Abstract
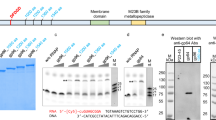
THE temperate bacteriophage Mu has the remarkable ability to insert its DNA in apparently random sites of the Escherichia coli chromosome (1–3). All Mu prophages have the same gene order4, and the finding that the prophage and the phage maps are identical suggests that specific sites (attachment sites) at the ends of the Mu genome are used for the integration of Mu5. Phage particles do not contain the excised, free form of the Mu DNA; instead, the Mu genome is flanked by what seems to be heterogeneous bacterial DNA6,7. The variable DNA at both ends of Mu is thought to be generated when Mu DNA is transposed to many new chromosomal sites during replication and is subsequently packaged into phage particles together with adjacent host DNA. The nucleotide sequence analysis of the ends of Mu DNA reported here substantiates this hypothesis and identifies the attachment sites of Mu. Structural features of the attachment sites are presented.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Taylor, A. J. Proc. natn. Acad. Sci. U.S.A. 50, 1043–1051 (1963).
Bukhari, A. I. & Zipser, D. Nature new Biol. 236, 240–243 (1972).
Daniell, E., Roberts, R. J. & Abelson, J. J. molec. Biol. 69, 1–8 (1972).
Abelson, J. et al. Virology 54, 90–92 (1973).
Hsu, M.-T. & Davidson, N. Virology 58, 229–239 (1974).
Daniell, E., Kohne, D. E. & Abelson, J. J. Virol. 15, 739–743 (1975).
Bukhari, A. I., Froshauer, S. & Botchan, M. Nature 264, 580–583 (1976).
Zipser, D., Moses, P., Kahmann, R. & Kamp, D. Gene 2, 263–271 (1977).
Maxam, A. & Gilbert, W. Proc. natn. Acad. Sci. U.S.A. 74, 560–564 (1977).
O'Day, K. J., Schultz, D. W. & Howe, M. M. Microbiology (ed. Schlessinger, D.), 48–51 (American Society for Microbiology, 1978).
Ohtsubo, H. & Ohtsubo, E. Proc. natn. Acad. Sci. U.S.A. 75, 615–619.
Grindley, N. D. F. Cell 13, 419–426 (1978).
Rosenberg, M., Court, D., Shimatake, H., Brady, C. & Wulff, D. L. Nature 272, 414 (1978).
Calos, M. P., Johnsrud, L. & Miller, J. H. Cell 13, 411–418 (1978).
Ohtsubo, H., Ohmori, H. & Ohtsubo, E. Cold Spring Harb. Symp. quant. Biol. 43 (1979).
Kamp, D. et al. Cold Spring Harb. Symp. quant. Biol. 43 (1979).
Allett, B. Cell 16, 123–129 (1979).
Allet, B. Nature 274, 553–558 (1978).
Kahmann, R., Kamp, D. & Zipser, D. DNA Insertion Elements, Plasmids and Episomes (eds Bukhari, A. I., Shapiro, J. A. & Adhya, S. L.) (Cold Spring Harbor, New York, 1977).
Chow, L. T., Kahmann, R. & Kamp, D. J. molec. Biol. 113, 591–609 (1977).
Chow, L. T. & Broker, T. R. J. Bact. 133, 1427–1436 (1978).
Bolivar, R. et al. Gene 2, 95–113 (1977).
Murray, N. E. & Murray, K. Nature 251, 476–481 (1974).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
KAHMANN, R., KAMP, D. Nucleotide sequences of the attachment sites of bacteriophage Mu DNA. Nature 280, 247–250 (1979). https://doi.org/10.1038/280247a0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/280247a0
This article is cited by
-
Deep sequencing reveals new roles for MuB in transposition immunity and target-capture, and redefines the insular Ter region of E. coli
Mobile DNA (2020)
-
Core and accessory genome architecture in a group of Pseudomonas aeruginosa Mu-like phages
BMC Genomics (2014)
-
Regulation of bacteriophage Mu transposition
Genetica (1994)
-
The right end of MudI(Ap,lac)
Archives of Microbiology (1989)
-
Nucleotide sequence of the ends of the conjugative shuttle transposon Tn1545
Molecular and General Genetics MGG (1987)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.