Abstract
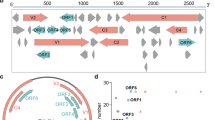
Only two groups of plant viruses, the caulimoviruses1,2 and the geminiviruses3, are known to contain a genome of DNA. Unlike that of the caulimoviruses, the genome of the geminivinises is composed of single-stranded, covalently-closed circles of DNA. There is evidence that the geminiviruses, specifically bean golden mosaic virus4 and tomato golden mosaic virus5, have a genome composed of two similar-sized circles of DNA, and the molecular cloning of both components from tomato golden mosaic virus has recently been reported6. No information is available, however, concerning the protein coding capacity of the genome, the possible modes of UNA transcription and DNA replication and the assembly of mature components into the characteristic geminate particles. We report here on the derivation of the nucleotide sequence of the geminivirus, cassava latent virus (CLV), as a fundamental step towards the investigation of the mode of replication of this group of viruses. Results show that CLV DNA comprises two similar-sized molecular species with a common region encompassing almost 200 nucleotides. The organization of the genome is discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Shepherd, R. J. A. Rev. Pl. Physiol. 30, 405–423 (1979).
Hull, R. in Nucleic Acids in Plants (eds Hall, T. C. & Dovies, J. W.) 3–29 (CRC Press, Boca Raton, Florida, 1979).
Goodman, R. M. Handbook of Plant Virus Infections and Comparative Diagnosis (ed. Kurstak, E.) 879–910 (Elsevier, Amsterdam, 1981).
Haber, S., Ikegami, M., Bajet, N. B. & Goodman, R. M. Nature 289, 324–326 (1981).
Hamilton, W. D. O., Bisaro, D. M. & Buck, K. W. Nucleic Acids Res. 10, 4901–4912 (1982).
Bisaro, D. M., Hamilton, W. D. O., Coutts, R. H. A. & Buck, K. W. Nucleic Acids Res. 10, 4913–4922 (1982).
Harrison, B. D. et al. Nature 270, 760–762 (1977).
Ikegami, M., Haber, S. & Goodman, R. M. Proc. natn. Acad. Sci. U.S.A. 78, 4102–4106 (1981).
Sanger, H. L. J. Virol. 3, 304–312 (1969).
Goldbach, R., Rezelman, G. & van Kammen, A. Nature 286, 297–300 (1980).
Robinson, D. J., Barker, H., Harrison, B. D. & Mayo, M. A. J. gen. Virol. 51, 317–326 (1980).
Sequeira, J. C. & Harrison, B. D., appl. Biol. 101, 33–42 (1982).
Horiuchi, K. & Zinder, N. D. Proc. natn. Acad. Sci. U.S.A. 72, 2555–2558 (1975).
Blakesley, R. W. & Wells, R. D. Nature 257, 421–422 (1975).
Maxam, A. M. & Gilbert, W., Enzym. 65, 499–560 (1980).
Tu, C-P. D. & Cohen, S. N. Gene 10, 177–183 (1980).
Hong, G. F. Biosci. Rep. 1, 243–252 (1981).
Messing, J., Crea, R. & Seeburg, P. H. Nucleic Acids Res. 9, 309–321 (1981).
Sanger, F., Coulson, A. R., Barrell, B. G., Smith, A. J. H. & Roe, B. A. J. molec. Biol. 143, 161–178 (1980).
Duckworth, M. L. et al. Nucleic Acids Res. 9, 1691–1706 (1981).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Stanley, J., Gay, M. Nucleotide sequence of cassava latent virus DNA. Nature 301, 260–262 (1983). https://doi.org/10.1038/301260a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/301260a0
This article is cited by
-
Asymptomatic populus alba: a tree serving as a reservoir of begomoviruses and associated satellites
Australasian Plant Pathology (2022)
-
Interaction of a tomato leaf curl New Delhi virus with a betasatellite enhances symptom severity in field-infected tomato plants
Tropical Plant Pathology (2021)
-
Cassava mosaic disease: a review of a threat to cassava production in Zambia
Journal of Plant Pathology (2019)
-
Divergent evolutionary and epidemiological dynamics of cassava mosaic geminiviruses in Madagascar
BMC Evolutionary Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.