Abstract
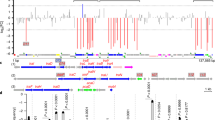
The transposable genetic element Tn9 consists of two direct repeats of the insertion sequence IS1 flanking a region of 1,102 base pairs which determines chloramphenicol resistance1,2. Transposition of Tn9 leads to the duplication of a 9-base pair sequence which preexists at the site of insertion. One copy of this sequence is found at each end of the inserted element3. The chloramphenicol resistance determined by TnP, and by various other R plasmids, is due to the synthesis of the enzyme chloramphenicol acetyl transferase (CAT)4,5. This enzyme catalyses the formation of acetylated derivatives of chloramphenicol which are inactive as inhibitors of protein synthesis5. By using the chain termination technique of DNA sequencing, we have now determined the nucleotide sequence of the 1,102 base pair region between the directly repeated IS1 sequences in the bacterial transposon Tn9 (encoding chloramphenicol resistance). The amino acid sequence of CAT predicted from the nucleotide sequence is identical to that determined by Shaw and coworkers6. An analysis of the sequence suggests that the internal 1,102 base pair region is not directly involved in transposition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Chow, L. T. & Bukhari, A. I. in DNA, Insertion Elements, Plasmids and Episomes (eds Bukhari, A. I., Shapiro, J. A. & Adhya, S. L.) 295–306 (Cold Spring Harbor Laboratory, New York, 1977).
MacHattie, L. A. & Jackowski, J. B. in DNA, Insertion Elements, Plasmids and Episomes (eds Bukhari, A. I., Shapiro, J. A. & Adhya, S. L.) 219–228 (Cold Spring Harbor Laboratory, New York, 1977).
Johnsrud, L., Calos, M. P. & Miller, J. H. Cell 15, 1209–1219 (1978).
Shaw, W. V. & Brodsky, R. F. J. Bact. 95, 28–36 (1968).
Shaw, W. V. J. biol. Chem. 242, 687–693 (1967).
Shaw, W. V. et al. Nature 282, 870–872 (1979).
Iida, S., Meyer, J. & Arber, W. Plasmid 1, 357–365 (1978).
Sanger, F., Nicklen, S. & Coulson, A. R. Proc. natn. Acad. Sci. U.S.A. 74, 5463–5467 (1977).
Smith, A. J. H. Nucleic Acids Res. 6, 831–848 (1979).
Messing, J., Gronenborn, B., Muller-Hill, B. & Hofschneider, P. H. Proc. natn. Acad. Sci. U.S.A. 74, 3642–3646 (1977).
Marvin, D. A. & Hohn, B. Bact. Rev. 33, 172–209 (1969).
Gronenborn, B. & Messing, J. Nature 272, 375–377 (1978).
Ray, D. S. & Kook, K. Gene 4, 109–119 (1978).
Barnes, W. M. Gene 5, 127–139 (1979).
Ohtsubo, H. & Ohtsubo, E. Proc. natn. Acad. Sci. U.S.A. 75, 615–619 (1978).
Johnsrud, L. Molec. gen. Genet. 169, 213–218 (1979).
Bolivar, F. Gene 4, 121–136 (1978).
Chang, A. C. Y. & Cohen, S. N. J. Bact. 134, 1141–1156 (1978).
Shine, J. & Dalgarno, L. Proc. natn. Acad. Sci. U.S.A. 71, 1342–1346 (1974).
Steitz, J. A. & Jakes, K. Proc. natn. Acad. Sci. U.S.A. 72, 4734–4738 (1975).
Godson, G. N., Barrell, B. G., Staden, R. & Fiddes, J. C. Nature 276, 236–247 (1978).
Steitz, J. A. in Biological Regulation and Control (ed. Goldberger, R.) (Plenum, New York, in the press).
Harwood, J. & Smith, D. H. Biochem. biophys. Res. Commun. 42, 57–62 (1971).
deCrombrugghe, B., Pastan, I., Shaw, W. V. & Rosner, J. L. Nature new Biol. 241, 237–239 (1973).
Reznikoff, W. S. & Abelson, J. N. in The Operon (eds Miller, J. H. & Reznikoff, W. S.) 221–243 (Cold Spring Harbor Laboratory, New York, 1978).
deCrombrugghe, B. & Pastan, I. in The Operon (eds Miller, J. H. & Reznikoff, W. S.) 303–324 (Cold Spring Harbor Laboratory, New York, 1978).
Musso, R. E., Dilauro, R., Adhya, S. & deCrombrugghe, B. Cell 12, 847–854 (1977).
Greenfield, L., Boone, T. & Wilcox, G. Proc. natn. Acad. Sci. U.S.A. 75, 4724–4728 (1978).
Pribnow, D. Proc. natn. Acad. Sci. U.S.A. 72, 784–788 (1975).
Gilbert, W. in RNA Polymerase (eds Losick, R. & Chamberlin, M.) 193–205 (Cold Spring Harbor Laboratory, New York, 1976).
Adhya, S. & Gottesman, M. A. Rev. Biochem. 47, 967–996 (1978).
Arber, W. et al. Cold Spring Harb. Symp. quant. Biol. 43, 1197–1208 (1978).
Davies, J. & Smith, D. I. A. Rev. Microbiol. 34, 469–518 (1978).
So, M., Heffron, F. & McCarthy, B. J. Nature 277, 453–456 (1979).
Heffron, F., Bedinger, P., Champoux, J. & Falkow, S. in DNA, Insertion Elements, Plasmids and Episomes (eds Bukhari, A. I., Shapiro, J. A. & Adhya, S. L.) 161–167 (Cold Spring Harbor Laboratory, New York, 1977).
Alton, N. K. & Vapnek, D. Plasmid 2, 366–376 (1979).
Alton, N. K. & Vapnek, D. Plasmid 1, 388–404 (1978).
Smith, H. O. & Birnsteil, M. L. Nucleic Acids Res. 3, 2387–2398 (1976).
Yajko, D. M. & Weis, B. Proc. natn. Acad. Sci. U.S.A. 72, 688–692 (1975).
Maxam, A. & Gilbert, W. Proc. natn. Acad. Sci. U.S.A. 74, 560–564 (1977).
Sanger, F. & Coulson, A. R. FEBS Lett. 87, 107–110 (1978).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Alton, N., Vapnek, D. Nucleotide sequence analysis of the chloramphenicol resistance transposon Tn9. Nature 282, 864–869 (1979). https://doi.org/10.1038/282864a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/282864a0
This article is cited by
-
The bacterial Tn9 chloramphenicol resistance gene: an attractive DNA segment for Mos1 mariner insertions
Molecular Genetics and Genomics (2009)
-
Transformation and gene replacement in the facultatively chemoheterotrophic, unicellular cyanobacterium Synechocystis sp. PCC6714 by electroporation
Applied Microbiology and Biotechnology (2008)
-
Heterologous gene expression in Thermus thermophilus: β-galactosidase, dibenzothiophene monooxygenase, PNB carboxy esterase, 2-aminobiphenyl-2,3-diol dioxygenase, and chloramphenicol acetyl transferase
Journal of Industrial Microbiology and Biotechnology (2004)
-
Plant selectable markers and reporter genes
Acta Physiologiae Plantarum (2001)
-
Efficient hammerhead ribozyme and antisense RNA targeting in a slow ribosome Escherichia colimutant
Nature Biotechnology (1997)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.