Abstract
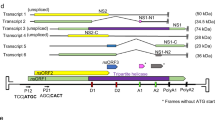
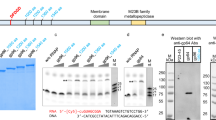
We have shown previously that Drosophila cells contain virus-like particles (VLPs) containing 5-kilobase (kb) RNA that hybridizes to a transposable element, termed copia. We have suggested that VLPs and copia are derivatives of viral particles and proviral forms, respectively, of ‘copia’ retrovirus, a putative Drosophila retrovirus1. To further clarify the relationship between copia and copia-related RNA in VLPs (VLP H-RNA), we determined and compared their nucleotide sequences. VLP H-RNA was found to be an unspliced, genome-sized transcript of copia, and, like retroviral genome RNA, VLP H-RNA is terminally redundant with termini localized in the long terminal repeats (LTRs) of copia. VLP H-RNA contains two long open reading frames (ORFs), one of which includes the coding sequence for a predominant VLP protein of relative molecular mass (Mr) 31,000 (31K). Here we show that, in contrast to 17.6 ORF2 (ref. 2), ORFs of copia have no extensive amino-acid sequence homology to the RT region2 of the reverse transcriptase of retrovirus in vertebrates. Because of a one-base insertion/deletion, the two ORFs in VLP H-RNA are fused and become a single, longer ORF in a genomic copia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Shiba, T. & Saigo, K. Nature 302, 119–124 (1983).
Saigo, K. et al. Nature 312, 659–661 (1984).
Saigo, K., Millstein, L. & Thomas, C. A. Jr Cold Spring Harb. Symp. quant. Biol. 45, 815–827 (1981).
Maniatis, T. et al. Cell 16, 687–701 (1978).
Rubin, G. M. et al. Cold Spring Harb Symp. quant. Biol. 45, 619–628 (1981).
Flavell, A. J., Levis, R., Simon, M. A. & Rubin, G. M. Nucleic Acids Res. 9, 6267–6291 (1981).
Fouts, D. L., Manning, J. E., Fox, G. M. & Schmid, C. W. Nucleic Acid Res. 9, 7053–7064 (1981).
Flavell, A. J. Nature 310, 514–516 (1984).
Toh, H., Hayashida, H. & Miyata, T. Nature 305, 827–829 (1983).
Toh, H. et al. EMBO J. (in the press).
Kamer, G. & Argos, P. Nucleic Acids Res. 12, 7269–7282 (1984).
Kugimiya, W., Ikenaga, H. & Saigo, K. Proc. natn. Acad. Sci. U.S.A. 80, 3193–3197 (1983).
Vieira, J. & Messing, J. Gene 19, 259–268 (1982).
Birnboim, H. C. & Doly, J. Nucleic Acids Res. 7, 1513–1523 (1979).
Sanger, F., Nickelen, S. & Coulson, A. R. Proc. natn. Acad. Sci. U.S.A. 74, 5463–5467 (1977).
Maniatis, T., Fritsch, E. F. & Sambrook, J. Molecular Cloning. A Laboratory Manual (Cold Spring Harbor Laboratory, New York, 1982).
Maxam, A. M. & Gilbert, W. Meth. Enzym. 65, 499–560 (1980).
Treisman, R., Proudfoot, N. J., Shander, M. & Maniatis, T. Cell 29, 903–911 (1982).
Koike, T. & Ikenaka, T. Eur. J. Biochem. 32, 401–407 (1973).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Emori, Y., Shiba, T., Kanaya, S. et al. The nucleotide sequences of copia and copia-related RNA in Drosophila virus-like particles. Nature 315, 773–776 (1985). https://doi.org/10.1038/315773a0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/315773a0
This article is cited by
-
Transcriptionally promiscuous “blurry” promoters in Tc1/mariner transposons allow transcription in distantly related genomes
Mobile DNA (2019)
-
BEL/Pao retrotransposons in metazoan genomes
BMC Evolutionary Biology (2011)
-
Envelope-Like Retrotransposons in the Plant Kingdom: Evidence of Their Presence in Gymnosperms (Pinus pinaster)
Journal of Molecular Evolution (2008)
-
Sequence heterogeneity and phylogenetic relationships between the copia retrotransposon in Drosophila species of the repleta and melanogaster groups
Genetics Selection Evolution (2006)
-
A full lengthTy3/gypsy-type retrotransposon in the fernAdiantum
Journal of Plant Research (1997)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.