth, T., Schmidt, M., Baumruker, T. & Horak, I. Nucleic Acids Res. 12, 3603–3610 (1984).
Norton, J. D., Connor, J. & Avery, R. J. Nucleic Acids Res. 12, 3445–3460 (1984).
Brulet, P., Kaghad, M., Xu, Y., Croissant, O. & Jacob, F. Proc. natn. Acad. Sci. U.S.A. 80, 5641–5645 (1983).
Baltimore, D. Cell 40, 481–48nd, 1034–1051 (1983).
Maxam, A. & Gilbert, W. Meth. Enzym. 65, 499–560 (1980).
Shinnick, T. M., Lerner, R. A. & Sutcliffe, J. G. Nature 293, 543–548 (1981).
Schwartz, D., Tizard, R. & Gilbert, W. Cell 32, 853–869 (1983).
Gardner, E. C. et al. Nucleic Acids Res. 9, 2871–2888 (1981).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wichman, H., Potter, S. & Pine, D. Mys, a family of mammalian transposable elements isolated by phylogenetic screening. Nature 317, 77–81 (1985). https://doi.org/10.1038/317077a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/317077a0
This article is cited by
-
Analysis of Orthologous Retrovirus-Like Elements in the White-Footed Mouse, Peromyscus leucopus
Journal of Molecular Evolution (1997)
-
Retrotransposon Mys was active during evolution of the Peromyscus leucopus-maniculatus complex
Journal of Molecular Evolution (1996)
-
Computer simulation of transposable element evolution: Random template and strict master models
Journal of Molecular Evolution (1996)
-
Rapidly evolving repetitive DNAs in a conservative genome: A test of factors that affect chromosomal evolution
Chromosome Research (1994)
-
Comparison of chromosomal distribution of a retroposon (LINE) and a retrovirus-like elementmys inPeromyscus maniculatus andP. leucopus
Chromosome Research (1994)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.